Seunghyun Park
Donut: Document Understanding Transformer without OCR
Nov 30, 2021
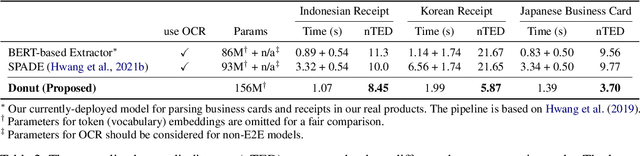
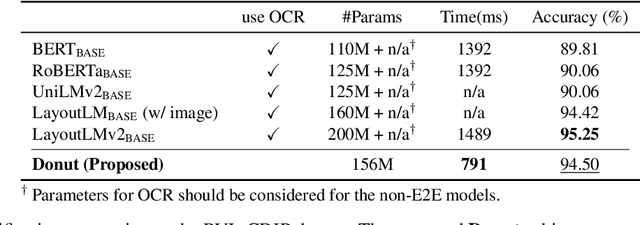
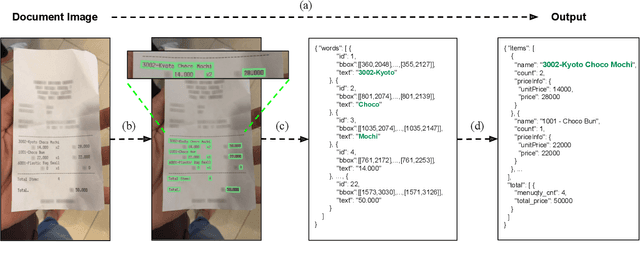

Abstract:Understanding document images (e.g., invoices) has been an important research topic and has many applications in document processing automation. Through the latest advances in deep learning-based Optical Character Recognition (OCR), current Visual Document Understanding (VDU) systems have come to be designed based on OCR. Although such OCR-based approach promise reasonable performance, they suffer from critical problems induced by the OCR, e.g., (1) expensive computational costs and (2) performance degradation due to the OCR error propagation. In this paper, we propose a novel VDU model that is end-to-end trainable without underpinning OCR framework. To this end, we propose a new task and a synthetic document image generator to pre-train the model to mitigate the dependencies on large-scale real document images. Our approach achieves state-of-the-art performance on various document understanding tasks in public benchmark datasets and private industrial service datasets. Through extensive experiments and analysis, we demonstrate the effectiveness of the proposed model especially with consideration for a real-world application.
SelfReg: Self-supervised Contrastive Regularization for Domain Generalization
Apr 20, 2021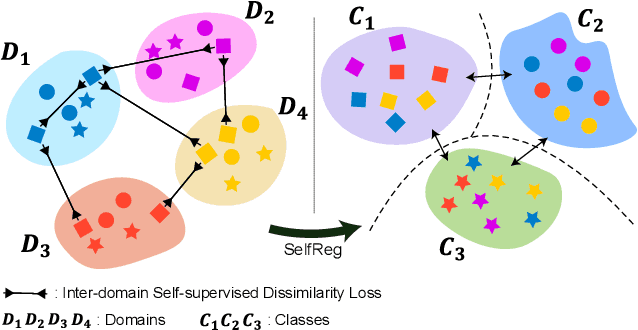
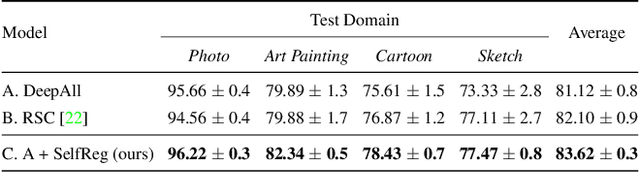
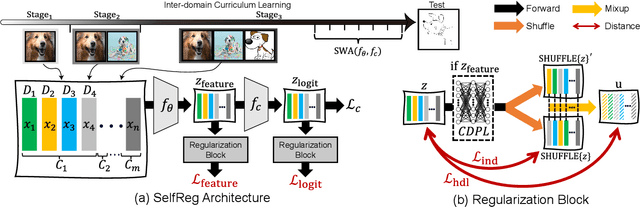
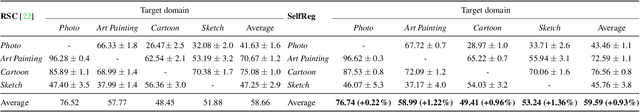
Abstract:In general, an experimental environment for deep learning assumes that the training and the test dataset are sampled from the same distribution. However, in real-world situations, a difference in the distribution between two datasets, domain shift, may occur, which becomes a major factor impeding the generalization performance of the model. The research field to solve this problem is called domain generalization, and it alleviates the domain shift problem by extracting domain-invariant features explicitly or implicitly. In recent studies, contrastive learning-based domain generalization approaches have been proposed and achieved high performance. These approaches require sampling of the negative data pair. However, the performance of contrastive learning fundamentally depends on quality and quantity of negative data pairs. To address this issue, we propose a new regularization method for domain generalization based on contrastive learning, self-supervised contrastive regularization (SelfReg). The proposed approach use only positive data pairs, thus it resolves various problems caused by negative pair sampling. Moreover, we propose a class-specific domain perturbation layer (CDPL), which makes it possible to effectively apply mixup augmentation even when only positive data pairs are used. The experimental results show that the techniques incorporated by SelfReg contributed to the performance in a compatible manner. In the recent benchmark, DomainBed, the proposed method shows comparable performance to the conventional state-of-the-art alternatives. Codes are available at https://github.com/dnap512/SelfReg.
Domain Generalization Needs Stochastic Weight Averaging for Robustness on Domain Shifts
Feb 17, 2021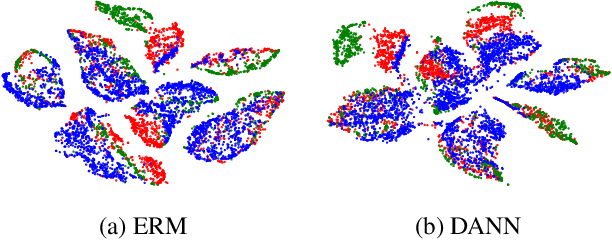
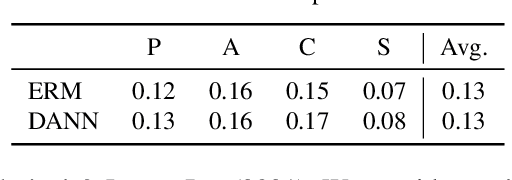
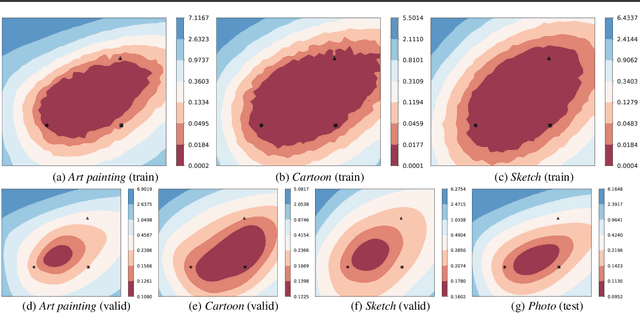
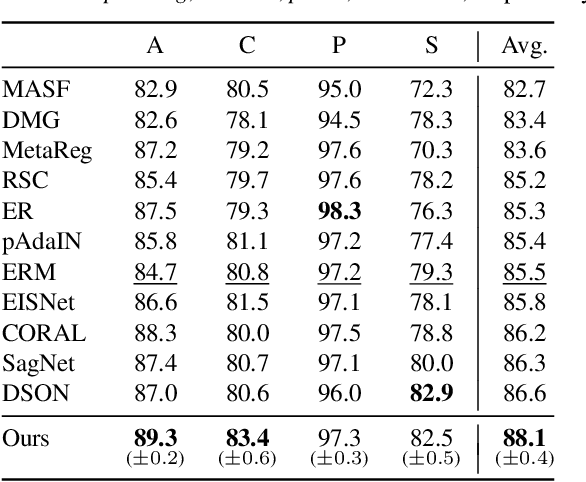
Abstract:Domain generalization aims to learn a generalizable model to unseen target domains from multiple source domains. Various approaches have been proposed to address this problem. However, recent benchmarks show that most of them do not provide significant improvements compared to the simple empirical risk minimization (ERM) in practical cases. In this paper, we analyze how ERM works in views of domain-invariant feature learning and domain-specific gradient normalization. In addition, we observe that ERM converges to a loss valley shared over multiple training domains and obtain an insight that a center of the valley generalizes better. To estimate the center, we employ stochastic weight averaging (SWA) and provide theoretical analysis describing how SWA supports the generalization bound for an unseen domain. As a result, we achieve state-of-the-art performances over all of widely used domain generalization benchmarks, namely PACS, VLCS, OfficeHome, TerraIncognita, and DomainNet with large margins. Further analysis reveals how SWA operates on domain generalization tasks.
Syntactic Question Abstraction and Retrieval for Data-Scarce Semantic Parsing
May 01, 2020
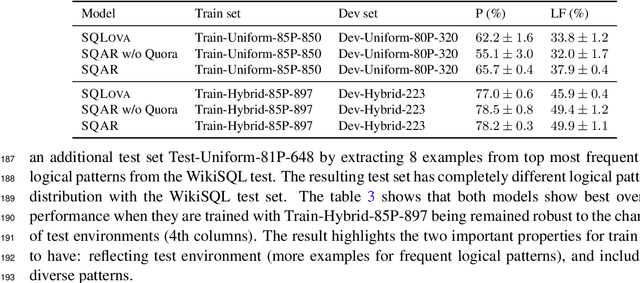

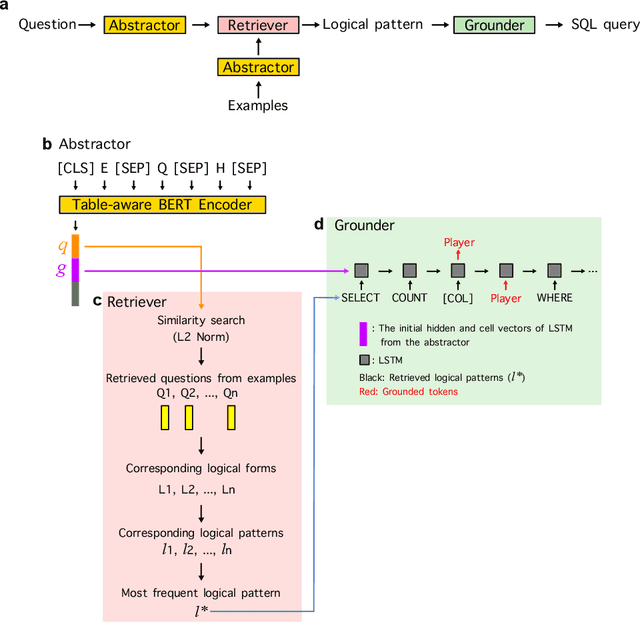

Abstract:Deep learning approaches to semantic parsing require a large amount of labeled data, but annotating complex logical forms is costly. Here, we propose Syntactic Question Abstraction and Retrieval (SQAR), a method to build a neural semantic parser that translates a natural language (NL) query to a SQL logical form (LF) with less than 1,000 annotated examples. SQAR first retrieves a logical pattern from the train data by computing the similarity between NL queries and then grounds a lexical information on the retrieved pattern in order to generate the final LF. We validate SQAR by training models using various small subsets of WikiSQL train data achieving up to 4.9% higher LF accuracy compared to the previous state-of-the-art models on WikiSQL test set. We also show that by using query-similarity to retrieve logical pattern, SQAR can leverage a paraphrasing dataset achieving up to 5.9% higher LF accuracy compared to the case where SQAR is trained by using only WikiSQL data. In contrast to a simple pattern classification approach, SQAR can generate unseen logical patterns upon the addition of new examples without re-training the model. We also discuss an ideal way to create cost efficient and robust train datasets when the data distribution can be approximated under a data-hungry setting.
Spatial Dependency Parsing for 2D Document Understanding
May 01, 2020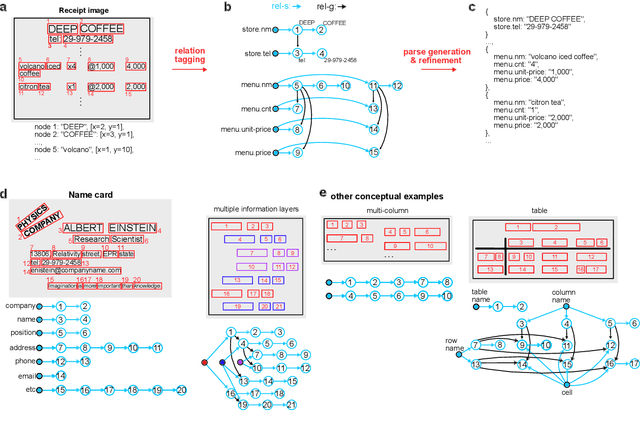
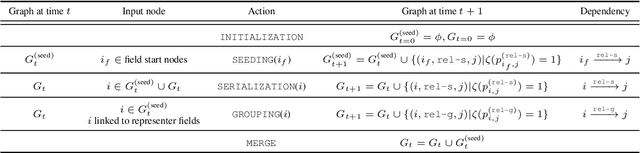

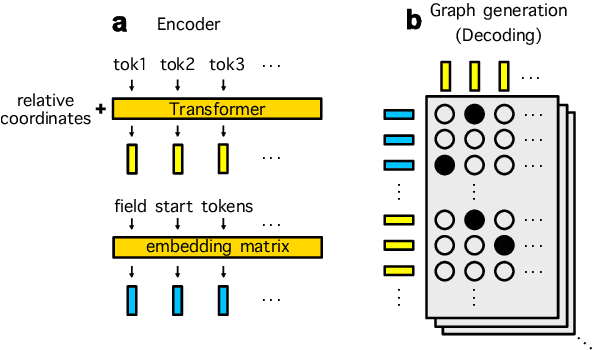
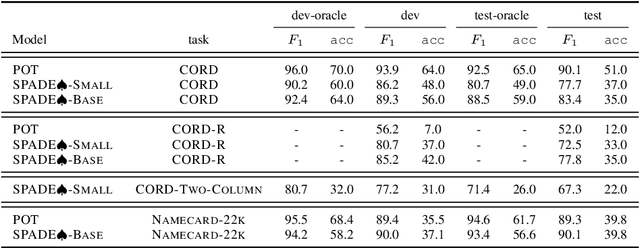
Abstract:Information Extraction (IE) for document images is often approached as a BIO tagging problem, where the model sequentially goes through and classifies each recognized input token into one of the information categories. However, such problem setup has two inherent limitations that (1) it can only extract a flat list of information and (2) it assumes that the input data is serialized, often by a simple rule-based script. Nevertheless, real-world documents often contain hierarchical information in the form of two-dimensional language data in which the serialization can be highly non-trivial. To tackle these issues, we propose SPADE$\spadesuit$ (SPatial DEpendency parser), an end-to-end spatial dependency parser that is serializer-free and capable of modeling an arbitrary number of information layers, making it suitable for parsing structure-rich documents such as receipts and multimodal documents such as name cards. We show that SPADE$\spadesuit$ outperforms the previous BIO tagging-based approach on name card parsing task and achieves comparable performance on receipt parsing task. Especially, when the receipt images have non-flat manifold representing physical distortion of receipt paper in real-world, SPADE$\spadesuit$ outperforms the tagging-based method by a large margin of 25.8% highlighting the strong performance of SPADE$\spadesuit$ over spatially complex document.
Pre-Training of Deep Bidirectional Protein Sequence Representations with Structural Information
Nov 25, 2019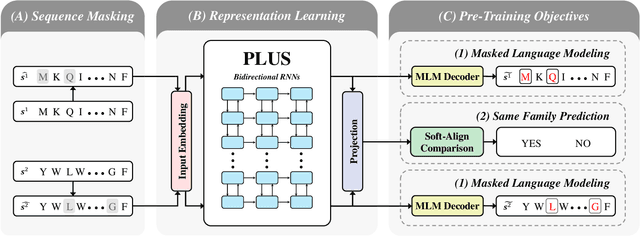
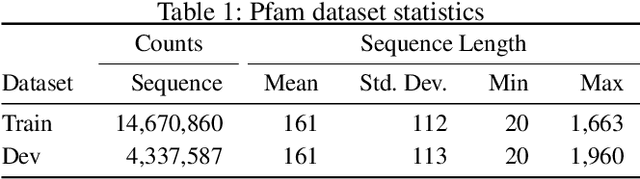
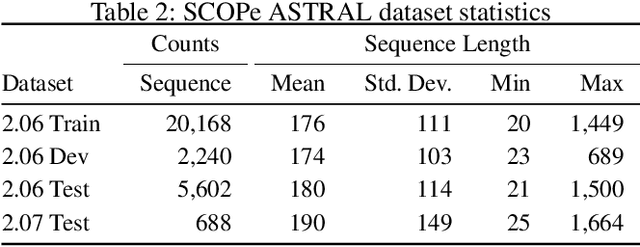
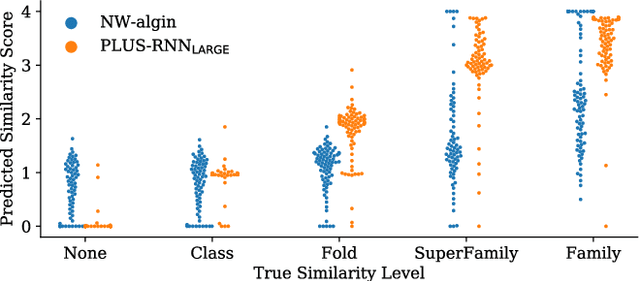
Abstract:A structure of a protein has a direct impact on its properties and functions. However, identification of structural similarity directly from amino acid sequences remains as a challenging problem in computational biology. In this paper, we introduce a novel BERT-wise pre-training scheme for a protein sequence representation model called PLUS, which stands for Protein sequence representations Learned Using Structural information. As natural language representation models capture syntactic and semantic information of words from a large unlabeled text corpus, PLUS captures structural information of amino acids from a large weakly labeled protein database. Since the Transformer encoder, BERT's original model architecture, has a severe computational requirement to handle long sequences, we first propose to combine a bidirectional recurrent neural network with the BERT-wise pre-training scheme. PLUS is designed to learn protein representations with two pre-training objectives, i.e., masked language modeling and same family prediction. Then, the pre-trained model can be fine-tuned for a wide range of tasks without training randomly initialized task-specific models from scratch. It obtains new state-of-the-art results on both (1) protein-level and (2) amino-acid-level tasks, outperforming many task-specific algorithms.
A Comprehensive Exploration on WikiSQL with Table-Aware Word Contextualization
Feb 04, 2019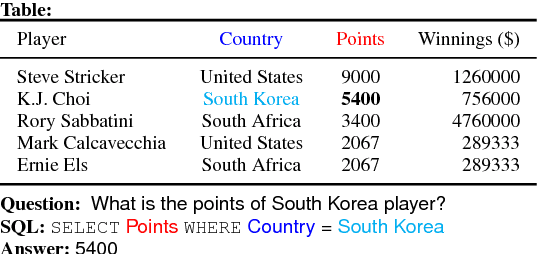
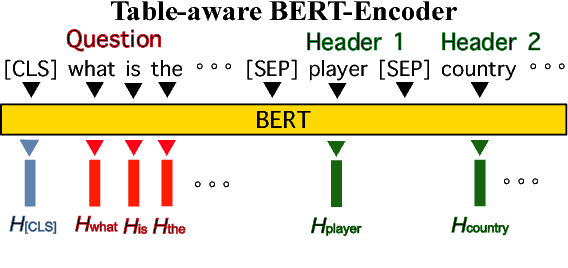
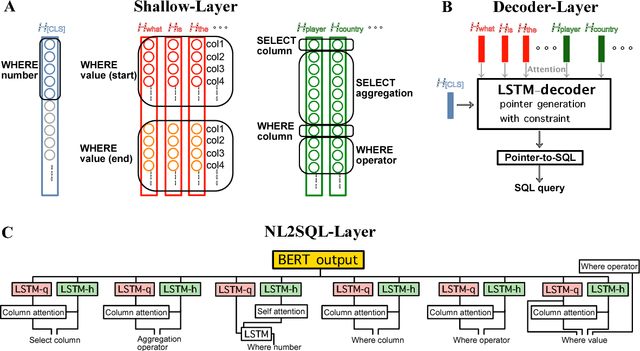
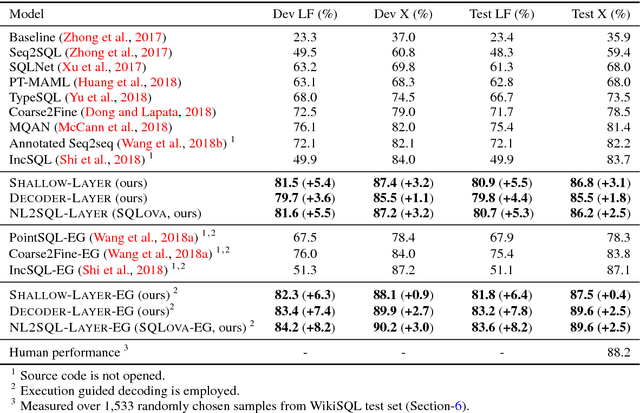
Abstract:WikiSQL is the task of mapping a natural language question to a SQL query given a table from a Wikipedia article. We first show that learning highly context- and table-aware word representations is arguably the most important consideration for achieving a high accuracy in the task. We explore three variants of BERT-based architecture and our best model outperforms the previous state of the art by 8.2% and 2.5% in logical form and execution accuracy, respectively. We provide a detailed analysis of the models to guide how word contextualization can be utilized in a such semantic parsing task. We then argue that this score is near the upper bound in WikiSQL, where we observe that the most of the evaluation errors are due to wrong annotations. We also measure human accuracy on a portion of the dataset and show that our model exceeds the human performance, at least by 1.4% execution accuracy.
deepTarget: End-to-end Learning Framework for microRNA Target Prediction using Deep Recurrent Neural Networks
Aug 19, 2016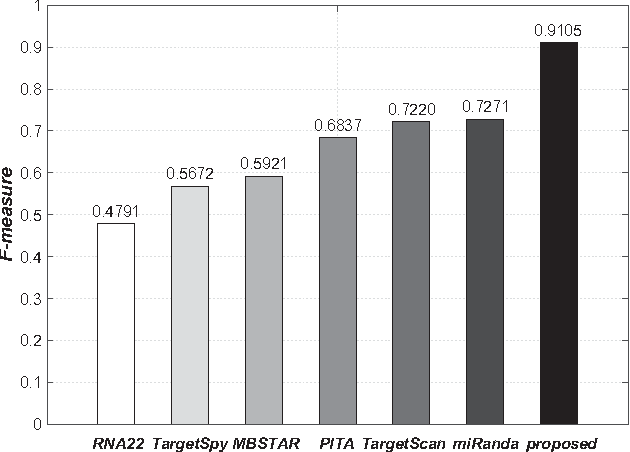
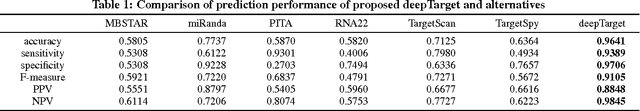
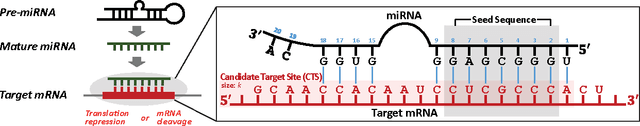
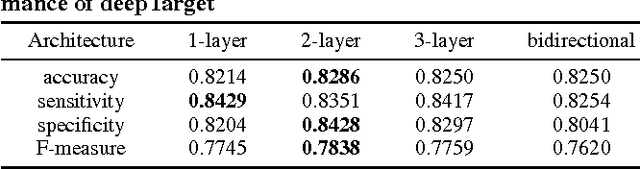
Abstract:MicroRNAs (miRNAs) are short sequences of ribonucleic acids that control the expression of target messenger RNAs (mRNAs) by binding them. Robust prediction of miRNA-mRNA pairs is of utmost importance in deciphering gene regulations but has been challenging because of high false positive rates, despite a deluge of computational tools that normally require laborious manual feature extraction. This paper presents an end-to-end machine learning framework for miRNA target prediction. Leveraged by deep recurrent neural networks-based auto-encoding and sequence-sequence interaction learning, our approach not only delivers an unprecedented level of accuracy but also eliminates the need for manual feature extraction. The performance gap between the proposed method and existing alternatives is substantial (over 25% increase in F-measure), and deepTarget delivers a quantum leap in the long-standing challenge of robust miRNA target prediction.
deepMiRGene: Deep Neural Network based Precursor microRNA Prediction
Apr 29, 2016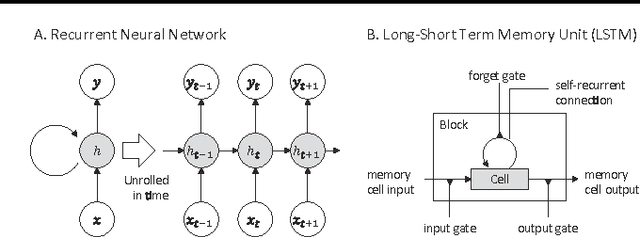

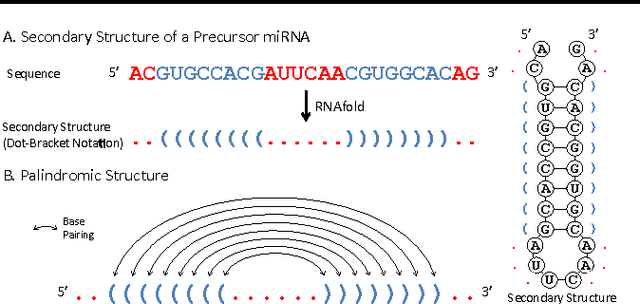
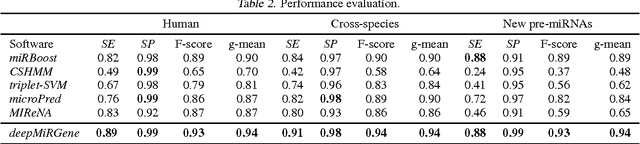
Abstract:Since microRNAs (miRNAs) play a crucial role in post-transcriptional gene regulation, miRNA identification is one of the most essential problems in computational biology. miRNAs are usually short in length ranging between 20 and 23 base pairs. It is thus often difficult to distinguish miRNA-encoding sequences from other non-coding RNAs and pseudo miRNAs that have a similar length, and most previous studies have recommended using precursor miRNAs instead of mature miRNAs for robust detection. A great number of conventional machine-learning-based classification methods have been proposed, but they often have the serious disadvantage of requiring manual feature engineering, and their performance is limited as well. In this paper, we propose a novel miRNA precursor prediction algorithm, deepMiRGene, based on recurrent neural networks, specifically long short-term memory networks. deepMiRGene automatically learns suitable features from the data themselves without manual feature engineering and constructs a model that can successfully reflect structural characteristics of precursor miRNAs. For the performance evaluation of our approach, we have employed several widely used evaluation metrics on three recent benchmark datasets and verified that deepMiRGene delivered comparable performance among the current state-of-the-art tools.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge