Seyed Amin Tabatabaei
AnalyticsGPT: An LLM Workflow for Scientometric Question Answering
Feb 10, 2026Abstract:This paper introduces AnalyticsGPT, an intuitive and efficient large language model (LLM)-powered workflow for scientometric question answering. This underrepresented downstream task addresses the subcategory of meta-scientific questions concerning the "science of science." When compared to traditional scientific question answering based on papers, the task poses unique challenges in the planning phase. Namely, the need for named-entity recognition of academic entities within questions and multi-faceted data retrieval involving scientometric indices, e.g. impact factors. Beyond their exceptional capacity for treating traditional natural language processing tasks, LLMs have shown great potential in more complex applications, such as task decomposition and planning and reasoning. In this paper, we explore the application of LLMs to scientometric question answering, and describe an end-to-end system implementing a sequential workflow with retrieval-augmented generation and agentic concepts. We also address the secondary task of effectively synthesizing the data into presentable and well-structured high-level analyses. As a database for retrieval-augmented generation, we leverage a proprietary research performance assessment platform. For evaluation, we consult experienced subject matter experts and leverage LLMs-as-judges. In doing so, we provide valuable insights on the efficacy of LLMs towards a niche downstream task. Our (skeleton) code and prompts are available at: https://github.com/lyvykhang/llm-agents-scientometric-qa/tree/acl.
Can Large Language Models Serve as Effective Classifiers for Hierarchical Multi-Label Classification of Scientific Documents at Industrial Scale?
Dec 06, 2024



Abstract:We address the task of hierarchical multi-label classification (HMC) of scientific documents at an industrial scale, where hundreds of thousands of documents must be classified across thousands of dynamic labels. The rapid growth of scientific publications necessitates scalable and efficient methods for classification, further complicated by the evolving nature of taxonomies--where new categories are introduced, existing ones are merged, and outdated ones are deprecated. Traditional machine learning approaches, which require costly retraining with each taxonomy update, become impractical due to the high overhead of labelled data collection and model adaptation. Large Language Models (LLMs) have demonstrated great potential in complex tasks such as multi-label classification. However, applying them to large and dynamic taxonomies presents unique challenges as the vast number of labels can exceed LLMs' input limits. In this paper, we present novel methods that combine the strengths of LLMs with dense retrieval techniques to overcome these challenges. Our approach avoids retraining by leveraging zero-shot HMC for real-time label assignment. We evaluate the effectiveness of our methods on SSRN, a large repository of preprints spanning multiple disciplines, and demonstrate significant improvements in both classification accuracy and cost-efficiency. By developing a tailored evaluation framework for dynamic taxonomies and publicly releasing our code, this research provides critical insights into applying LLMs for document classification, where the number of classes corresponds to the number of nodes in a large taxonomy, at an industrial scale.
Identifying Patient Groups based on Frequent Patterns of Patient Samples
Apr 03, 2019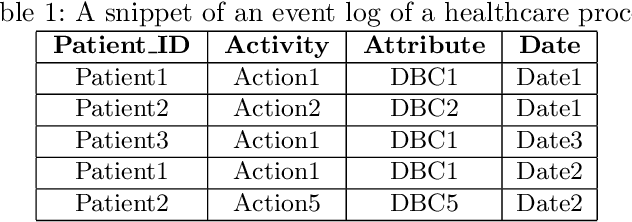
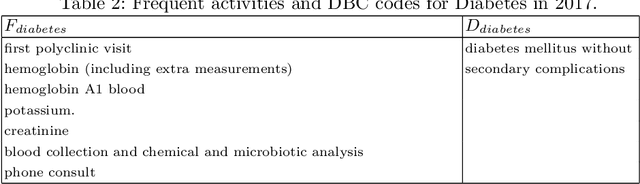
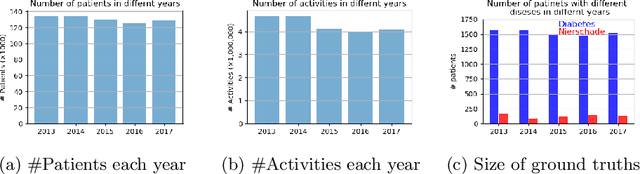

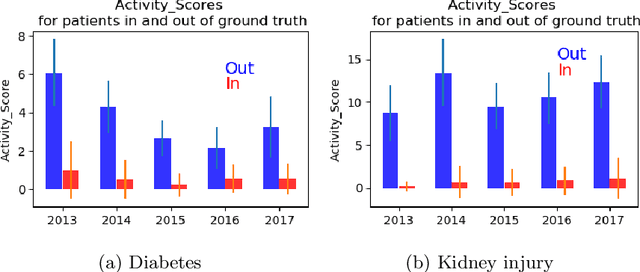
Abstract:Grouping patients meaningfully can give insights about the different types of patients, their needs, and the priorities. Finding groups that are meaningful is however very challenging as background knowledge is often required to determine what a useful grouping is. In this paper we propose an approach that is able to find groups of patients based on a small sample of positive examples given by a domain expert. Because of that, the approach relies on very limited efforts by the domain experts. The approach groups based on the activities and diagnostic/billing codes within health pathways of patients. To define such a grouping based on the sample of patients efficiently, frequent patterns of activities are discovered and used to measure the similarity between the care pathways of other patients to the patients in the sample group. This approach results in an insightful definition of the group. The proposed approach is evaluated using several datasets obtained from a large university medical center. The evaluation shows F1-scores of around 0.7 for grouping kidney injury and around 0.6 for diabetes.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge