Sarah Parisot
Learning Conditioned Graph Structures for Interpretable Visual Question Answering
Nov 01, 2018
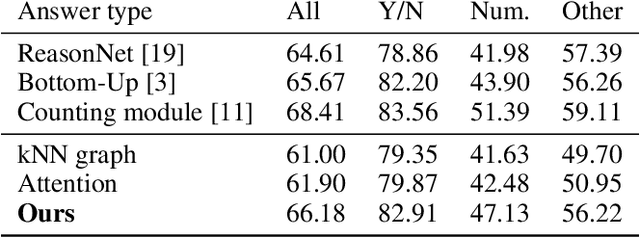
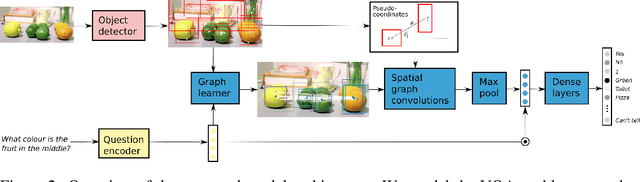
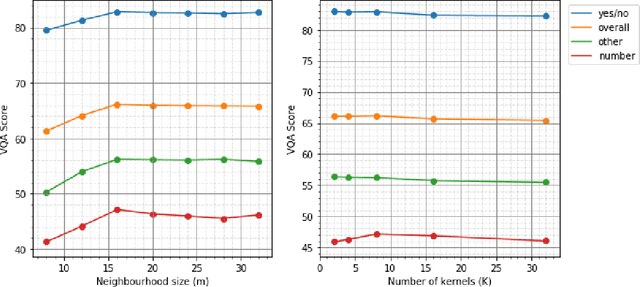
Abstract:Visual Question answering is a challenging problem requiring a combination of concepts from Computer Vision and Natural Language Processing. Most existing approaches use a two streams strategy, computing image and question features that are consequently merged using a variety of techniques. Nonetheless, very few rely on higher level image representations, which can capture semantic and spatial relationships. In this paper, we propose a novel graph-based approach for Visual Question Answering. Our method combines a graph learner module, which learns a question specific graph representation of the input image, with the recent concept of graph convolutions, aiming to learn image representations that capture question specific interactions. We test our approach on the VQA v2 dataset using a simple baseline architecture enhanced by the proposed graph learner module. We obtain promising results with 66.18% accuracy and demonstrate the interpretability of the proposed method. Code can be found at github.com/aimbrain/vqa-project.
Uncertainty Quantification in CNN-Based Surface Prediction Using Shape Priors
Jul 30, 2018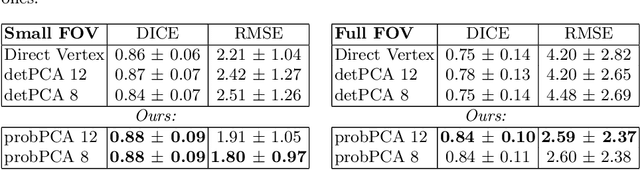
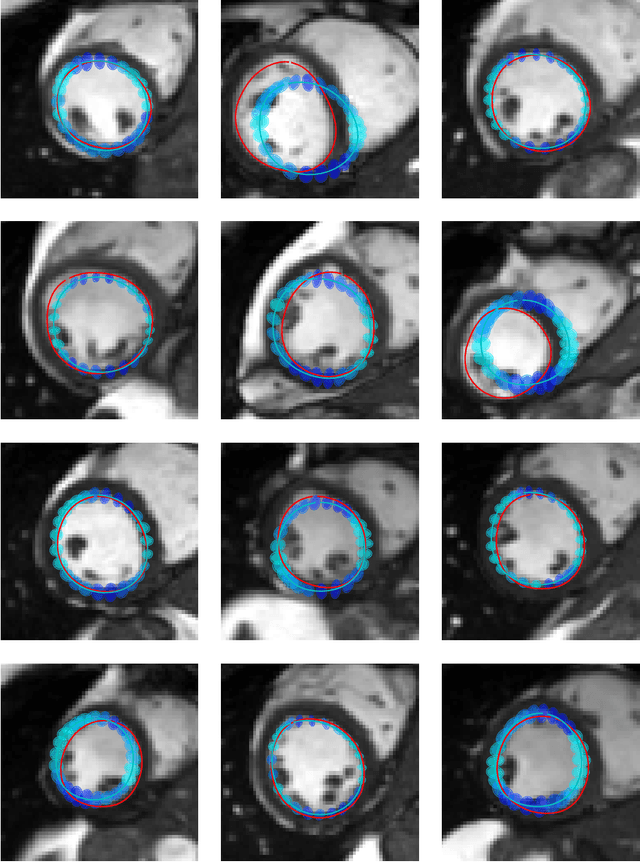
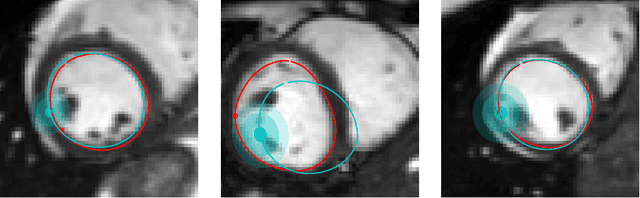
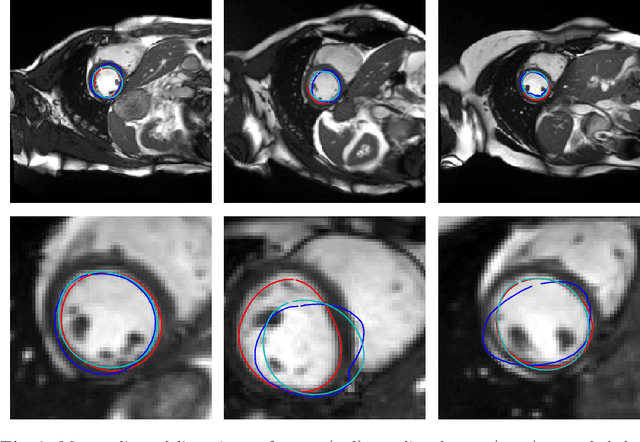
Abstract:Surface reconstruction is a vital tool in a wide range of areas of medical image analysis and clinical research. Despite the fact that many methods have proposed solutions to the reconstruction problem, most, due to their deterministic nature, do not directly address the issue of quantifying uncertainty associated with their predictions. We remedy this by proposing a novel probabilistic deep learning approach capable of simultaneous surface reconstruction and associated uncertainty prediction. The method incorporates prior shape information in the form of a principal component analysis (PCA) model. Experiments using the UK Biobank data show that our probabilistic approach outperforms an analogous deterministic PCA-based method in the task of 2D organ delineation and quantifies uncertainty by formulating distributions over predicted surface vertex positions.
Disease Prediction using Graph Convolutional Networks: Application to Autism Spectrum Disorder and Alzheimer's Disease
Jun 05, 2018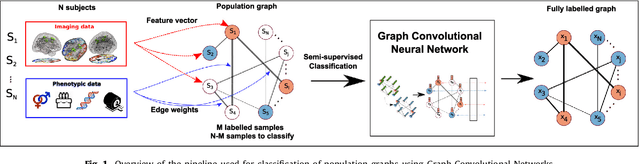
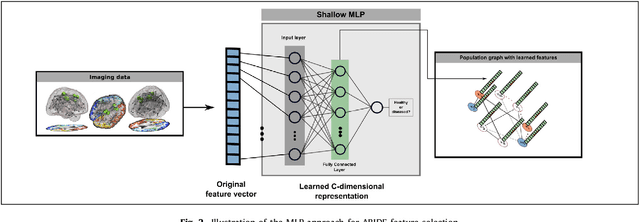
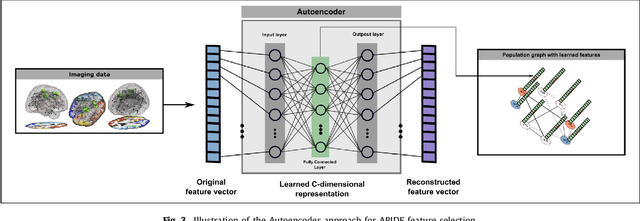
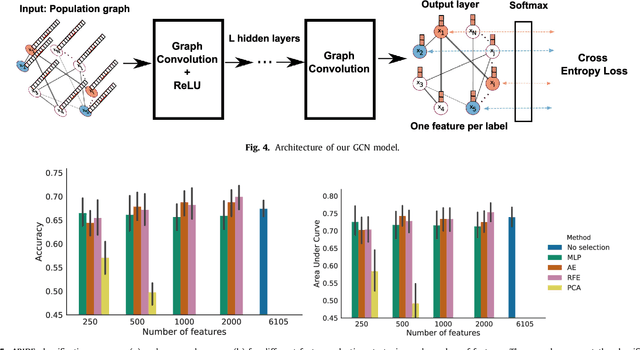
Abstract:Graphs are widely used as a natural framework that captures interactions between individual elements represented as nodes in a graph. In medical applications, specifically, nodes can represent individuals within a potentially large population (patients or healthy controls) accompanied by a set of features, while the graph edges incorporate associations between subjects in an intuitive manner. This representation allows to incorporate the wealth of imaging and non-imaging information as well as individual subject features simultaneously in disease classification tasks. Previous graph-based approaches for supervised or unsupervised learning in the context of disease prediction solely focus on pairwise similarities between subjects, disregarding individual characteristics and features, or rather rely on subject-specific imaging feature vectors and fail to model interactions between them. In this paper, we present a thorough evaluation of a generic framework that leverages both imaging and non-imaging information and can be used for brain analysis in large populations. This framework exploits Graph Convolutional Networks (GCNs) and involves representing populations as a sparse graph, where its nodes are associated with imaging-based feature vectors, while phenotypic information is integrated as edge weights. The extensive evaluation explores the effect of each individual component of this framework on disease prediction performance and further compares it to different baselines. The framework performance is tested on two large datasets with diverse underlying data, ABIDE and ADNI, for the prediction of Autism Spectrum Disorder and conversion to Alzheimer's disease, respectively. Our analysis shows that our novel framework can improve over state-of-the-art results on both databases, with 70.4% classification accuracy for ABIDE and 80.0% for ADNI.
Spectral Graph Convolutions for Population-based Disease Prediction
Jun 21, 2017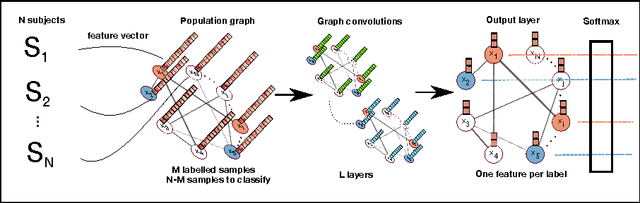
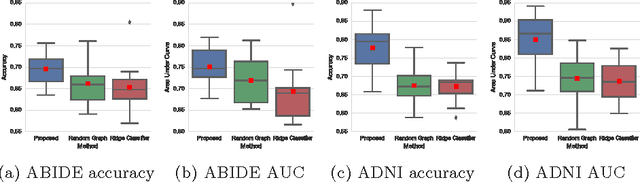
Abstract:Exploiting the wealth of imaging and non-imaging information for disease prediction tasks requires models capable of representing, at the same time, individual features as well as data associations between subjects from potentially large populations. Graphs provide a natural framework for such tasks, yet previous graph-based approaches focus on pairwise similarities without modelling the subjects' individual characteristics and features. On the other hand, relying solely on subject-specific imaging feature vectors fails to model the interaction and similarity between subjects, which can reduce performance. In this paper, we introduce the novel concept of Graph Convolutional Networks (GCN) for brain analysis in populations, combining imaging and non-imaging data. We represent populations as a sparse graph where its vertices are associated with image-based feature vectors and the edges encode phenotypic information. This structure was used to train a GCN model on partially labelled graphs, aiming to infer the classes of unlabelled nodes from the node features and pairwise associations between subjects. We demonstrate the potential of the method on the challenging ADNI and ABIDE databases, as a proof of concept of the benefit from integrating contextual information in classification tasks. This has a clear impact on the quality of the predictions, leading to 69.5% accuracy for ABIDE (outperforming the current state of the art of 66.8%) and 77% for ADNI for prediction of MCI conversion, significantly outperforming standard linear classifiers where only individual features are considered.
Distance Metric Learning using Graph Convolutional Networks: Application to Functional Brain Networks
Jun 14, 2017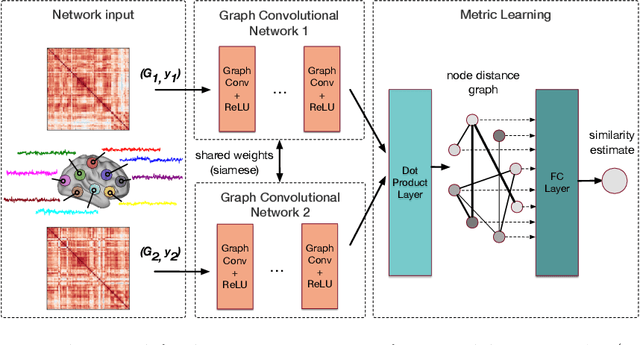

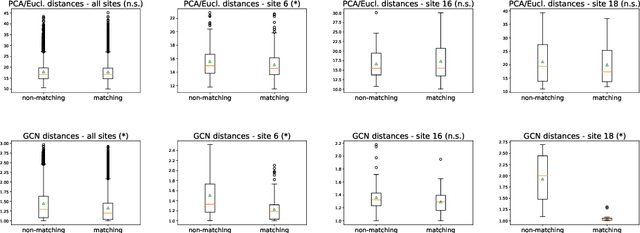
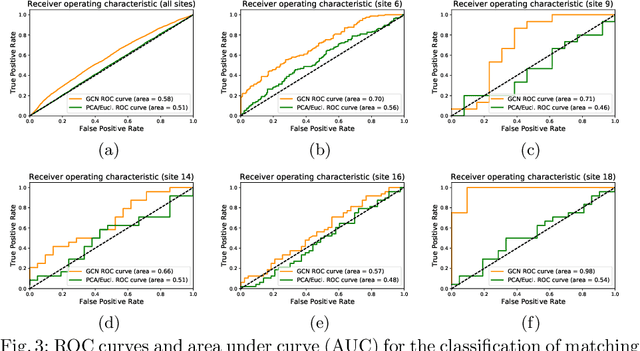
Abstract:Evaluating similarity between graphs is of major importance in several computer vision and pattern recognition problems, where graph representations are often used to model objects or interactions between elements. The choice of a distance or similarity metric is, however, not trivial and can be highly dependent on the application at hand. In this work, we propose a novel metric learning method to evaluate distance between graphs that leverages the power of convolutional neural networks, while exploiting concepts from spectral graph theory to allow these operations on irregular graphs. We demonstrate the potential of our method in the field of connectomics, where neuronal pathways or functional connections between brain regions are commonly modelled as graphs. In this problem, the definition of an appropriate graph similarity function is critical to unveil patterns of disruptions associated with certain brain disorders. Experimental results on the ABIDE dataset show that our method can learn a graph similarity metric tailored for a clinical application, improving the performance of a simple k-nn classifier by 11.9% compared to a traditional distance metric.
Exploring Heritability of Functional Brain Networks with Inexact Graph Matching
Mar 29, 2017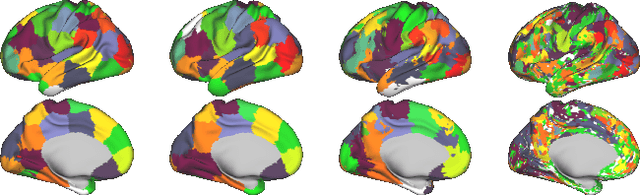
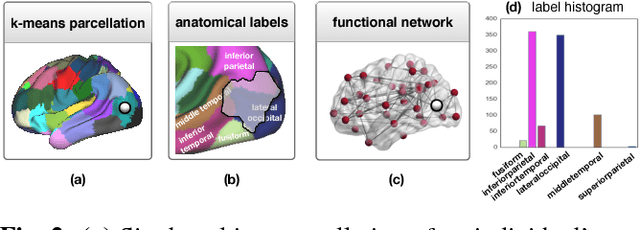
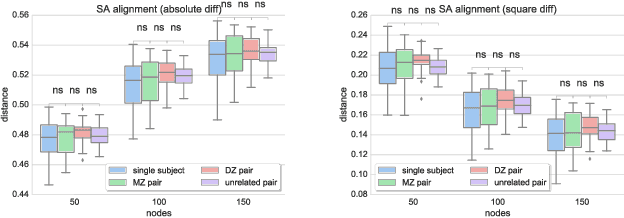
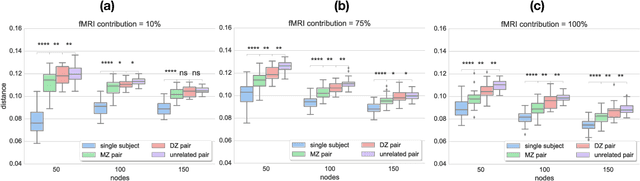
Abstract:Data-driven brain parcellations aim to provide a more accurate representation of an individual's functional connectivity, since they are able to capture individual variability that arises due to development or disease. This renders comparisons between the emerging brain connectivity networks more challenging, since correspondences between their elements are not preserved. Unveiling these correspondences is of major importance to keep track of local functional connectivity changes. We propose a novel method based on graph edit distance for the comparison of brain graphs directly in their domain, that can accurately reflect similarities between individual networks while providing the network element correspondences. This method is validated on a dataset of 116 twin subjects provided by the Human Connectome Project.
Proceedings of the Workshop on Brain Analysis using COnnectivity Networks - BACON 2016
Nov 24, 2016Abstract:Understanding brain connectivity in a network-theoretic context has shown much promise in recent years. This type of analysis identifies brain organisational principles, bringing a new perspective to neuroscience. At the same time, large public databases of connectomic data are now available. However, connectome analysis is still an emerging field and there is a crucial need for robust computational methods to fully unravelits potential. This workshop provides a platform to discuss the development of new analytic techniques; methods for evaluating and validating commonly used approaches; as well as the effects of variations in pre-processing steps.
Comparison of Brain Networks with Unknown Correspondences
Nov 15, 2016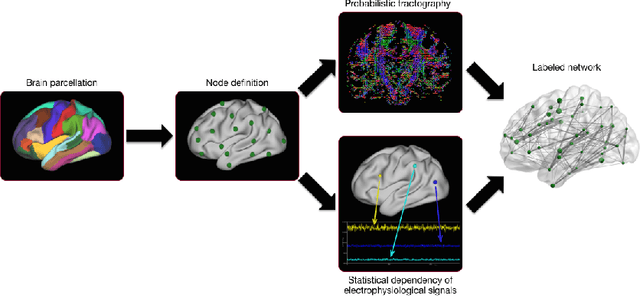
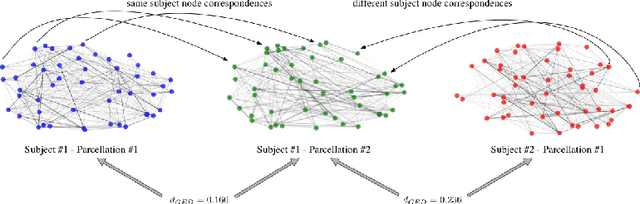
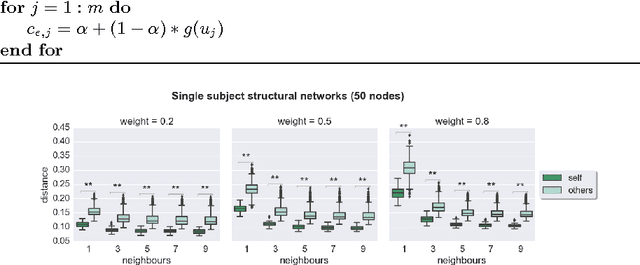
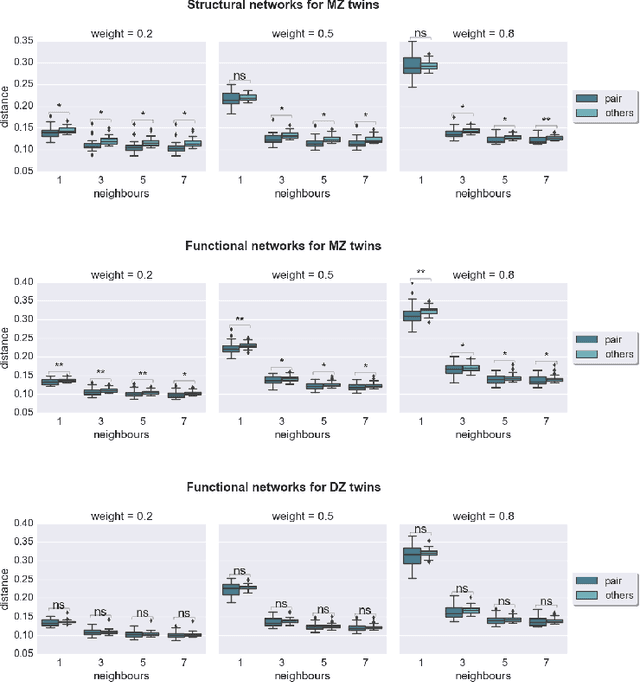
Abstract:Graph theory has drawn a lot of attention in the field of Neuroscience during the last decade, mainly due to the abundance of tools that it provides to explore the interactions of elements in a complex network like the brain. The local and global organization of a brain network can shed light on mechanisms of complex cognitive functions, while disruptions within the network can be linked to neurodevelopmental disorders. In this effort, the construction of a representative brain network for each individual is critical for further analysis. Additionally, graph comparison is an essential step for inference and classification analyses on brain graphs. In this work we explore a method based on graph edit distance for evaluating graph similarity, when correspondences between network elements are unknown due to different underlying subdivisions of the brain. We test this method on 30 unrelated subjects as well as 40 twin pairs and show that this method can accurately reflect the higher similarity between two related networks compared to unrelated ones, while identifying node correspondences.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge