Shizuo Kaji
Topological filtering of a signal over a network
Aug 26, 2024Abstract:Graph Signal Processing deals with the problem of analyzing and processing signals defined on graphs. In this paper, we introduce a novel filtering method for graph-based signals by employing ideas from topological data analysis. We begin by working with signals over general graphs and then extend our approach to what we term signals over graphs with faces. To construct the filter, we introduce a new structure called the Basin Hierarchy Tree, which encodes the persistent homology. We provide an efficient algorithm and demonstrate the effectiveness of our approach through examples with synthetic and real datasets. This work bridges topological data analysis and signal processing, presenting a new application of persistent homology as a topological data processing tool.
Iterative CT Reconstruction via Latent Variable Optimization of Shallow Diffusion Models
Aug 06, 2024Abstract:Image generative AI has garnered significant attention in recent years. In particular, the diffusion model, a core component of recent generative AI, produces high-quality images with rich diversity. In this study, we propose a novel CT reconstruction method by combining the denoising diffusion probabilistic model with iterative CT reconstruction. In sharp contrast to previous studies, we optimize the fidelity loss of CT reconstruction with respect to the latent variable of the diffusion model, instead of the image and model parameters. To suppress anatomical structure changes produced by the diffusion model, we shallow the diffusion and reverse processes, and fix a set of added noises in the reverse process to make it deterministic during inference. We demonstrate the effectiveness of the proposed method through sparse view CT reconstruction of 1/10 view projection data. Despite the simplicity of the implementation, the proposed method shows the capability of reconstructing high-quality images while preserving the patient's anatomical structure, and outperforms existing methods including iterative reconstruction, iterative reconstruction with total variation, and the diffusion model alone in terms of quantitative indices such as SSIM and PSNR. We also explore further sparse view CT using 1/20 view projection data with the same trained diffusion model. As the number of iterations increases, image quality improvement comparable to that of 1/10 sparse view CT reconstruction is achieved. In principle, the proposed method can be widely applied not only to CT but also to other imaging modalities such as MRI, PET, and SPECT.
An explicit construction of Kaleidocycles
Aug 09, 2023Abstract:We model a family of closed kinematic chains, known as Kaleidocycles, with the theory of discrete spatial curves. By leveraging the connection between the deformation of discrete curves and the semi-discrete integrable systems, we describe the motion of a Kaleidocycle by elliptic theta functions. This study showcases an interesting example in which an integrable system generates an orbit in the space of the real solutions of polynomial equations defined by geometric constraints.
Training deep cross-modality conversion models with a small amount of data and its application to MVCT to kVCT conversion
Jul 12, 2021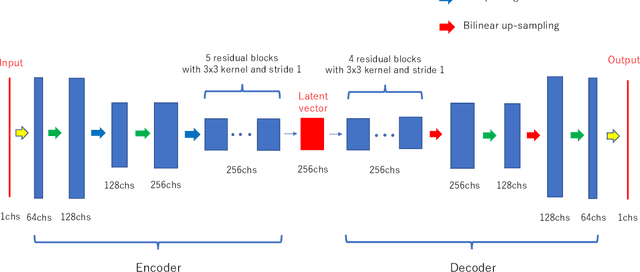
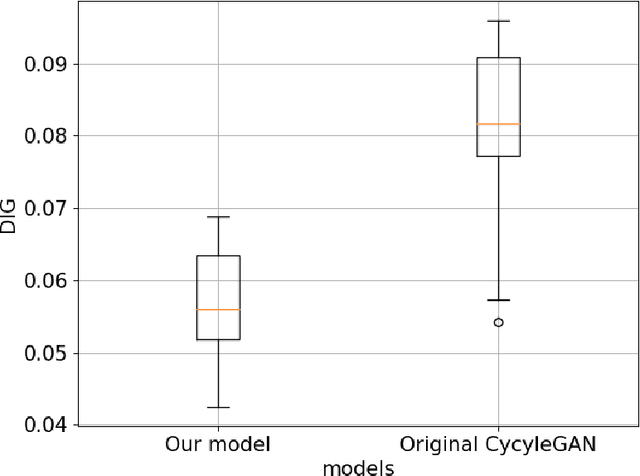
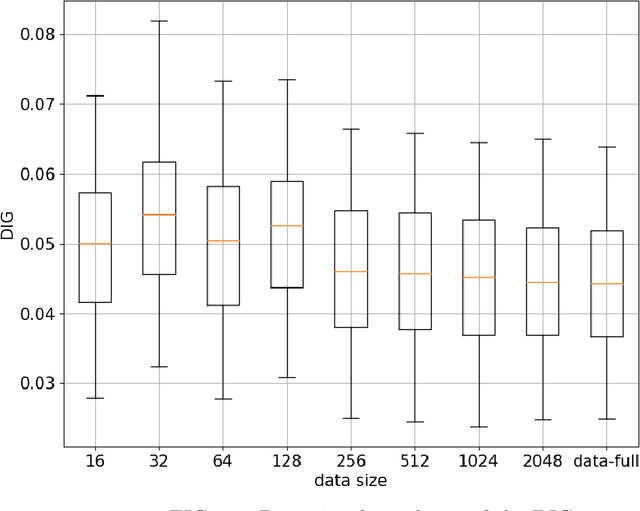
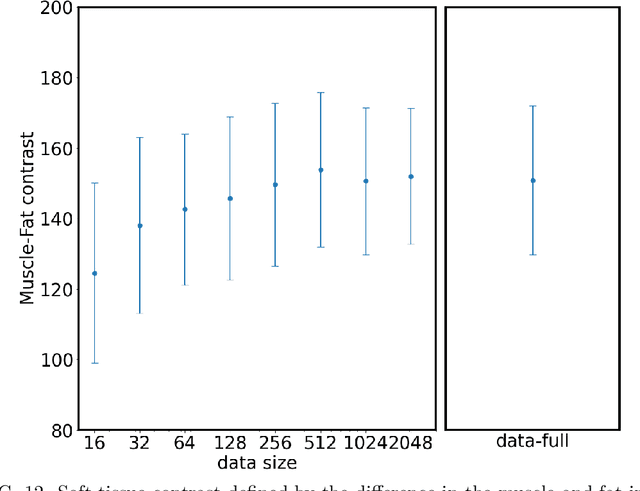
Abstract:Deep-learning-based image processing has emerged as a valuable tool in recent years owing to its high performance. However, the quality of deep-learning-based methods relies heavily on the amount of training data, and the cost of acquiring a large amount of data is often prohibitive in medical fields. Therefore, we performed CT modality conversion based on deep learning requiring only a small number of unsupervised images. The proposed method is based on generative adversarial networks (GANs) with several extensions tailored for CT images. This method emphasizes the preservation of the structure in the processed images and reduction in the amount of training data. This method was applied to realize the conversion of mega-voltage computed tomography (MVCT) to kilo-voltage computed tomography (kVCT) images. Training was performed using several datasets acquired from patients with head and neck cancer. The size of the datasets ranged from 16 slices (for two patients) to 2745 slices (for 137 patients) of MVCT and 2824 slices of kVCT for 98 patients. The quality of the processed MVCT images was considerably enhanced, and the structural changes in the images were minimized. With an increase in the size of training data, the image quality exhibited a satisfactory convergence from a few hundred slices. In addition to statistical and visual evaluations, these results were clinically evaluated by medical doctors in terms of the accuracy of contouring. We developed an MVCT to kVCT conversion model based on deep learning, which can be trained using a few hundred unpaired images. The stability of the model against the change in the data size was demonstrated. This research promotes the reliable use of deep learning in clinical medicine by partially answering the commonly asked questions: "Is our data enough? How much data must we prepare?"
Nested Subspace Arrangement for Representation of Relational Data
Jul 04, 2020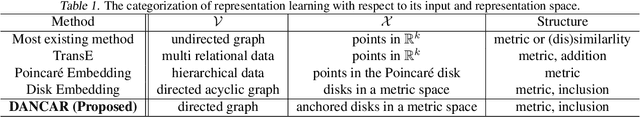
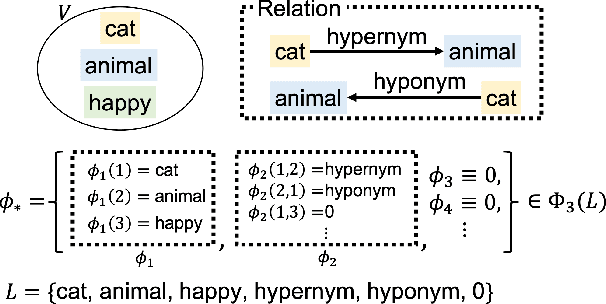
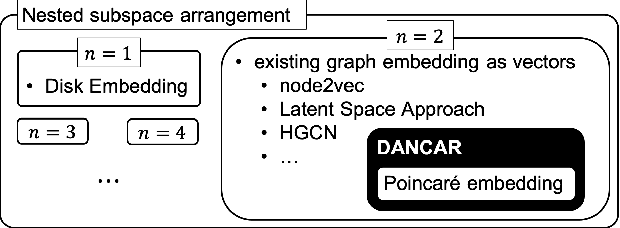
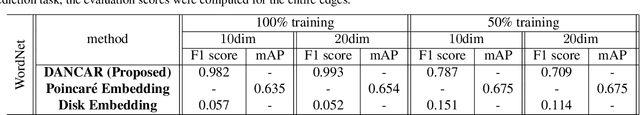
Abstract:Studies on acquiring appropriate continuous representations of discrete objects, such as graphs and knowledge base data, have been conducted by many researchers in the field of machine learning. In this study, we introduce Nested SubSpace (NSS) arrangement, a comprehensive framework for representation learning. We show that existing embedding techniques can be regarded as special cases of the NSS arrangement. Based on the concept of the NSS arrangement, we implement a Disk-ANChor ARrangement (DANCAR), a representation learning method specialized to reproducing general graphs. Numerical experiments have shown that DANCAR has successfully embedded WordNet in ${\mathbb R}^{20}$ with an F1 score of 0.993 in the reconstruction task. DANCAR is also suitable for visualization in understanding the characteristics of graphs.
Cubical Ripser: Software for computing persistent homology of image and volume data
Jun 12, 2020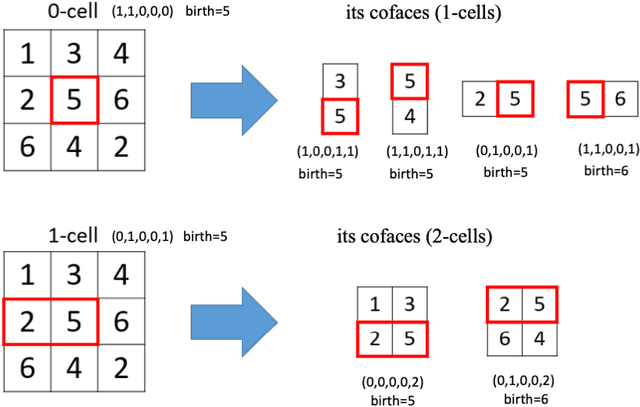
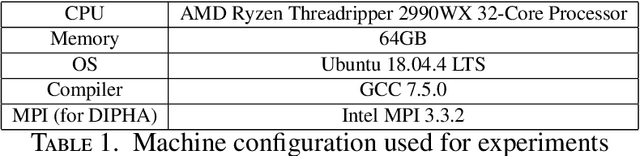

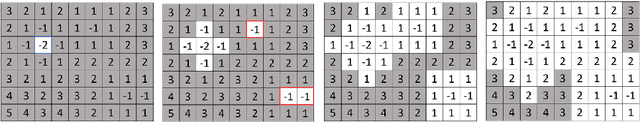
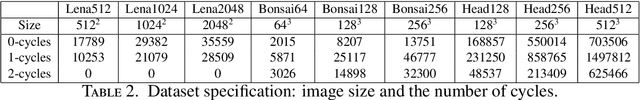
Abstract:We introduce Cubical Ripser for computing persistent homology of image and volume data (more precisely, weighted cubical complexes). To our best knowledge, Cubical Ripser is currently the fastest and the most memory-efficient program for computing persistent homology of weighted cubical complexes. We demonstrate our software with an example of image analysis in which persistent homology and convolutional neural networks are successfully combined. Our open-source implementation is available online.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge