Tewodros Ayalew
PROGRESSOR: A Perceptually Guided Reward Estimator with Self-Supervised Online Refinement
Nov 26, 2024



Abstract:We present PROGRESSOR, a novel framework that learns a task-agnostic reward function from videos, enabling policy training through goal-conditioned reinforcement learning (RL) without manual supervision. Underlying this reward is an estimate of the distribution over task progress as a function of the current, initial, and goal observations that is learned in a self-supervised fashion. Crucially, PROGRESSOR refines rewards adversarially during online RL training by pushing back predictions for out-of-distribution observations, to mitigate distribution shift inherent in non-expert observations. Utilizing this progress prediction as a dense reward together with an adversarial push-back, we show that PROGRESSOR enables robots to learn complex behaviors without any external supervision. Pretrained on large-scale egocentric human video from EPIC-KITCHENS, PROGRESSOR requires no fine-tuning on in-domain task-specific data for generalization to real-robot offline RL under noisy demonstrations, outperforming contemporary methods that provide dense visual reward for robotic learning. Our findings highlight the potential of PROGRESSOR for scalable robotic applications where direct action labels and task-specific rewards are not readily available.
Unsupervised Domain Adaptation For Plant Organ Counting
Sep 02, 2020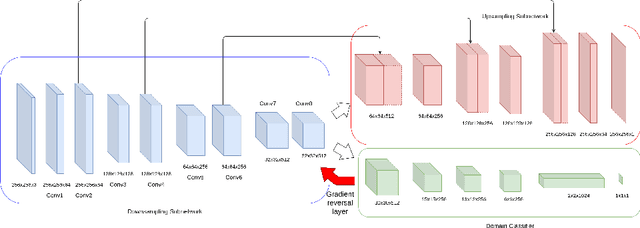
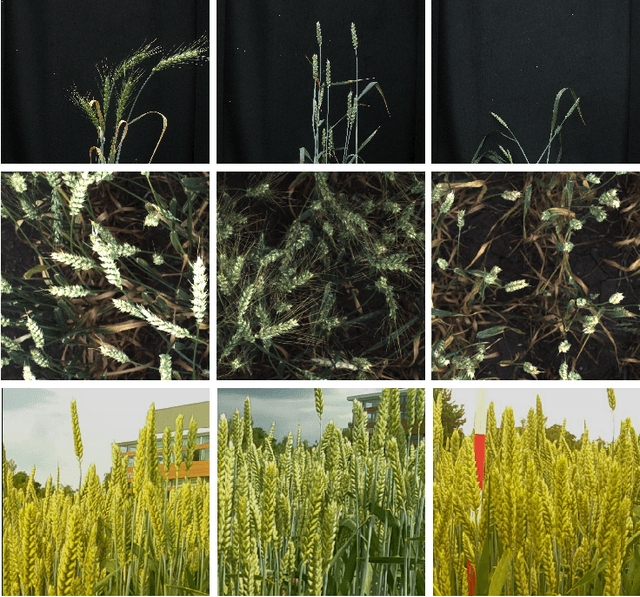
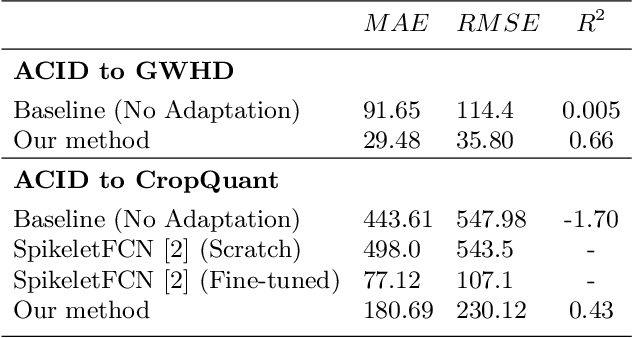

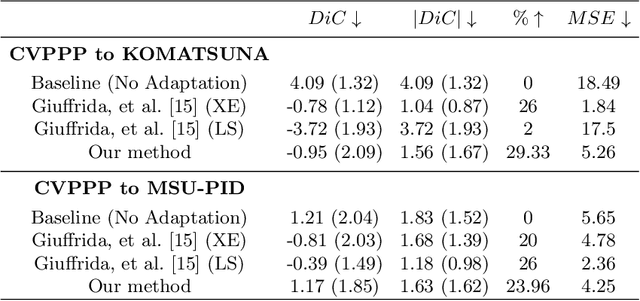
Abstract:Supervised learning is often used to count objects in images, but for counting small, densely located objects, the required image annotations are burdensome to collect. Counting plant organs for image-based plant phenotyping falls within this category. Object counting in plant images is further challenged by having plant image datasets with significant domain shift due to different experimental conditions, e.g. applying an annotated dataset of indoor plant images for use on outdoor images, or on a different plant species. In this paper, we propose a domain-adversarial learning approach for domain adaptation of density map estimation for the purposes of object counting. The approach does not assume perfectly aligned distributions between the source and target datasets, which makes it more broadly applicable within general object counting and plant organ counting tasks. Evaluation on two diverse object counting tasks (wheat spikelets, leaves) demonstrates consistent performance on the target datasets across different classes of domain shift: from indoor-to-outdoor images and from species-to-species adaptation.
AutoCount: Unsupervised Segmentation and Counting of Organs in Field Images
Jul 17, 2020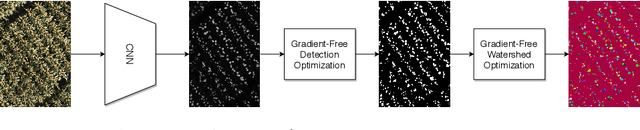
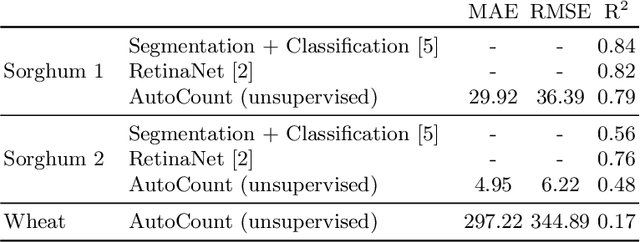
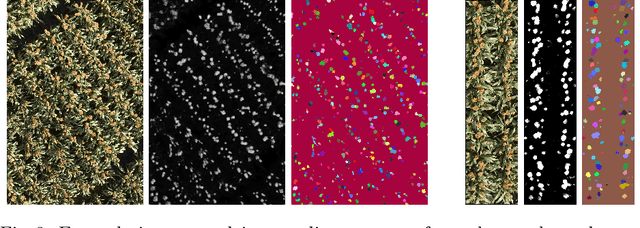

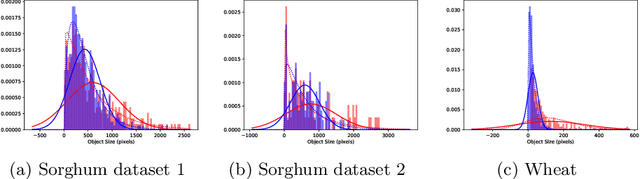

Abstract:Counting plant organs such as heads or tassels from outdoor imagery is a popular benchmark computer vision task in plant phenotyping, which has been previously investigated in the literature using state-of-the-art supervised deep learning techniques. However, the annotation of organs in field images is time-consuming and prone to errors. In this paper, we propose a fully unsupervised technique for counting dense objects such as plant organs. We use a convolutional network-based unsupervised segmentation method followed by two post-hoc optimization steps. The proposed technique is shown to provide competitive counting performance on a range of organ counting tasks in sorghum (S. bicolor) and wheat (T. aestivum) with no dataset-dependent tuning or modifications.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge