Steve Jiang
Medical Artificial Intelligence and Automation Laboratory and Department of Radiation Oncology, UT Southwestern Medical Center, Dallas TX 75235, USA
Exploring the combination of deep-learning based direct segmentation and deformable image registration for cone-beam CT based auto-segmentation for adaptive radiotherapy
Jun 07, 2022
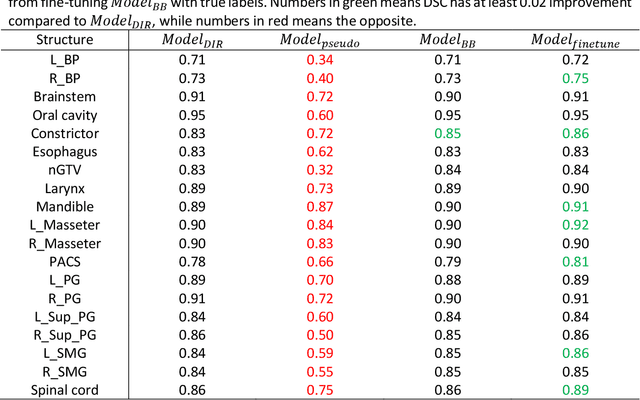
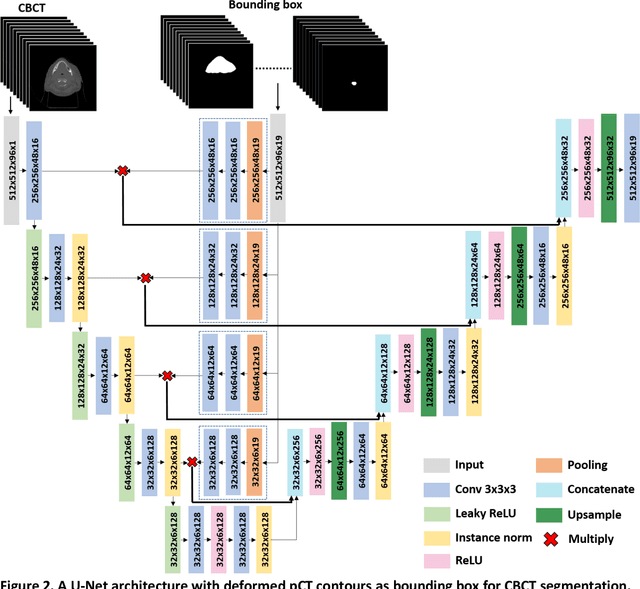
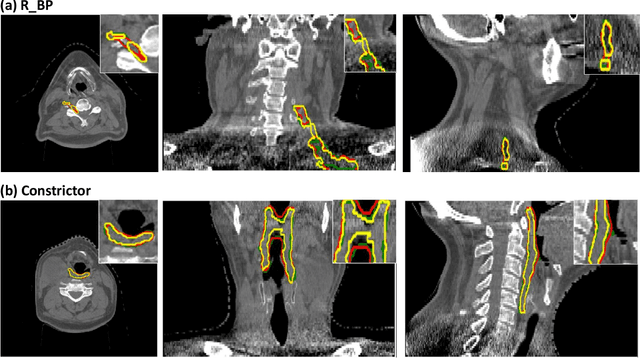
Abstract:CBCT-based online adaptive radiotherapy (ART) calls for accurate auto-segmentation models to reduce the time cost for physicians to edit contours, since the patient is immobilized on the treatment table waiting for treatment to start. However, auto-segmentation of CBCT images is a difficult task, majorly due to low image quality and lack of true labels for training a deep learning (DL) model. Meanwhile CBCT auto-segmentation in ART is a unique task compared to other segmentation problems, where manual contours on planning CT (pCT) are available. To make use of this prior knowledge, we propose to combine deformable image registration (DIR) and direct segmentation (DS) on CBCT for head and neck patients. First, we use deformed pCT contours derived from multiple DIR methods between pCT and CBCT as pseudo labels for training. Second, we use deformed pCT contours as bounding box to constrain the region of interest for DS. Meanwhile deformed pCT contours are used as pseudo labels for training, but are generated from different DIR algorithms from bounding box. Third, we fine-tune the model with bounding box on true labels. We found that DS on CBCT trained with pseudo labels and without utilizing any prior knowledge has very poor segmentation performance compared to DIR-only segmentation. However, adding deformed pCT contours as bounding box in the DS network can dramatically improve segmentation performance, comparable to DIR-only segmentation. The DS model with bounding box can be further improved by fine-tuning it with some real labels. Experiments showed that 7 out of 19 structures have at least 0.2 dice similarity coefficient increase compared to DIR-only segmentation. Utilizing deformed pCT contours as pseudo labels for training and as bounding box for shape and location feature extraction in a DS model is a good way to combine DIR and DS.
Region Specific Optimization (RSO)-based Deep Interactive Registration
Mar 08, 2022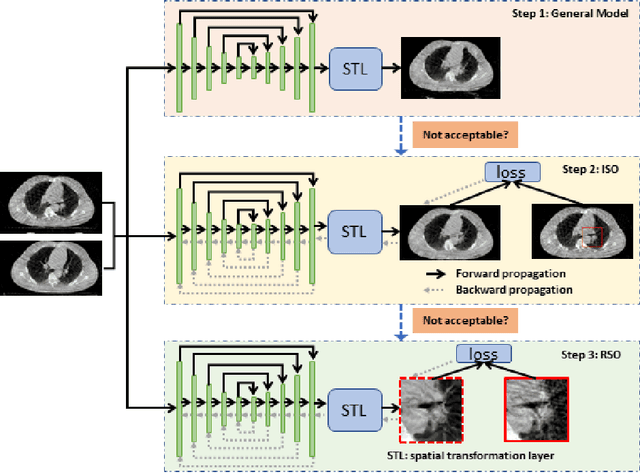
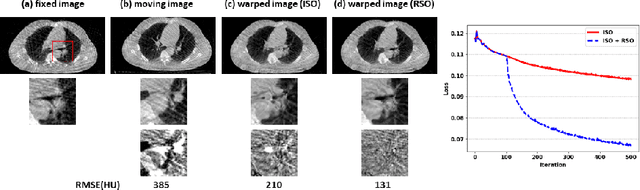

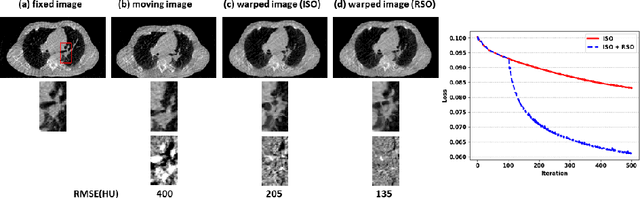
Abstract:Medical image registration is a fundamental and vital task which will affect the efficacy of many downstream clinical tasks. Deep learning (DL)-based deformable image registration (DIR) methods have been investigated, showing state-of-the-art performance. A test time optimization (TTO) technique was proposed to further improve the DL models' performance. Despite the substantial accuracy improvement with this TTO technique, there still remained some regions that exhibited large registration errors even after many TTO iterations. To mitigate this challenge, we firstly identified the reason why the TTO technique was slow, or even failed, to improve those regions' registration results. We then proposed a two-levels TTO technique, i.e., image-specific optimization (ISO) and region-specific optimization (RSO), where the region can be interactively indicated by the clinician during the registration result reviewing process. For both efficiency and accuracy, we further envisioned a three-step DL-based image registration workflow. Experimental results showed that our proposed method outperformed the conventional method qualitatively and quantitatively.
Segmentation by Test-Time Optimization (TTO) for CBCT-based Adaptive Radiation Therapy
Feb 08, 2022



Abstract:Online adaptive radiotherapy (ART) requires accurate and efficient auto-segmentation of target volumes and organs-at-risk (OARs) in mostly cone-beam computed tomography (CBCT) images. Propagating expert-drawn contours from the pre-treatment planning CT (pCT) through traditional or deep learning (DL) based deformable image registration (DIR) can achieve improved results in many situations. Typical DL-based DIR models are population based, that is, trained with a dataset for a population of patients, so they may be affected by the generalizability problem. In this paper, we propose a method called test-time optimization (TTO) to refine a pre-trained DL-based DIR population model, first for each individual test patient, and then progressively for each fraction of online ART treatment. Our proposed method is less susceptible to the generalizability problem, and thus can improve overall performance of different DL-based DIR models by improving model accuracy, especially for outliers. Our experiments used data from 239 patients with head and neck squamous cell carcinoma to test the proposed method. Firstly, we trained a population model with 200 patients, and then applied TTO to the remaining 39 test patients by refining the trained population model to obtain 39 individualized models. We compared each of the individualized models with the population model in terms of segmentation accuracy. The number of patients with at least 0.05 DSC improvement or 2 mm HD95 improvement by TTO averaged over the 17 selected structures for the state-of-the-art architecture Voxelmorph is 10 out of 39 test patients. The average time for deriving the individualized model using TTO from the pre-trained population model is approximately four minutes. When adapting the individualized model to a later fraction of the same patient, the average time is reduced to about one minute and the accuracy is slightly improved.
A Proof-of-Concept Study of Artificial Intelligence Assisted Contour Revision
Jul 28, 2021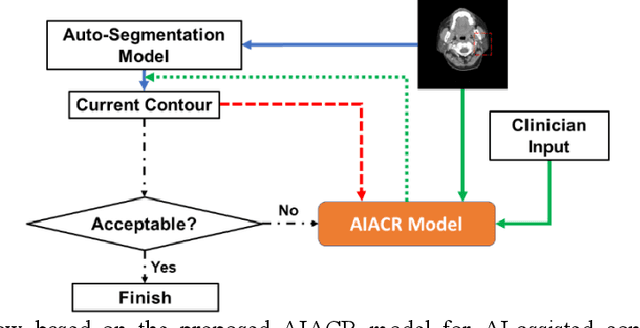

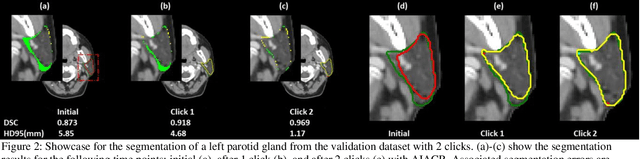
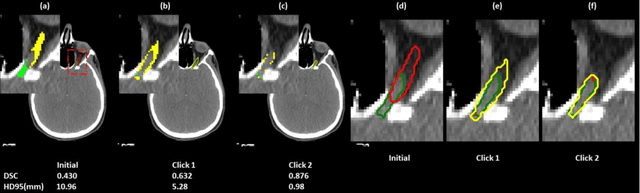
Abstract:Automatic segmentation of anatomical structures is critical for many medical applications. However, the results are not always clinically acceptable and require tedious manual revision. Here, we present a novel concept called artificial intelligence assisted contour revision (AIACR) and demonstrate its feasibility. The proposed clinical workflow of AIACR is as follows given an initial contour that requires a clinicians revision, the clinician indicates where a large revision is needed, and a trained deep learning (DL) model takes this input to update the contour. This process repeats until a clinically acceptable contour is achieved. The DL model is designed to minimize the clinicians input at each iteration and to minimize the number of iterations needed to reach acceptance. In this proof-of-concept study, we demonstrated the concept on 2D axial images of three head-and-neck cancer datasets, with the clinicians input at each iteration being one mouse click on the desired location of the contour segment. The performance of the model is quantified with Dice Similarity Coefficient (DSC) and 95th percentile of Hausdorff Distance (HD95). The average DSC/HD95 (mm) of the auto-generated initial contours were 0.82/4.3, 0.73/5.6 and 0.67/11.4 for three datasets, which were improved to 0.91/2.1, 0.86/2.4 and 0.86/4.7 with three mouse clicks, respectively. Each DL-based contour update requires around 20 ms. We proposed a novel AIACR concept that uses DL models to assist clinicians in revising contours in an efficient and effective way, and we demonstrated its feasibility by using 2D axial CT images from three head-and-neck cancer datasets.
Site-Agnostic 3D Dose Distribution Prediction with Deep Learning Neural Networks
Jun 15, 2021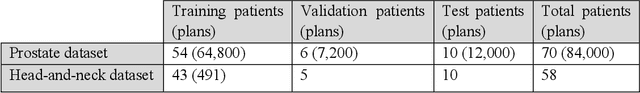
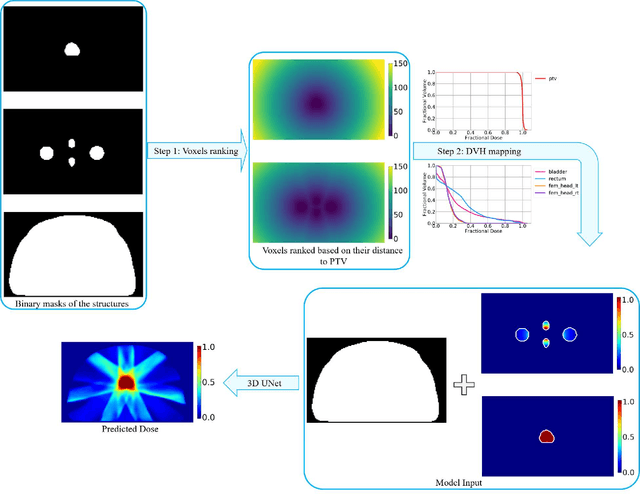
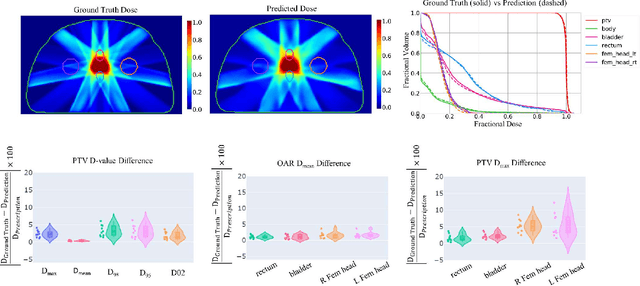
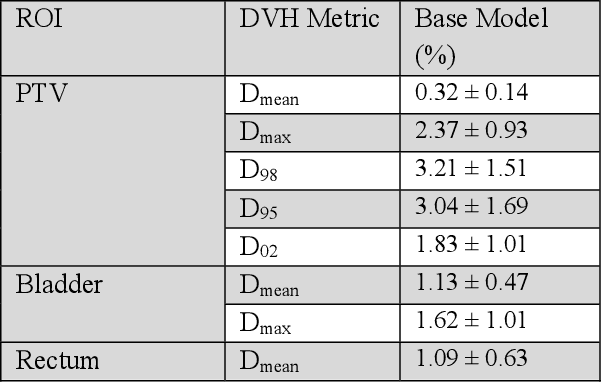
Abstract:Typically, the current dose prediction models are limited to small amounts of data and require re-training for a specific site, often leading to suboptimal performance. We propose a site-agnostic, 3D dose distribution prediction model using deep learning that can leverage data from any treatment site, thus increasing the total data available to train the model. Applying our proposed model to a new target treatment site requires only a brief fine-tuning of the model to the new data and involves no modifications to the model input channels or its parameters. Thus, it can be efficiently adapted to a different treatment site, even with a small training dataset.
Intentional Deep Overfit Learning (IDOL): A Novel Deep Learning Strategy for Adaptive Radiation Therapy
Apr 23, 2021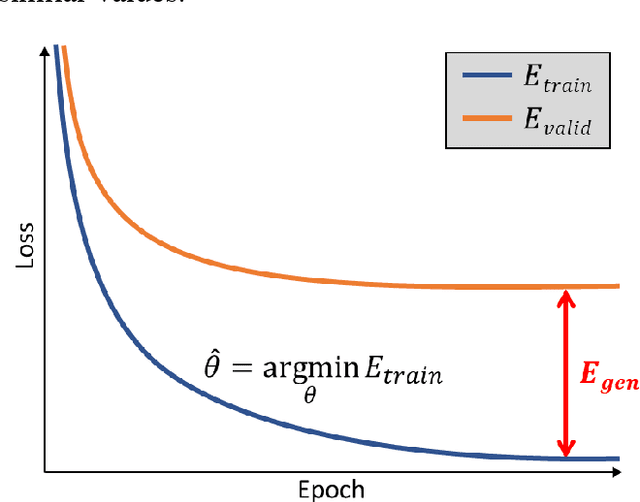
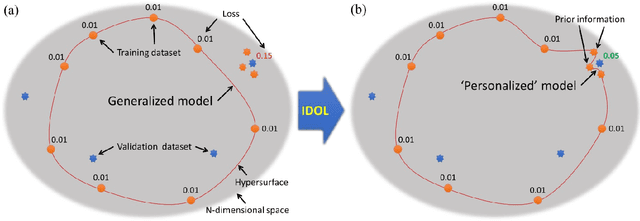
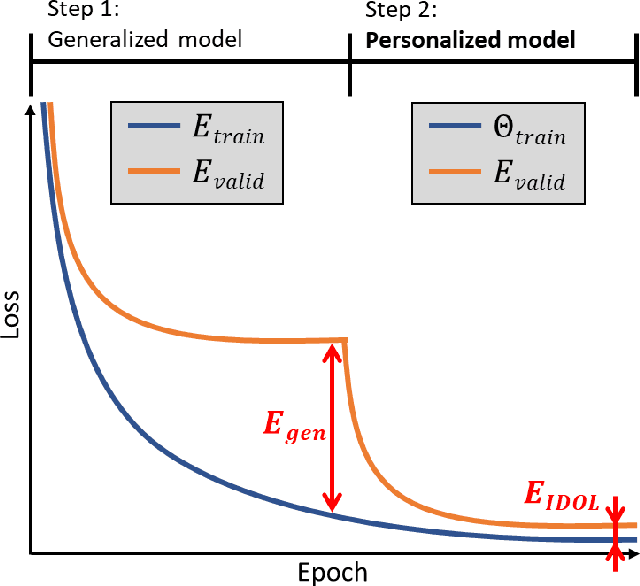
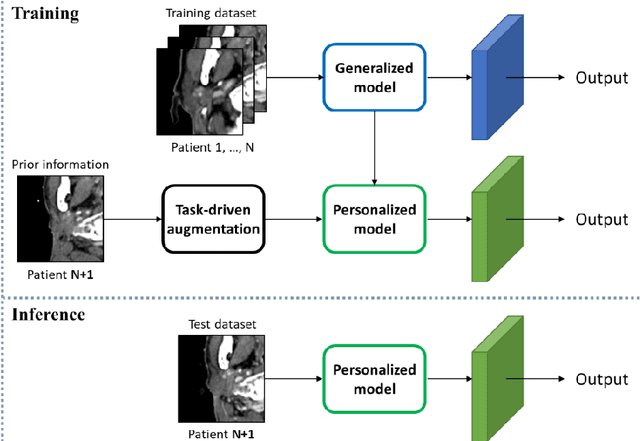
Abstract:In this study, we propose a tailored DL framework for patient-specific performance that leverages the behavior of a model intentionally overfitted to a patient-specific training dataset augmented from the prior information available in an ART workflow - an approach we term Intentional Deep Overfit Learning (IDOL). Implementing the IDOL framework in any task in radiotherapy consists of two training stages: 1) training a generalized model with a diverse training dataset of N patients, just as in the conventional DL approach, and 2) intentionally overfitting this general model to a small training dataset-specific the patient of interest (N+1) generated through perturbations and augmentations of the available task- and patient-specific prior information to establish a personalized IDOL model. The IDOL framework itself is task-agnostic and is thus widely applicable to many components of the ART workflow, three of which we use as a proof of concept here: the auto-contouring task on re-planning CTs for traditional ART, the MRI super-resolution (SR) task for MRI-guided ART, and the synthetic CT (sCT) reconstruction task for MRI-only ART. In the re-planning CT auto-contouring task, the accuracy measured by the Dice similarity coefficient improves from 0.847 with the general model to 0.935 by adopting the IDOL model. In the case of MRI SR, the mean absolute error (MAE) is improved by 40% using the IDOL framework over the conventional model. Finally, in the sCT reconstruction task, the MAE is reduced from 68 to 22 HU by utilizing the IDOL framework.
Deep learning-based COVID-19 pneumonia classification using chest CT images: model generalizability
Feb 18, 2021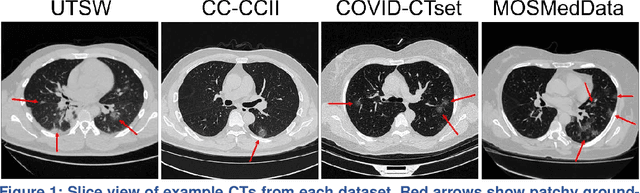
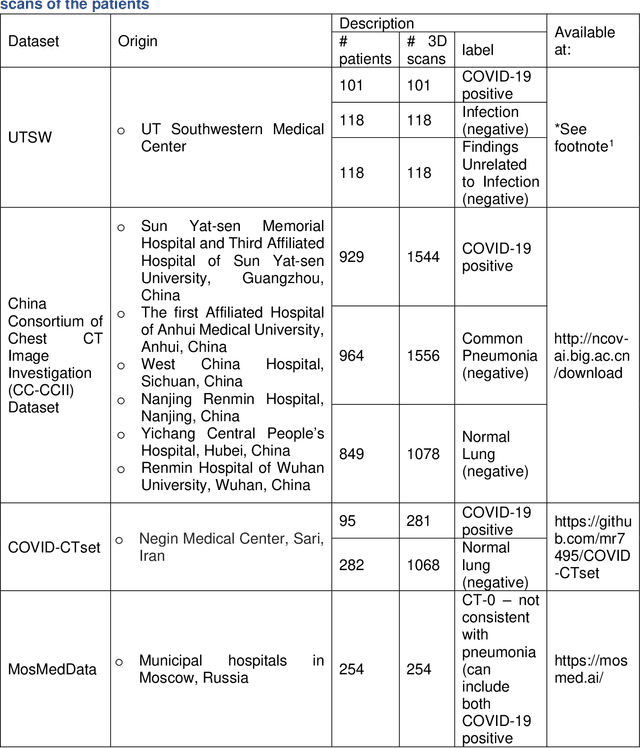
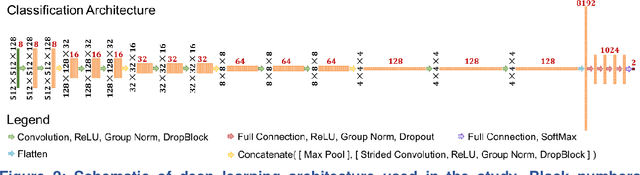
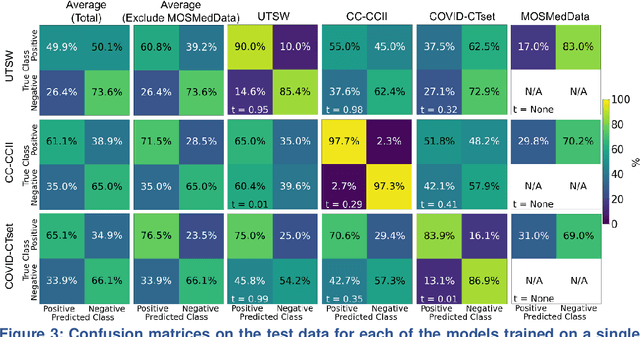
Abstract:Since the outbreak of the COVID-19 pandemic, worldwide research efforts have focused on using artificial intelligence (AI) technologies on various medical data of COVID-19-positive patients in order to identify or classify various aspects of the disease, with promising reported results. However, concerns have been raised over their generalizability, given the heterogeneous factors in training datasets. This study aims to examine the severity of this problem by evaluating deep learning (DL) classification models trained to identify COVID-19-positive patients on 3D computed tomography (CT) datasets from different countries. We collected one dataset at UT Southwestern (UTSW), and three external datasets from different countries: CC-CCII Dataset (China), COVID-CTset (Iran), and MosMedData (Russia). We divided the data into 2 classes: COVID-19-positive and COVID-19-negative patients. We trained nine identical DL-based classification models by using combinations of the datasets with a 72% train, 8% validation, and 20% test data split. The models trained on a single dataset achieved accuracy/area under the receiver operating characteristics curve (AUC) values of 0.87/0.826 (UTSW), 0.97/0.988 (CC-CCCI), and 0.86/0.873 (COVID-CTset) when evaluated on their own dataset. The models trained on multiple datasets and evaluated on a test set from one of the datasets used for training performed better. However, the performance dropped close to an AUC of 0.5 (random guess) for all models when evaluated on a different dataset outside of its training datasets. Including the MosMedData, which only contained positive labels, into the training did not necessarily help the performance on the other datasets. Multiple factors likely contribute to these results, such as patient demographics and differences in image acquisition or reconstruction, causing a data shift among different study cohorts.
PSA-Net: Deep Learning based Physician Style-Aware Segmentation Network for Post-Operative Prostate Cancer Clinical Target Volume
Feb 15, 2021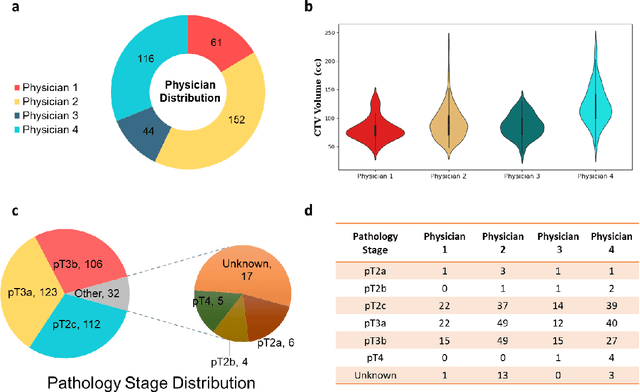
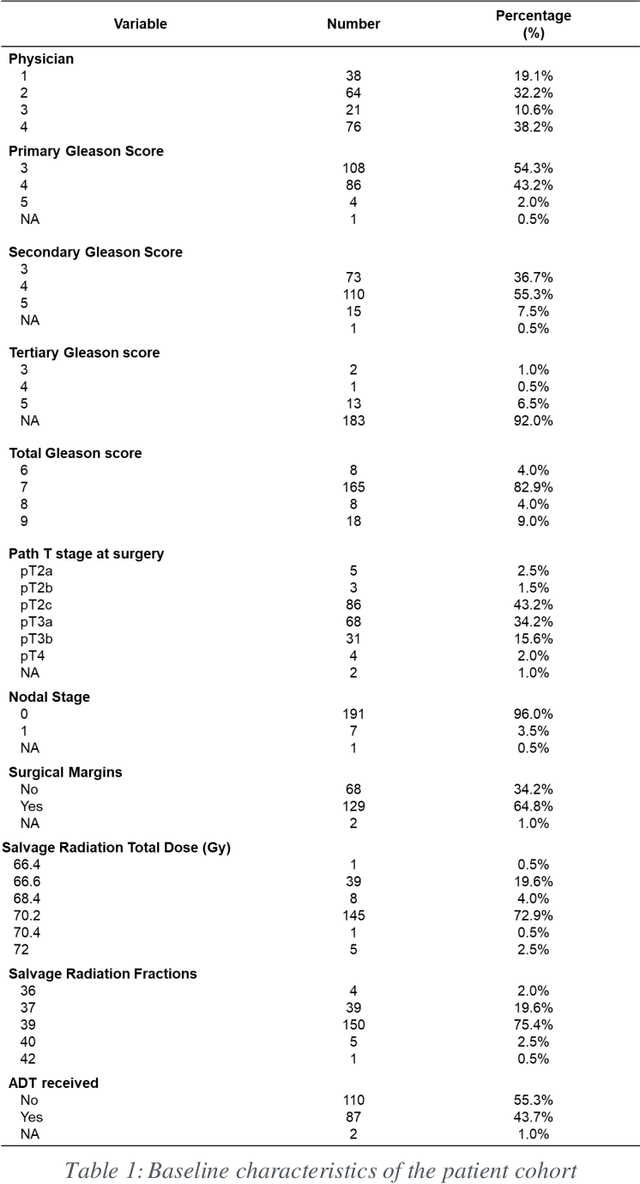
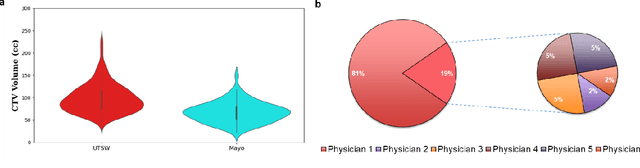
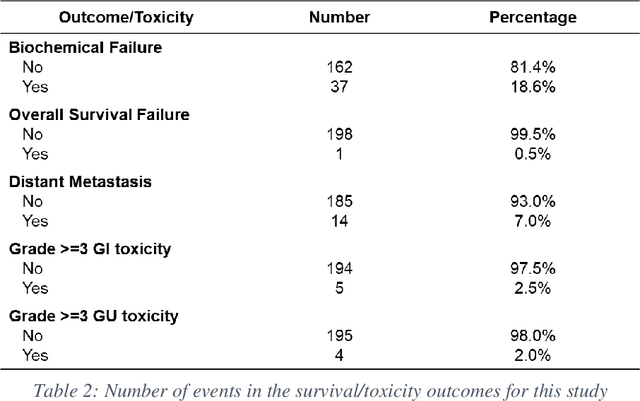
Abstract:Automatic segmentation of medical images with DL algorithms has proven to be highly successful. With most of these algorithms, inter-observer variation is an acknowledged problem, leading to sub-optimal results. This problem is even more significant in post-operative clinical target volume (post-op CTV) segmentation due to the absence of macroscopic visual tumor in the image. This study, using post-op CTV segmentation as the test bed, tries to determine if physician styles are consistent and learnable, if there is an impact of physician styles on treatment outcome and toxicity; and how to explicitly deal with physician styles in DL algorithms to facilitate its clinical acceptance. A classifier is trained to identify which physician has contoured the CTV from just the contour and corresponding CT scan, to determine if physician styles are consistent and learnable. Next, we evaluate if adapting automatic segmentation to physician styles would be clinically feasible based on a lack of difference between outcomes. For modeling different physician styles of CTV segmentation, a concept called physician style-aware (PSA) segmentation is proposed which is an encoder-multidecoder network trained with perceptual loss. With the proposed physician style-aware network (PSA-Net), Dice similarity coefficient (DSC) accuracy increases on an average of 3.4% for all physicians from a general model that is not style adapted. We show that stylistic contouring variations also exist between institutions that follow the same segmentation guidelines and show the effectiveness of the proposed method in adapting to new institutional styles. We observed an accuracy improvement of 5% in terms of DSC when adapting to the style of a separate institution.
Dosimetric impact of physician style variations in contouring CTV for post-operative prostate cancer: A deep learning based simulation study
Feb 01, 2021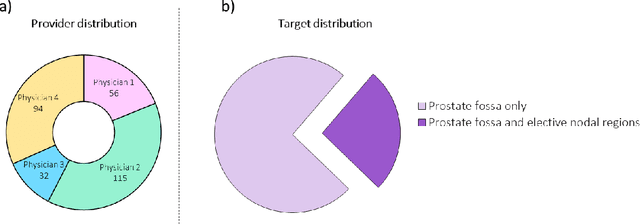
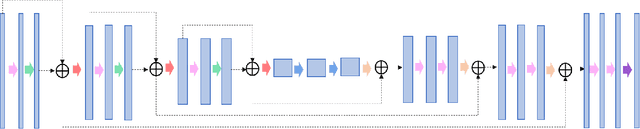
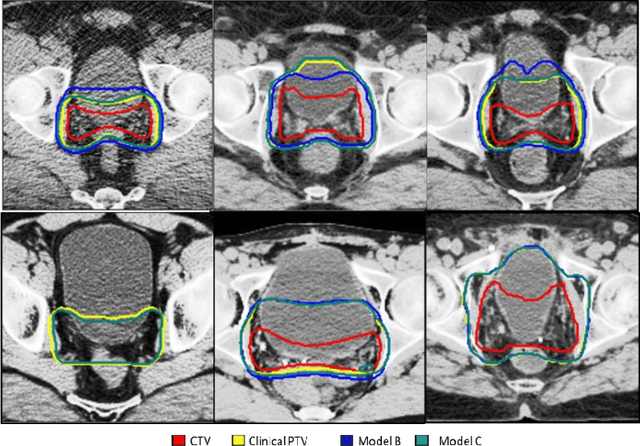
Abstract:In tumor segmentation, inter-observer variation is acknowledged to be a significant problem. This is even more significant in clinical target volume (CTV) segmentation, specifically, in post-operative settings, where a gross tumor does not exist. In this scenario, CTV is not an anatomically established structure but rather one determined by the physician based on the clinical guideline used, the preferred trade off between tumor control and toxicity, their experience, training background etc... This results in high inter-observer variability between physicians. Inter-observer variability has been considered an issue, however its dosimetric consequence is still unclear, due to the absence of multiple physician CTV contours for each patient and the significant amount of time required for dose planning. In this study, we analyze the impact that these physician stylistic variations have on organs-at-risk (OAR) dose by simulating the clinical workflow using deep learning. For a given patient previously treated by one physician, we use DL-based tools to simulate how other physicians would contour the CTV and how the corresponding dose distributions should look like for this patient. To simulate multiple physician styles, we use a previously developed in-house CTV segmentation model that can produce physician style-aware segmentations. The corresponding dose distribution is predicted using another in-house deep learning tool, which, averaging across all structures, is capable of predicting dose within 3% of the prescription dose on the test data. For every test patient, four different physician-style CTVs are considered and four different dose distributions are analyzed. OAR dose metrics are compared, showing that even though physician style variations results in organs getting different doses, all the important dose metrics except Maximum Dose point are within the clinically acceptable limit.
Deep High-Resolution Network for Low Dose X-ray CT Denoising
Feb 01, 2021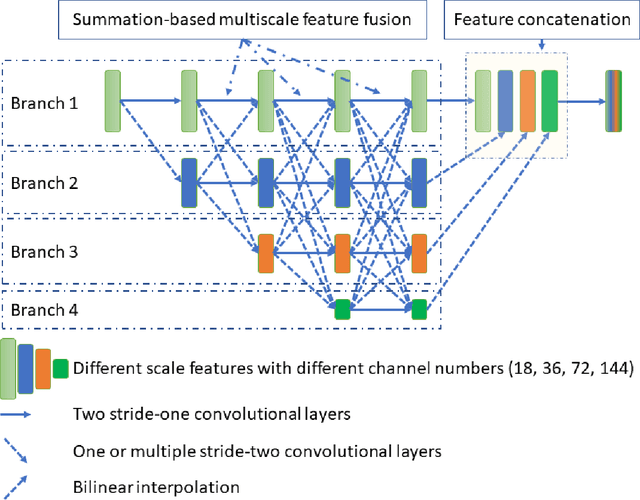
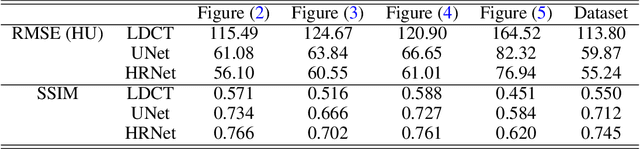
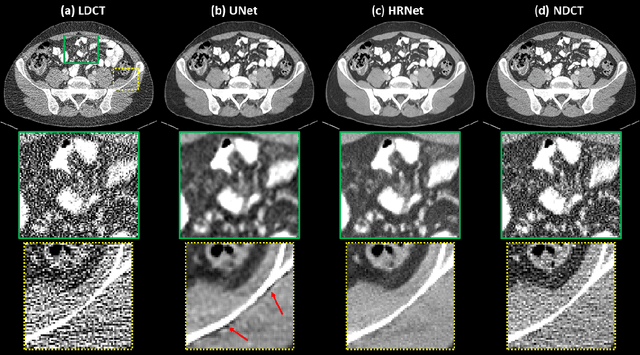
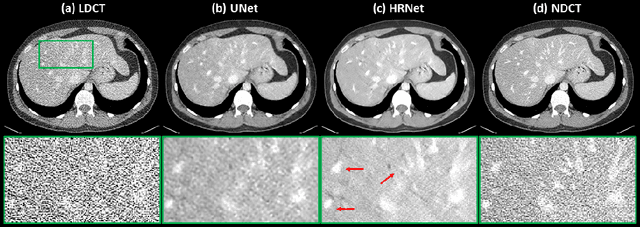
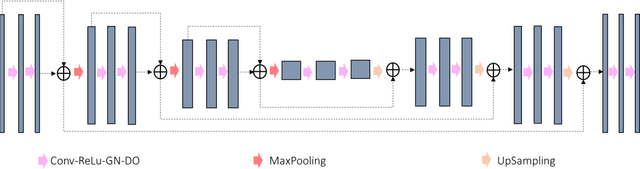
Abstract:Low Dose Computed Tomography (LDCT) is clinically desirable due to the reduced radiation to patients. However, the quality of LDCT images is often sub-optimal because of the inevitable strong quantum noise. Inspired by their unprecedent success in computer vision, deep learning (DL)-based techniques have been used for LDCT denoising. Despite the promising noise removal ability of DL models, people have observed that the resolution of the DL-denoised images is compromised, decreasing their clinical value. Aiming at relieving this problem, in this work, we developed a more effective denoiser by introducing a high-resolution network (HRNet). Since HRNet consists of multiple branches of subnetworks to extract multiscale features which are later fused together, the quality of the generated features can be substantially enhanced, leading to improved denoising performance. Experimental results demonstrated that the introduced HRNet-based denoiser outperforms the benchmarked UNet-based denoiser in terms of superior image resolution preservation ability while comparable, if not better, noise suppression ability. Quantitative metrics in terms of root-mean-squared-errors (RMSE)/structure similarity index (SSIM) showed that the HRNet-based denoiser can improve the values from 113.80/0.550 (LDCT) to 55.24/0.745 (HRNet), in comparison to 59.87/0.712 for the UNet-based denoiser.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge