Sasank Chilamkurthy
Deep-ASPECTS: A Segmentation-Assisted Model for Stroke Severity Measurement
Mar 17, 2022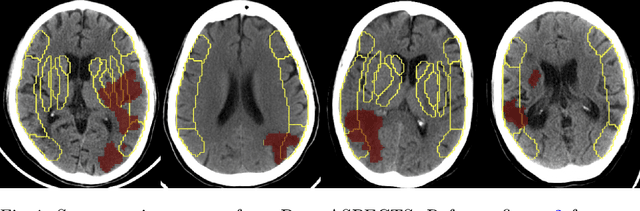
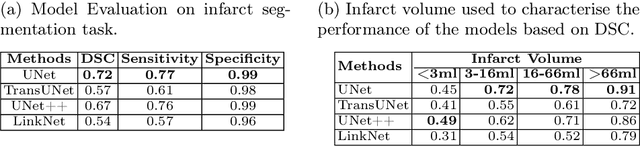
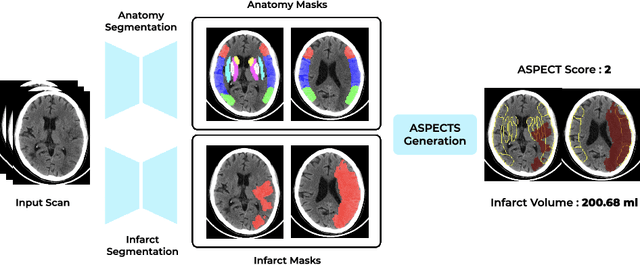
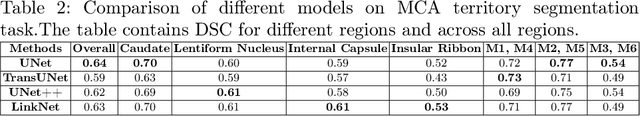
Abstract:A stroke occurs when an artery in the brain ruptures and bleeds or when the blood supply to the brain is cut off. Blood and oxygen cannot reach the brain's tissues due to the rupture or obstruction resulting in tissue death. The Middle cerebral artery (MCA) is the largest cerebral artery and the most commonly damaged vessel in stroke. The quick onset of a focused neurological deficit caused by interruption of blood flow in the territory supplied by the MCA is known as an MCA stroke. Alberta stroke programme early CT score (ASPECTS) is used to estimate the extent of early ischemic changes in patients with MCA stroke. This study proposes a deep learning-based method to score the CT scan for ASPECTS. Our work has three highlights. First, we propose a novel method for medical image segmentation for stroke detection. Second, we show the effectiveness of AI solution for fully-automated ASPECT scoring with reduced diagnosis time for a given non-contrast CT (NCCT) Scan. Our algorithms show a dice similarity coefficient of 0.64 for the MCA anatomy segmentation and 0.72 for the infarcts segmentation. Lastly, we show that our model's performance is inline with inter-reader variability between radiologists.
PyTorch: An Imperative Style, High-Performance Deep Learning Library
Dec 03, 2019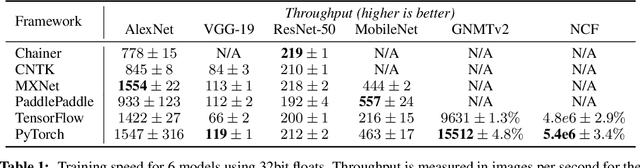

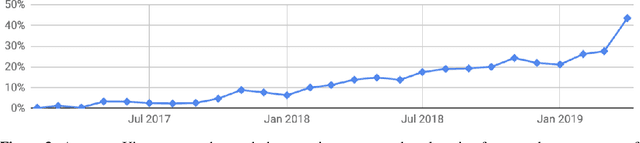
Abstract:Deep learning frameworks have often focused on either usability or speed, but not both. PyTorch is a machine learning library that shows that these two goals are in fact compatible: it provides an imperative and Pythonic programming style that supports code as a model, makes debugging easy and is consistent with other popular scientific computing libraries, while remaining efficient and supporting hardware accelerators such as GPUs. In this paper, we detail the principles that drove the implementation of PyTorch and how they are reflected in its architecture. We emphasize that every aspect of PyTorch is a regular Python program under the full control of its user. We also explain how the careful and pragmatic implementation of the key components of its runtime enables them to work together to achieve compelling performance. We demonstrate the efficiency of individual subsystems, as well as the overall speed of PyTorch on several common benchmarks.
Development and Validation of Deep Learning Algorithms for Detection of Critical Findings in Head CT Scans
Apr 12, 2018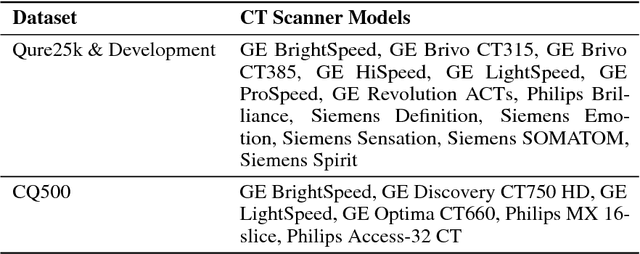
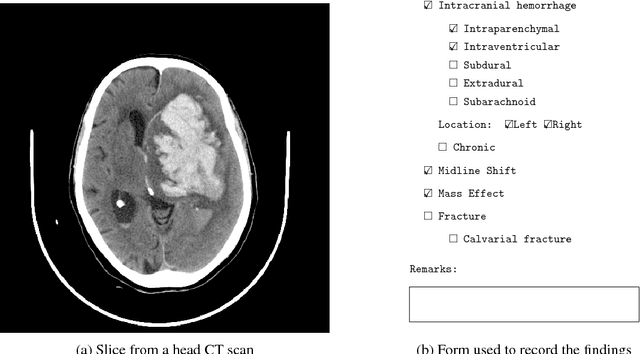
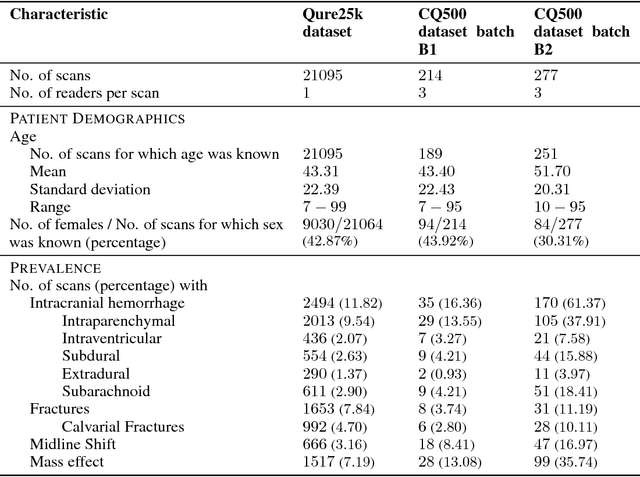
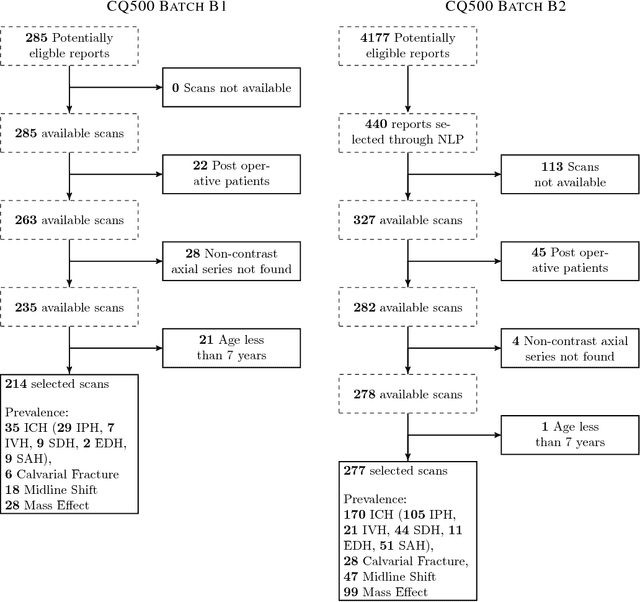
Abstract:Importance: Non-contrast head CT scan is the current standard for initial imaging of patients with head trauma or stroke symptoms. Objective: To develop and validate a set of deep learning algorithms for automated detection of following key findings from non-contrast head CT scans: intracranial hemorrhage (ICH) and its types, intraparenchymal (IPH), intraventricular (IVH), subdural (SDH), extradural (EDH) and subarachnoid (SAH) hemorrhages, calvarial fractures, midline shift and mass effect. Design and Settings: We retrospectively collected a dataset containing 313,318 head CT scans along with their clinical reports from various centers. A part of this dataset (Qure25k dataset) was used to validate and the rest to develop algorithms. Additionally, a dataset (CQ500 dataset) was collected from different centers in two batches B1 & B2 to clinically validate the algorithms. Main Outcomes and Measures: Original clinical radiology report and consensus of three independent radiologists were considered as gold standard for Qure25k and CQ500 datasets respectively. Area under receiver operating characteristics curve (AUC) for each finding was primarily used to evaluate the algorithms. Results: Qure25k dataset contained 21,095 scans (mean age 43.31; 42.87% female) while batches B1 and B2 of CQ500 dataset consisted of 214 (mean age 43.40; 43.92% female) and 277 (mean age 51.70; 30.31% female) scans respectively. On Qure25k dataset, the algorithms achieved AUCs of 0.9194, 0.8977, 0.9559, 0.9161, 0.9288 and 0.9044 for detecting ICH, IPH, IVH, SDH, EDH and SAH respectively. AUCs for the same on CQ500 dataset were 0.9419, 0.9544, 0.9310, 0.9521, 0.9731 and 0.9574 respectively. For detecting calvarial fractures, midline shift and mass effect, AUCs on Qure25k dataset were 0.9244, 0.9276 and 0.8583 respectively, while AUCs on CQ500 dataset were 0.9624, 0.9697 and 0.9216 respectively.
2D-3D Fully Convolutional Neural Networks for Cardiac MR Segmentation
Jul 31, 2017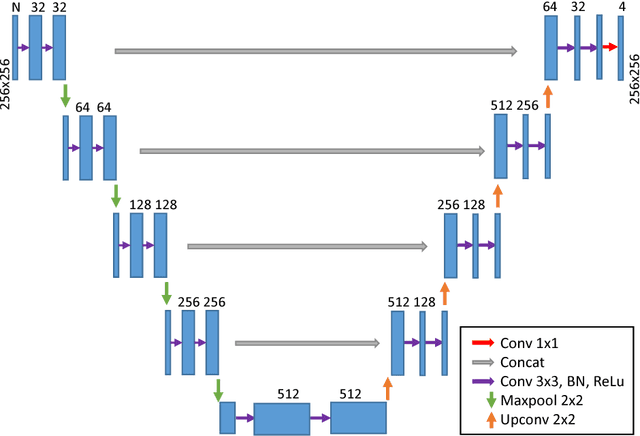
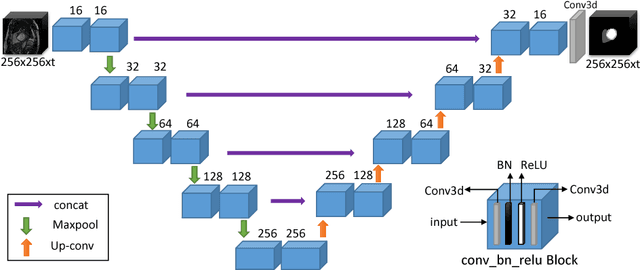
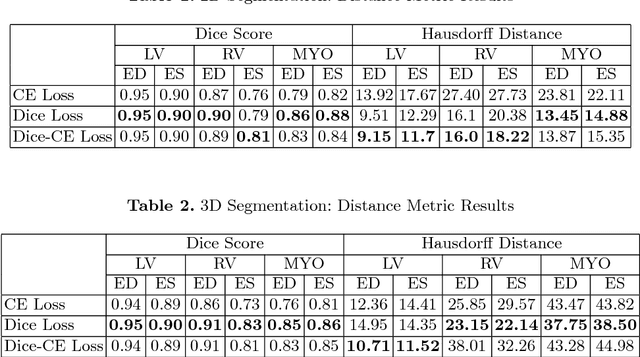

Abstract:In this paper, we develop a 2D and 3D segmentation pipelines for fully automated cardiac MR image segmentation using Deep Convolutional Neural Networks (CNN). Our models are trained end-to-end from scratch using the ACD Challenge 2017 dataset comprising of 100 studies, each containing Cardiac MR images in End Diastole and End Systole phase. We show that both our segmentation models achieve near state-of-the-art performance scores in terms of distance metrics and have convincing accuracy in terms of clinical parameters. A comparative analysis is provided by introducing a novel dice loss function and its combination with cross entropy loss. By exploring different network structures and comprehensive experiments, we discuss several key insights to obtain optimal model performance, which also is central to the theme of this challenge.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge