Shweta Yadav
Identifying Depressive Symptoms from Tweets: Figurative Language Enabled Multitask Learning Framework
Nov 12, 2020
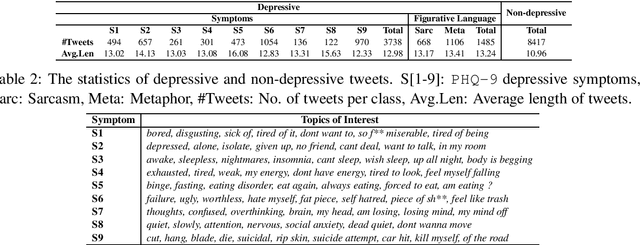
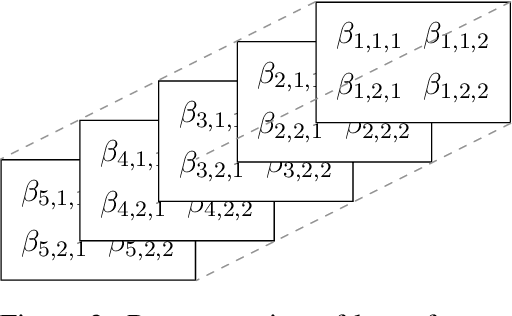
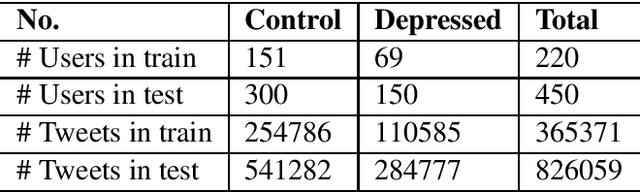
Abstract:Existing studies on using social media for deriving mental health status of users focus on the depression detection task. However, for case management and referral to psychiatrists, healthcare workers require practical and scalable depressive disorder screening and triage system. This study aims to design and evaluate a decision support system (DSS) to reliably determine the depressive triage level by capturing fine-grained depressive symptoms expressed in user tweets through the emulation of Patient Health Questionnaire-9 (PHQ-9) that is routinely used in clinical practice. The reliable detection of depressive symptoms from tweets is challenging because the 280-character limit on tweets incentivizes the use of creative artifacts in the utterances and figurative usage contributes to effective expression. We propose a novel BERT based robust multi-task learning framework to accurately identify the depressive symptoms using the auxiliary task of figurative usage detection. Specifically, our proposed novel task sharing mechanism, co-task aware attention, enables automatic selection of optimal information across the BERT layers and tasks by soft-sharing of parameters. Our results show that modeling figurative usage can demonstrably improve the model's robustness and reliability for distinguishing the depression symptoms.
Medical Knowledge-enriched Textual Entailment Framework
Nov 10, 2020
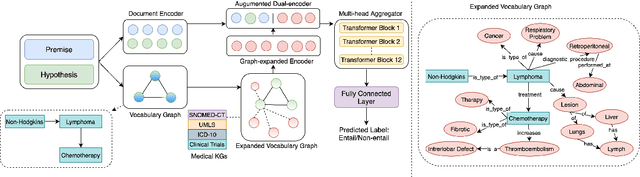
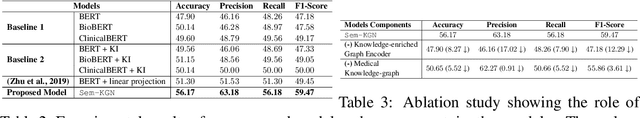
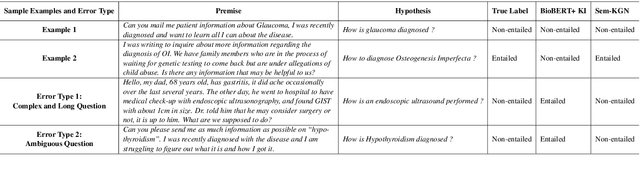
Abstract:One of the cardinal tasks in achieving robust medical question answering systems is textual entailment. The existing approaches make use of an ensemble of pre-trained language models or data augmentation, often to clock higher numbers on the validation metrics. However, two major shortcomings impede higher success in identifying entailment: (1) understanding the focus/intent of the question and (2) ability to utilize the real-world background knowledge to capture the context beyond the sentence. In this paper, we present a novel Medical Knowledge-Enriched Textual Entailment framework that allows the model to acquire a semantic and global representation of the input medical text with the help of a relevant domain-specific knowledge graph. We evaluate our framework on the benchmark MEDIQA-RQE dataset and manifest that the use of knowledge enriched dual-encoding mechanism help in achieving an absolute improvement of 8.27% over SOTA language models. We have made the source code available here.
"When they say weed causes depression, but it's your fav antidepressant": Knowledge-aware Attention Framework for Relationship Extraction
Sep 21, 2020
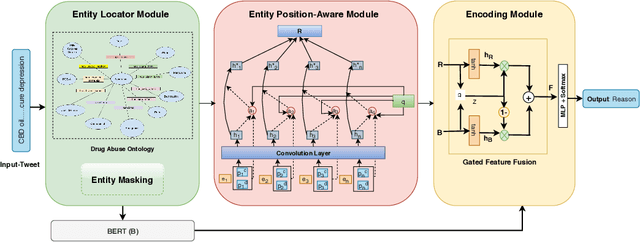

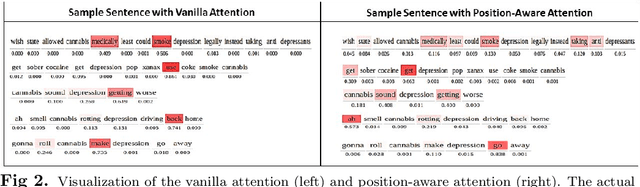
Abstract:With the increasing legalization of medical and recreational use of cannabis, more research is needed to understand the association between depression and consumer behavior related to cannabis consumption. Big social media data has potential to provide deeper insights about these associations to public health analysts. In this interdisciplinary study, we demonstrate the value of incorporating domain-specific knowledge in the learning process to identify the relationships between cannabis use and depression. We develop an end-to-end knowledge infused deep learning framework (Gated-K-BERT) that leverages the pre-trained BERT language representation model and domain-specific declarative knowledge source (Drug Abuse Ontology (DAO)) to jointly extract entities and their relationship using gated fusion sharing mechanism. Our model is further tailored to provide more focus to the entities mention in the sentence through entity-position aware attention layer, where ontology is used to locate the target entities position. Experimental results show that inclusion of the knowledge-aware attentive representation in association with BERT can extract the cannabis-depression relationship with better coverage in comparison to the state-of-the-art relation extractor.
Assessing the Severity of Health States based on Social Media Posts
Sep 21, 2020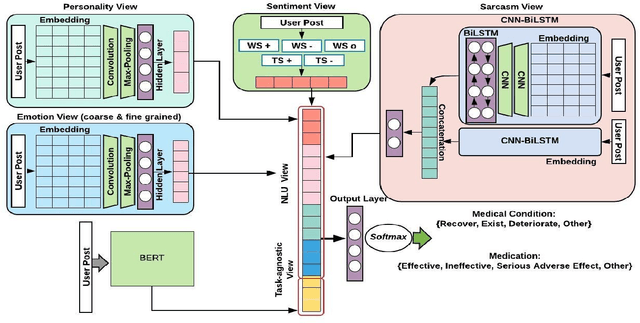
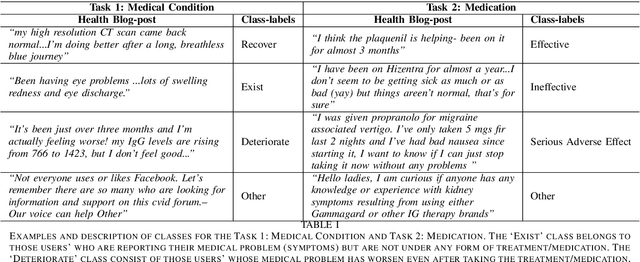
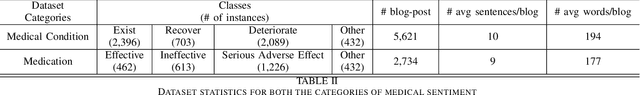
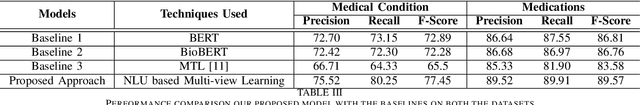
Abstract:The unprecedented growth of Internet users has resulted in an abundance of unstructured information on social media including health forums, where patients request health-related information or opinions from other users. Previous studies have shown that online peer support has limited effectiveness without expert intervention. Therefore, a system capable of assessing the severity of health state from the patients' social media posts can help health professionals (HP) in prioritizing the user's post. In this study, we inspect the efficacy of different aspects of Natural Language Understanding (NLU) to identify the severity of the user's health state in relation to two perspectives(tasks) (a) Medical Condition (i.e., Recover, Exist, Deteriorate, Other) and (b) Medication (i.e., Effective, Ineffective, Serious Adverse Effect, Other) in online health communities. We propose a multiview learning framework that models both the textual content as well as contextual-information to assess the severity of the user's health state. Specifically, our model utilizes the NLU views such as sentiment, emotions, personality, and use of figurative language to extract the contextual information. The diverse NLU views demonstrate its effectiveness on both the tasks and as well as on the individual disease to assess a user's health.
Relation Extraction from Biomedical and Clinical Text: Unified Multitask Learning Framework
Sep 20, 2020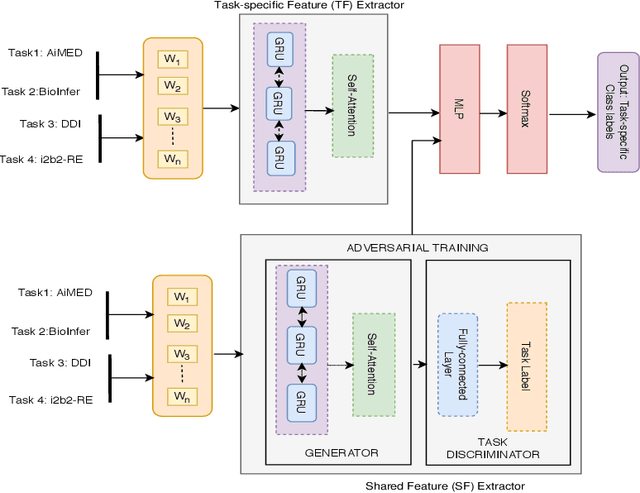

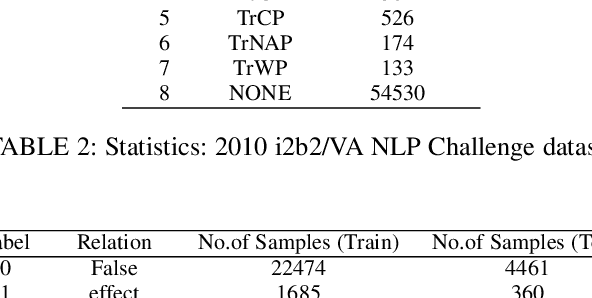
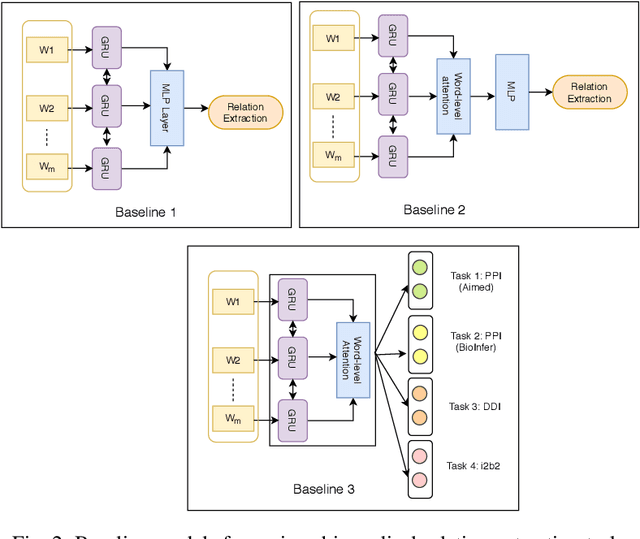
Abstract:To minimize the accelerating amount of time invested in the biomedical literature search, numerous approaches for automated knowledge extraction have been proposed. Relation extraction is one such task where semantic relations between the entities are identified from the free text. In the biomedical domain, extraction of regulatory pathways, metabolic processes, adverse drug reaction or disease models necessitates knowledge from the individual relations, for example, physical or regulatory interactions between genes, proteins, drugs, chemical, disease or phenotype. In this paper, we study the relation extraction task from three major biomedical and clinical tasks, namely drug-drug interaction, protein-protein interaction, and medical concept relation extraction. Towards this, we model the relation extraction problem in multi-task learning (MTL) framework and introduce for the first time the concept of structured self-attentive network complemented with the adversarial learning approach for the prediction of relationships from the biomedical and clinical text. The fundamental notion of MTL is to simultaneously learn multiple problems together by utilizing the concepts of the shared representation. Additionally, we also generate the highly efficient single task model which exploits the shortest dependency path embedding learned over the attentive gated recurrent unit to compare our proposed MTL models. The framework we propose significantly improves overall the baselines (deep learning techniques) and single-task models for predicting the relationships, without compromising on the performance of all the tasks.
Relation extraction between the clinical entities based on the shortest dependency path based LSTM
Mar 24, 2019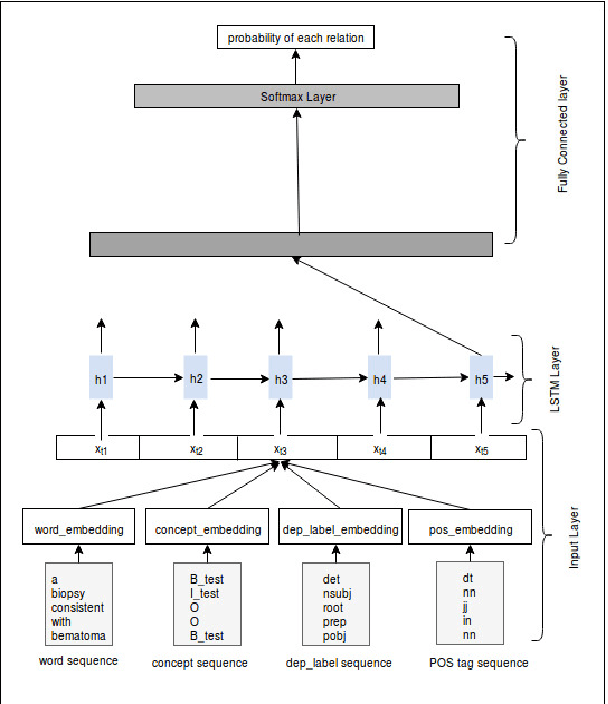
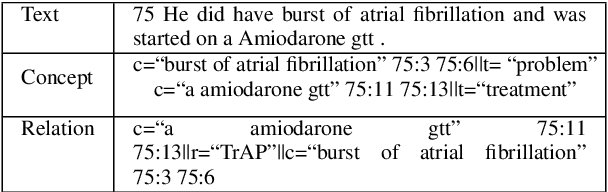

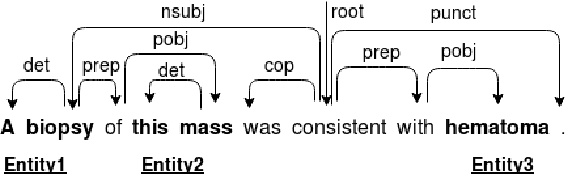
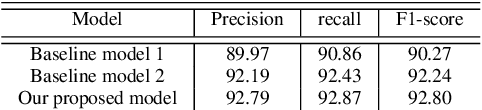
Abstract:Owing to the exponential rise in the electronic medical records, information extraction in this domain is becoming an important area of research in recent years. Relation extraction between the medical concepts such as medical problem, treatment, and test etc. is also one of the most important tasks in this area. In this paper, we present an efficient relation extraction system based on the shortest dependency path (SDP) generated from the dependency parsed tree of the sentence. Instead of relying on many handcrafted features and the whole sequence of tokens present in a sentence, our system relies only on the SDP between the target entities. For every pair of entities, the system takes only the words in the SDP, their dependency labels, Part-of-Speech information and the types of the entities as the input. We develop a dependency parser for extracting dependency information. We perform our experiments on the benchmark i2b2 dataset for clinical relation extraction challenge 2010. Experimental results show that our system outperforms the existing systems.
Leveraging Medical Sentiment to Understand Patients Health on Social Media
Jul 30, 2018
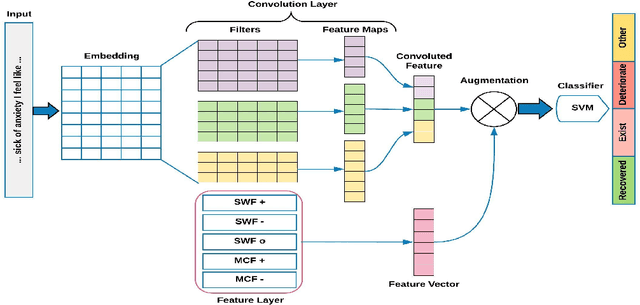

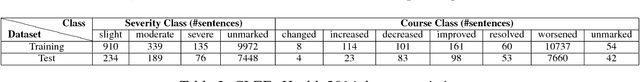
Abstract:The unprecedented growth of Internet users in recent years has resulted in an abundance of unstructured information in the form of social media text. A large percentage of this population is actively engaged in health social networks to share health-related information. In this paper, we address an important and timely topic by analyzing the users' sentiments and emotions w.r.t their medical conditions. Towards this, we examine users on popular medical forums (Patient.info,dailystrength.org), where they post on important topics such as asthma, allergy, depression, and anxiety. First, we provide a benchmark setup for the task by crawling the data, and further define the sentiment specific fine-grained medical conditions (Recovered, Exist, Deteriorate, and Other). We propose an effective architecture that uses a Convolutional Neural Network (CNN) as a data-driven feature extractor and a Support Vector Machine (SVM) as a classifier. We further develop a sentiment feature which is sensitive to the medical context. Here, we show that the use of medical sentiment feature along with extracted features from CNN improves the model performance. In addition to our dataset, we also evaluate our approach on the benchmark "CLEF eHealth 2014" corpora and show that our model outperforms the state-of-the-art techniques.
Feature Assisted bi-directional LSTM Model for Protein-Protein Interaction Identification from Biomedical Texts
Jul 05, 2018
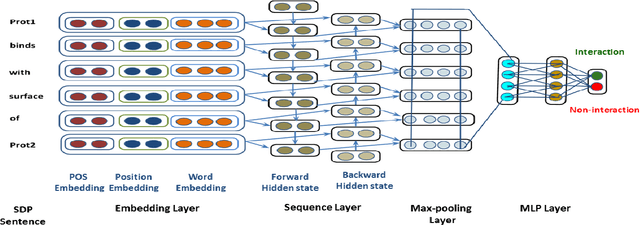
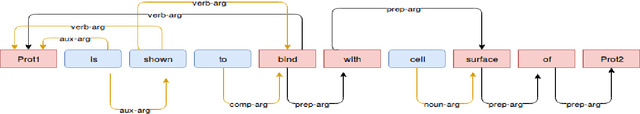
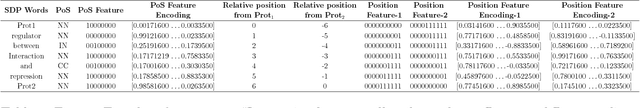
Abstract:Knowledge about protein-protein interactions is essential in understanding the biological processes such as metabolic pathways, DNA replication, and transcription etc. However, a majority of the existing Protein-Protein Interaction (PPI) systems are dependent primarily on the scientific literature, which is yet not accessible as a structured database. Thus, efficient information extraction systems are required for identifying PPI information from the large collection of biomedical texts. Most of the existing systems model the PPI extraction task as a classification problem and are tailored to the handcrafted feature set including domain dependent features. In this paper, we present a novel method based on deep bidirectional long short-term memory (B-LSTM) technique that exploits word sequences and dependency path related information to identify PPI information from text. This model leverages joint modeling of proteins and relations in a single unified framework, which we name as Shortest Dependency Path B-LSTM (sdpLSTM) model. We perform experiments on two popular benchmark PPI datasets, namely AiMed & BioInfer. The evaluation shows the F1-score values of 86.45% and 77.35% on AiMed and BioInfer, respectively. Comparisons with the existing systems show that our proposed approach attains state-of-the-art performance.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge