Shalmali Joshi
Ethical Machine Learning in Health Care
Oct 08, 2020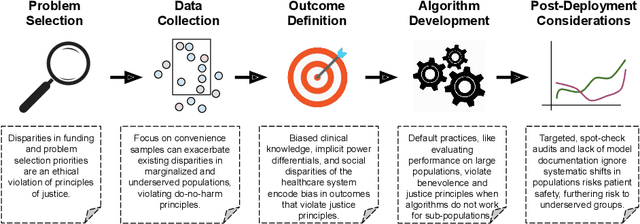
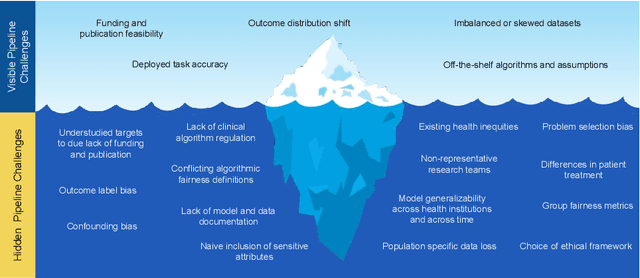
Abstract:The use of machine learning (ML) in health care raises numerous ethical concerns, especially as models can amplify existing health inequities. Here, we outline ethical considerations for equitable ML in the advancement of health care. Specifically, we frame ethics of ML in health care through the lens of social justice. We describe ongoing efforts and outline challenges in a proposed pipeline of ethical ML in health, ranging from problem selection to post-deployment considerations. We close by summarizing recommendations to address these challenges.
Probabilistic Machine Learning for Healthcare
Sep 23, 2020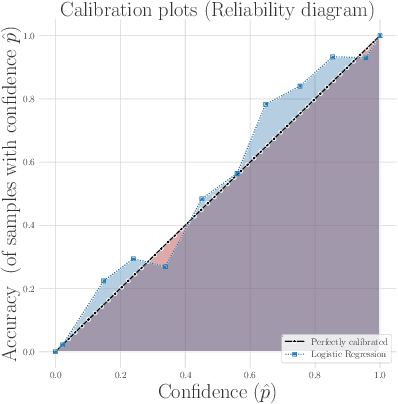
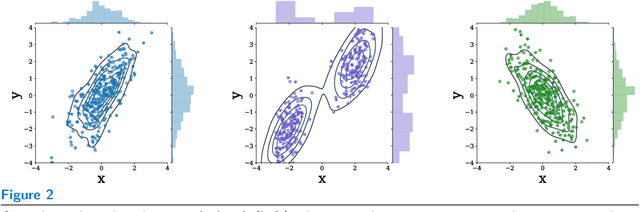
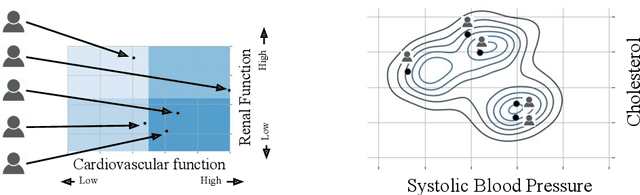
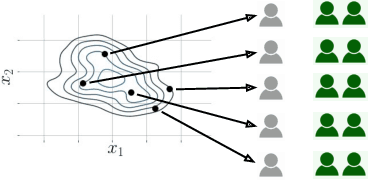
Abstract:Machine learning can be used to make sense of healthcare data. Probabilistic machine learning models help provide a complete picture of observed data in healthcare. In this review, we examine how probabilistic machine learning can advance healthcare. We consider challenges in the predictive model building pipeline where probabilistic models can be beneficial including calibration and missing data. Beyond predictive models, we also investigate the utility of probabilistic machine learning models in phenotyping, in generative models for clinical use cases, and in reinforcement learning.
Sequential Explanations with Mental Model-Based Policies
Jul 17, 2020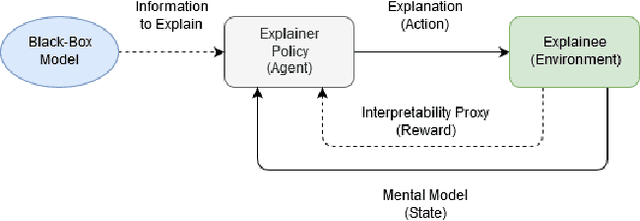

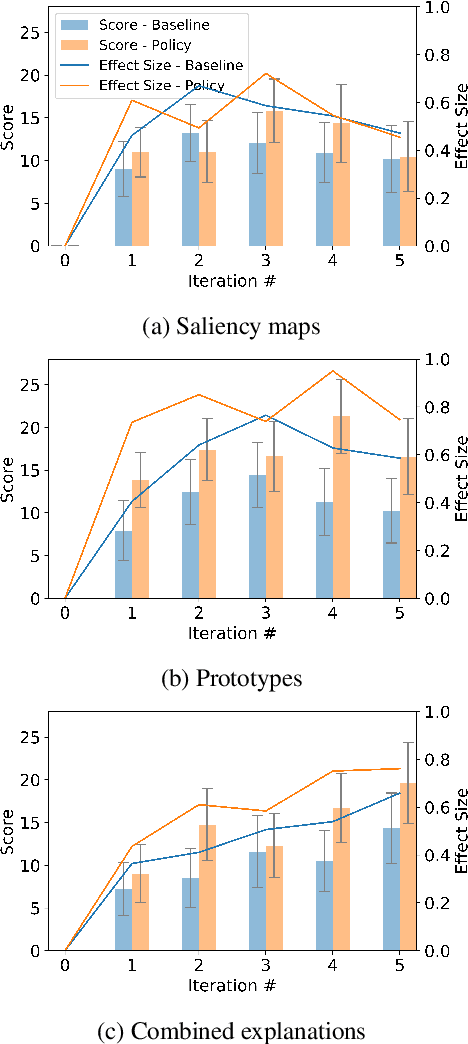
Abstract:The act of explaining across two parties is a feedback loop, where one provides information on what needs to be explained and the other provides an explanation relevant to this information. We apply a reinforcement learning framework which emulates this format by providing explanations based on the explainee's current mental model. We conduct novel online human experiments where explanations generated by various explanation methods are selected and presented to participants, using policies which observe participants' mental models, in order to optimize an interpretability proxy. Our results suggest that mental model-based policies (anchored in our proposed state representation) may increase interpretability over multiple sequential explanations, when compared to a random selection baseline. This work provides insight into how to select explanations which increase relevant information for users, and into conducting human-grounded experimentation to understand interpretability.
Counterfactually Guided Policy Transfer in Clinical Settings
Jun 20, 2020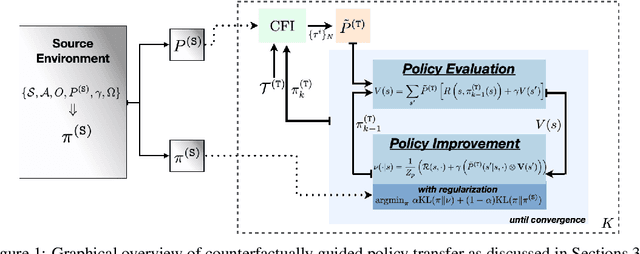
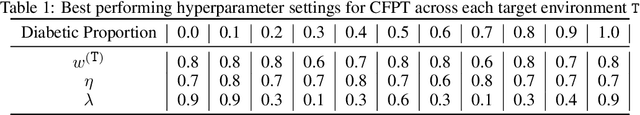
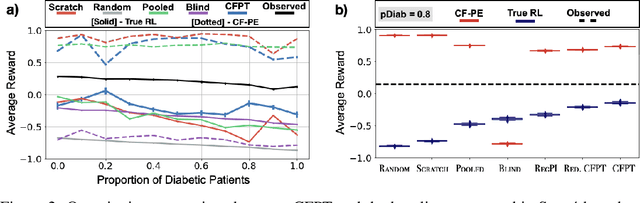
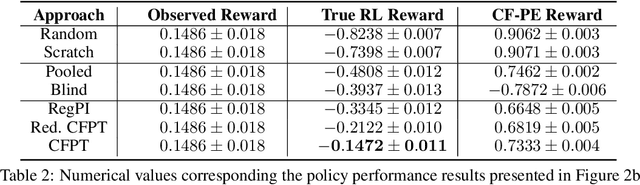
Abstract:Reliably transferring treatment policies learned in one clinical environment to another is currently limited by challenges related to domain shift. In this paper we address off-policy learning for sequential decision making under domain shift -- a scenario susceptible to catastrophic overconfidence -- which is highly relevant to a high-stakes clinical settings where the target domain may also be data-scarce. We propose a two-fold counterfactual regularization procedure to improve off-policy learning, addressing domain shift and data scarcity. First, we utilize an informative prior derived from a data-rich source environment to indirectly improve drawing counterfactual example observations. Then, these samples are then used to learn a policy for the target domain, regularized by the source policy through KL-divergence. In simulated sepsis treatment, our counterfactual policy transfer procedure significantly improves the performance of a learned treatment policy.
What went wrong and when? Instance-wise Feature Importance for Time-series Models
Mar 05, 2020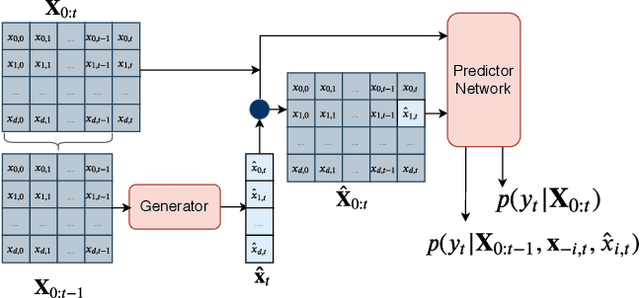
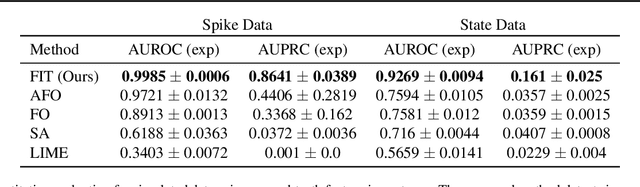
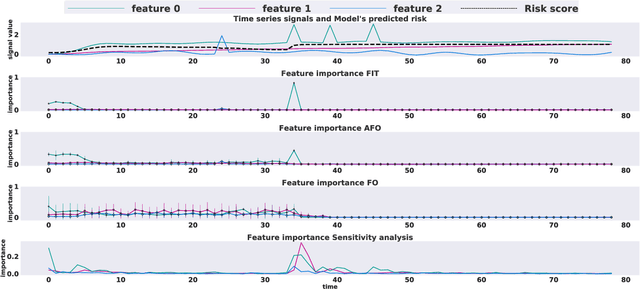
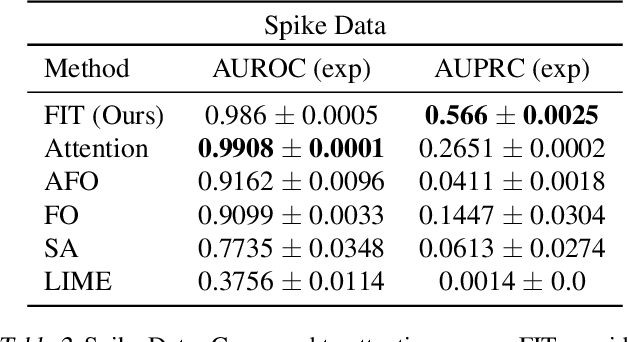
Abstract:Multivariate time series models are poised to be used for decision support in high-stakes applications, such as healthcare. In these contexts, it is important to know which features at which times most influenced a prediction. We demonstrate a general approach for assigning importance to observations in multivariate time series, based on their counterfactual influence on future predictions. Specifically, we define the importance of an observation as the change in the predictive distribution, had the observation not been seen. We integrate over plausible counterfactuals by sampling from the corresponding conditional distributions of generative time series models. We compare our importance metric to gradient-based explanations, attention mechanisms, and other baselines in simulated and clinical ICU data, and show that our approach generates the most precise explanations. Our method is inexpensive, model agnostic, and can be used with arbitrarily complex time series models and predictors.
Towards Realistic Individual Recourse and Actionable Explanations in Black-Box Decision Making Systems
Jul 22, 2019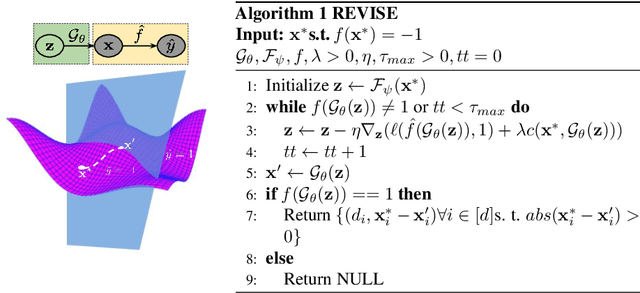
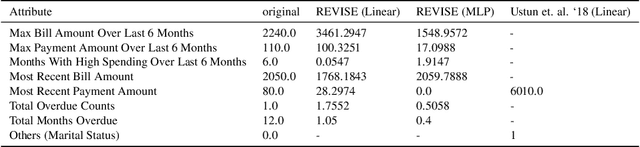
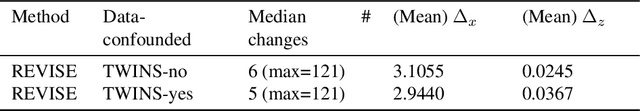
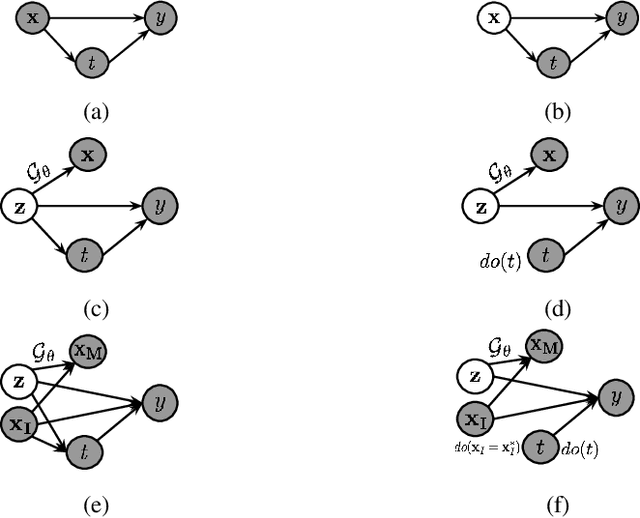
Abstract:Machine learning based decision making systems are increasingly affecting humans. An individual can suffer an undesirable outcome under such decision making systems (e.g. denied credit) irrespective of whether the decision is fair or accurate. Individual recourse pertains to the problem of providing an actionable set of changes a person can undertake in order to improve their outcome. We propose a recourse algorithm that models the underlying data distribution or manifold. We then provide a mechanism to generate the smallest set of changes that will improve an individual's outcome. This mechanism can be easily used to provide recourse for any differentiable machine learning based decision making system. Further, the resulting algorithm is shown to be applicable to both supervised classification and causal decision making systems. Our work attempts to fill gaps in existing fairness literature that have primarily focused on discovering and/or algorithmically enforcing fairness constraints on decision making systems. This work also provides an alternative approach to generating counterfactual explanations.
What Clinicians Want: Contextualizing Explainable Machine Learning for Clinical End Use
May 13, 2019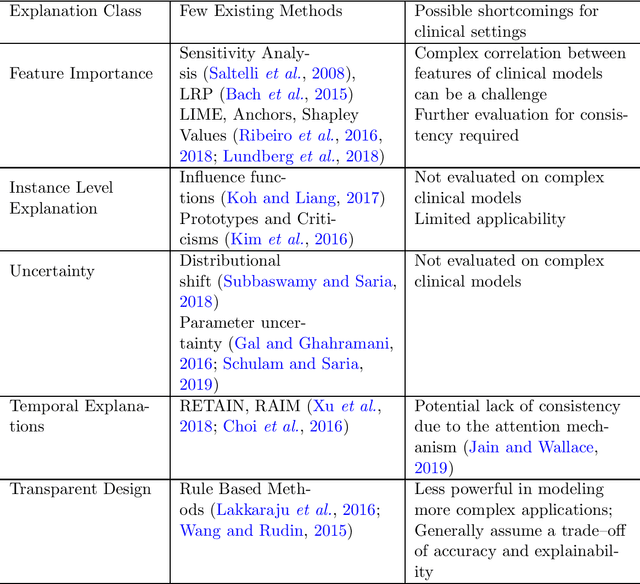
Abstract:Translating machine learning (ML) models effectively to clinical practice requires establishing clinicians' trust. Explainability, or the ability of an ML model to justify its outcomes and assist clinicians in rationalizing the model prediction, has been generally understood to be critical to establishing trust. However, the field suffers from the lack of concrete definitions for usable explanations in different settings. To identify specific aspects of explainability that may catalyze building trust in ML models, we surveyed clinicians from two distinct acute care specialties (Intenstive Care Unit and Emergency Department). We use their feedback to characterize when explainability helps to improve clinicians' trust in ML models. We further identify the classes of explanations that clinicians identified as most relevant and crucial for effective translation to clinical practice. Finally, we discern concrete metrics for rigorous evaluation of clinical explainability methods. By integrating perceptions of explainability between clinicians and ML researchers we hope to facilitate the endorsement and broader adoption and sustained use of ML systems in healthcare.
xGEMs: Generating Examplars to Explain Black-Box Models
Jun 22, 2018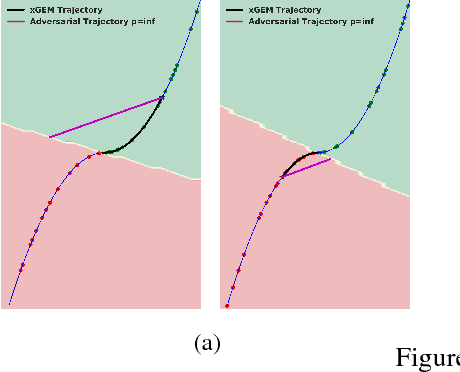
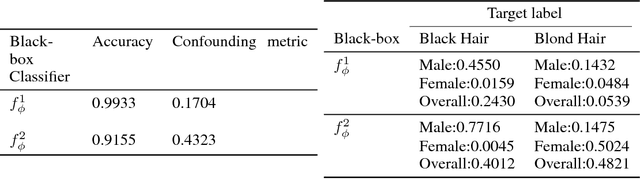
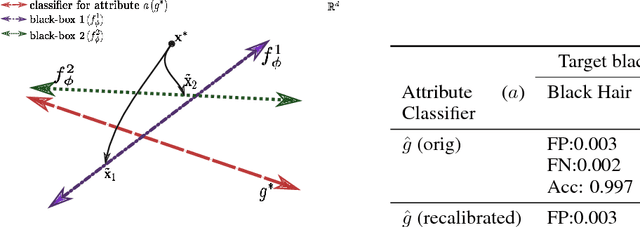

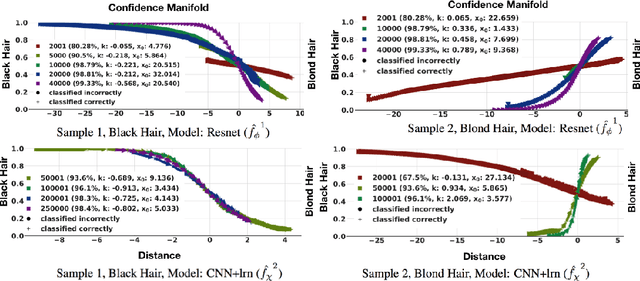
Abstract:This work proposes xGEMs or manifold guided exemplars, a framework to understand black-box classifier behavior by exploring the landscape of the underlying data manifold as data points cross decision boundaries. To do so, we train an unsupervised implicit generative model -- treated as a proxy to the data manifold. We summarize black-box model behavior quantitatively by perturbing data samples along the manifold. We demonstrate xGEMs' ability to detect and quantify bias in model learning and also for understanding the changes in model behavior as training progresses.
Identifiable Phenotyping using Constrained Non-Negative Matrix Factorization
Sep 20, 2016
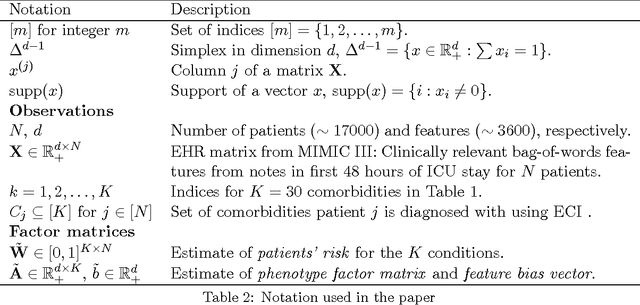
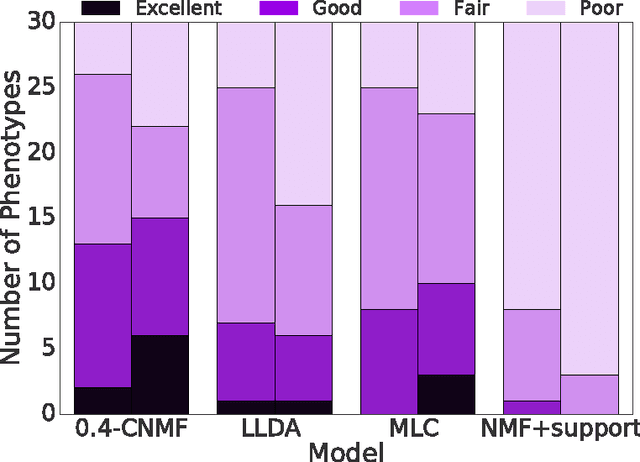

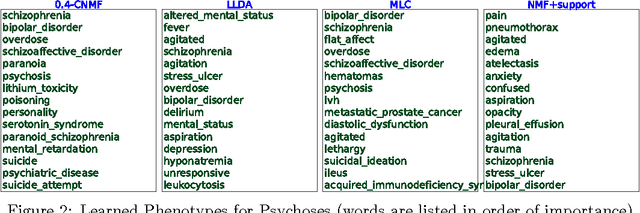
Abstract:This work proposes a new algorithm for automated and simultaneous phenotyping of multiple co-occurring medical conditions, also referred as comorbidities, using clinical notes from the electronic health records (EHRs). A basic latent factor estimation technique of non-negative matrix factorization (NMF) is augmented with domain specific constraints to obtain sparse latent factors that are anchored to a fixed set of chronic conditions. The proposed anchoring mechanism ensures a one-to-one identifiable and interpretable mapping between the latent factors and the target comorbidities. Qualitative assessment of the empirical results by clinical experts suggests that the proposed model learns clinically interpretable phenotypes while being predictive of 30 day mortality. The proposed method can be readily adapted to any non-negative EHR data across various healthcare institutions.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge