Samuel Kaski
Department of Computer Science, Aalto University, Department of Computer Science, University of Manchester
Scalable Bayesian Non-linear Matrix Completion
Jul 31, 2019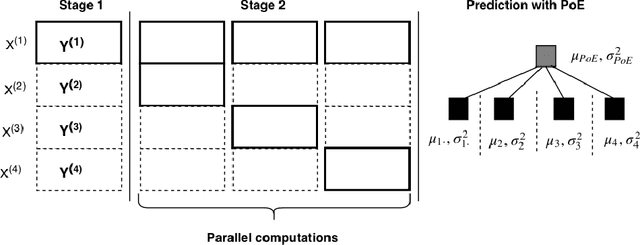
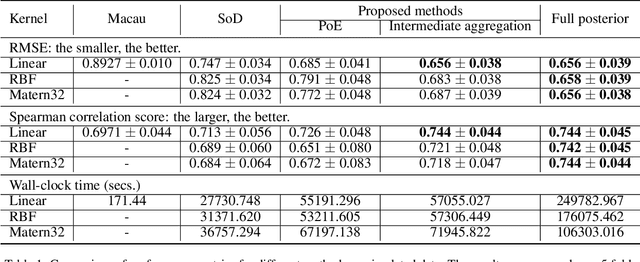
Abstract:Matrix completion aims to predict missing elements in a partially observed data matrix which in typical applications, such as collaborative filtering, is large and extremely sparsely observed. A standard solution is matrix factorization, which predicts unobserved entries as linear combinations of latent variables. We generalize to non-linear combinations in massive-scale matrices. Bayesian approaches have been proven beneficial in linear matrix completion, but not applied in the more general non-linear case, due to limited scalability. We introduce a Bayesian non-linear matrix completion algorithm, which is based on a recent Bayesian formulation of Gaussian process latent variable models. To solve the challenges regarding scalability and computation, we propose a data-parallel distributed computational approach with a restricted communication scheme. We evaluate our method on challenging out-of-matrix prediction tasks using both simulated and real-world data.
Learning spectrograms with convolutional spectral kernels
May 23, 2019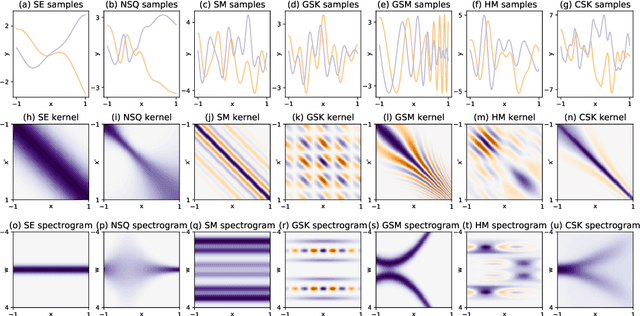

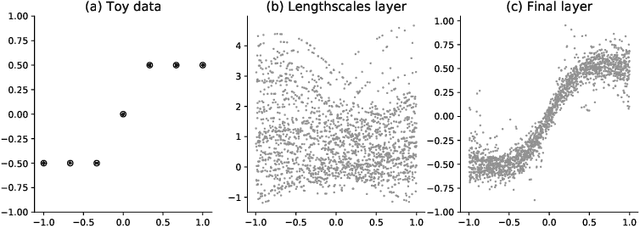

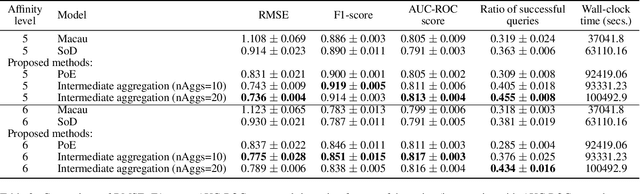
Abstract:We introduce the convolutional spectral kernel (CSK), a novel family of interpretable and non-stationary kernels derived from the convolution of two imaginary radial basis functions. We propose the input-frequency spectrogram as a novel tool to analyze nonparametric kernels as well as the kernels of deep Gaussian processes (DGPs). Observing through the lens of the spectrogram, we shed light on the interpretability of deep models, along with useful insights for effective inference. We also present scalable variational and stochastic Hamiltonian Monte Carlo inference to learn rich, yet interpretable frequency patterns from data using DGPs constructed via covariance functions. Empirically we show on simulated and real-world datasets that CSK extracts meaningful non-stationary periodicities.
Active Learning for Decision-Making from Imbalanced Observational Data
Apr 10, 2019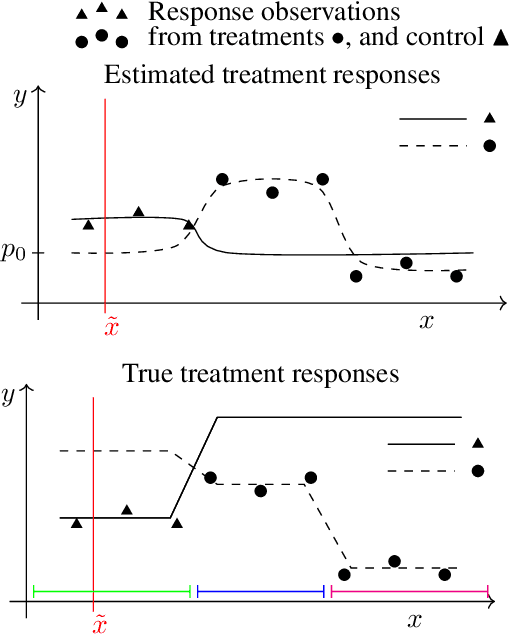
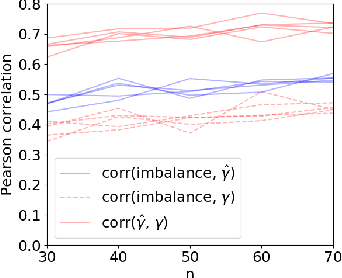
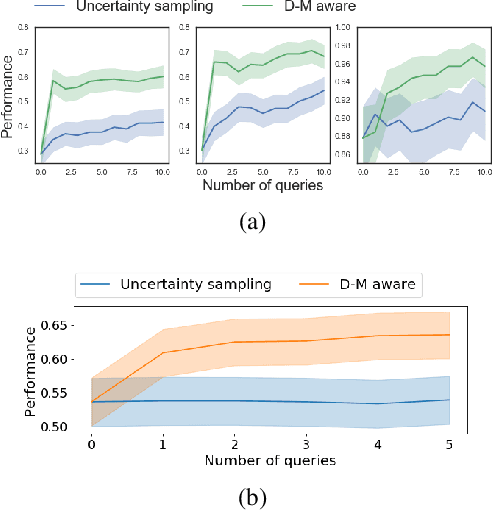
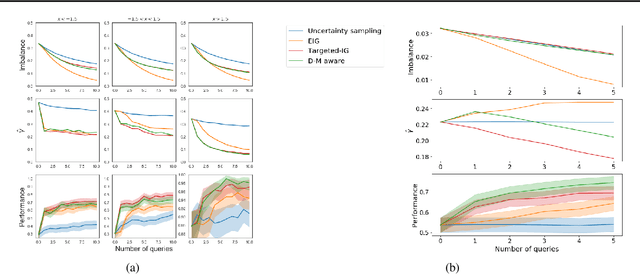
Abstract:Machine learning can help personalized decision support by learning models to predict individual treatment effects (ITE). This work studies the reliability of prediction-based decision-making in a task of deciding which action $a$ to take for a target unit after observing its covariates $\tilde{x}$ and predicted outcomes $\hat{p}(\tilde{y} \mid \tilde{x}, a)$. An example case is personalized medicine and the decision of which treatment to give to a patient. A common problem when learning these models from observational data is imbalance, that is, difference in treated/control covariate distributions, which is known to increase the upper bound of the expected ITE estimation error. We propose to assess the decision-making reliability by estimating the ITE model's Type S error rate, which is the probability of the model inferring the sign of the treatment effect wrong. Furthermore, we use the estimated reliability as a criterion for active learning, in order to collect new (possibly expensive) observations, instead of making a forced choice based on unreliable predictions. We demonstrate the effectiveness of this decision-making aware active learning in two decision-making tasks: in simulated data with binary outcomes and in a medical dataset with synthetic and continuous treatment outcomes.
Human-in-the-loop Active Covariance Learning for Improving Prediction in Small Data Sets
Mar 18, 2019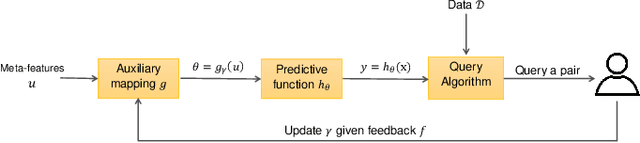
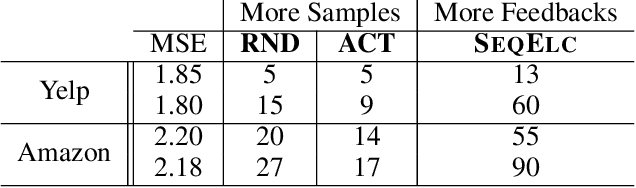
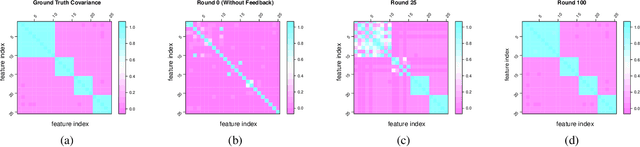
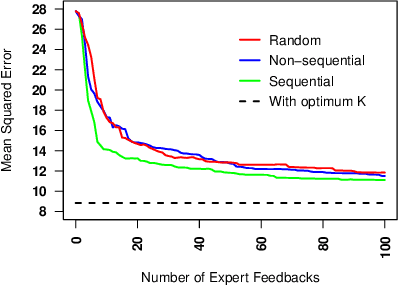
Abstract:Learning predictive models from small high-dimensional data sets is a key problem in high-dimensional statistics. Expert knowledge elicitation can help, and a strong line of work focuses on directly eliciting informative prior distributions for parameters. This either requires considerable statistical expertise or is laborious, as the emphasis has been on accuracy and not on efficiency of the process. Another line of work queries about importance of features one at a time, assuming them to be independent and hence missing covariance information. In contrast, we propose eliciting expert knowledge about pairwise feature similarities, to borrow statistical strength in the predictions, and using sequential decision making techniques to minimize the effort of the expert. Empirical results demonstrate improvement in predictive performance on both simulated and real data, in high-dimensional linear regression tasks, where we learn the covariance structure with a Gaussian process, based on sequential elicitation.
Embarrassingly parallel MCMC using deep invertible transformations
Mar 11, 2019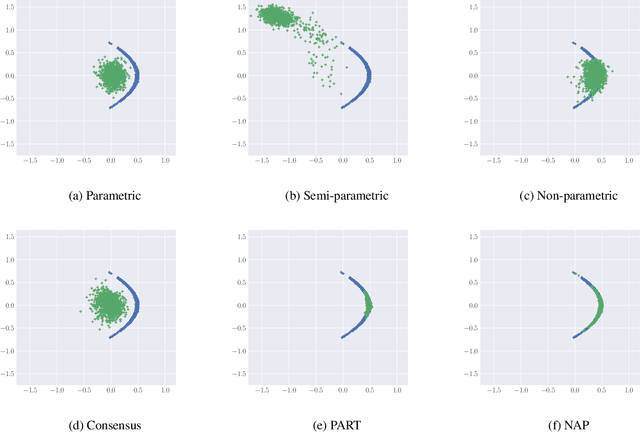
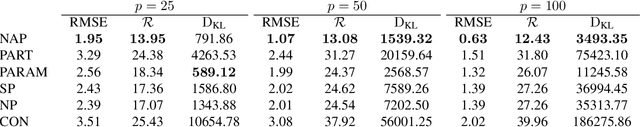
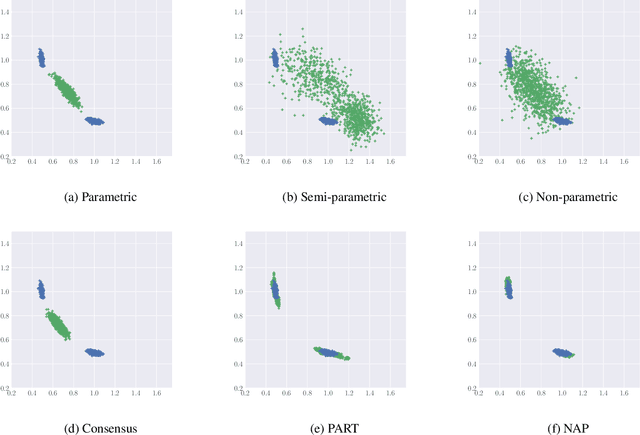

Abstract:While MCMC methods have become a main work-horse for Bayesian inference, scaling them to large distributed datasets is still a challenge. Embarrassingly parallel MCMC strategies take a divide-and-conquer stance to achieve this by writing the target posterior as a product of subposteriors, running MCMC for each of them in parallel and subsequently combining the results. The challenge then lies in devising efficient aggregation strategies. Current strategies trade-off between approximation quality, and costs of communication and computation. In this work, we introduce a novel method that addresses these issues simultaneously. Our key insight is to introduce a deep invertible transformation to approximate each of the subposteriors. These approximations can be made accurate even for complex distributions and serve as intermediate representations, keeping the total communication cost limited. Moreover, they enable us to sample from the product of the subposteriors using an efficient and stable importance sampling scheme. We demonstrate the approach outperforms available state-of-the-art methods in a range of challenging scenarios, including high-dimensional and heterogeneous subposteriors.
Representation Transfer for Differentially Private Drug Sensitivity Prediction
Jan 29, 2019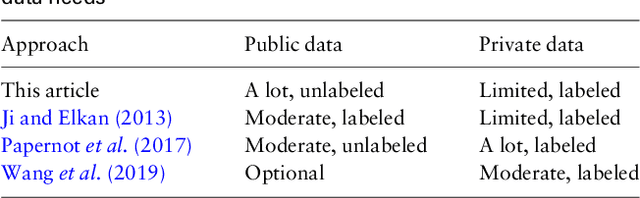
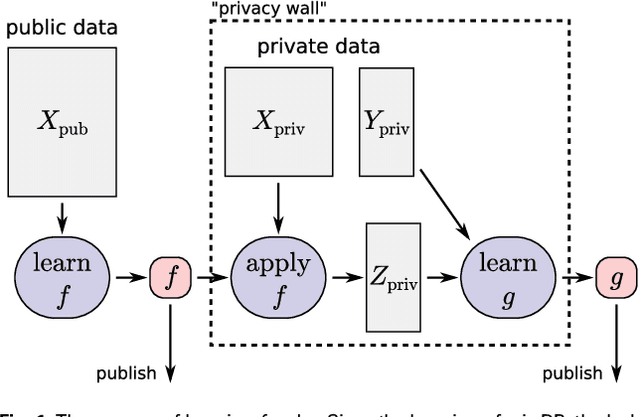
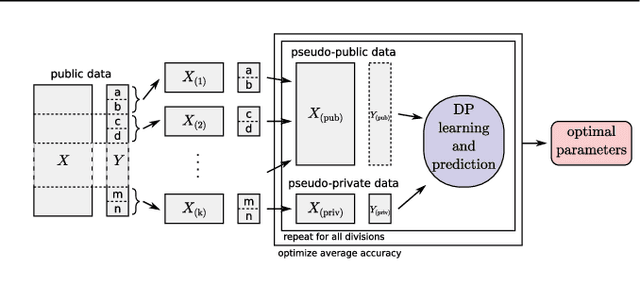
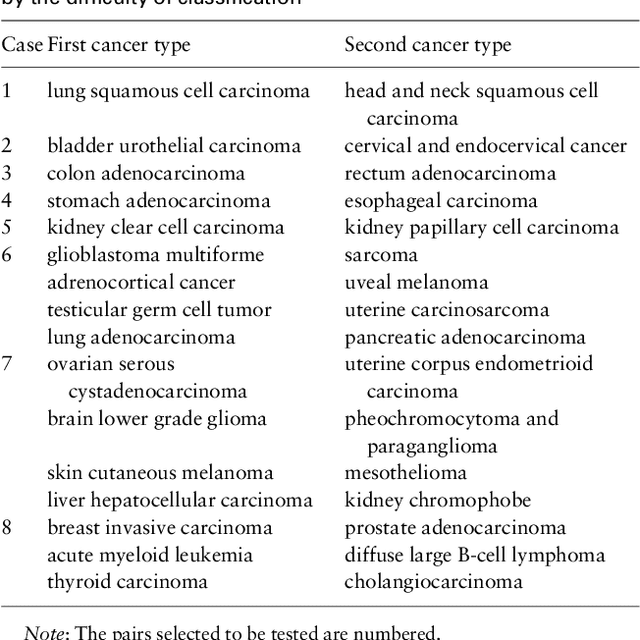
Abstract:Motivation: Human genomic datasets often contain sensitive information that limits use and sharing of the data. In particular, simple anonymisation strategies fail to provide sufficient level of protection for genomic data, because the data are inherently identifiable. Differentially private machine learning can help by guaranteeing that the published results do not leak too much information about any individual data point. Recent research has reached promising results on differentially private drug sensitivity prediction using gene expression data. Differentially private learning with genomic data is challenging because it is more difficult to guarantee the privacy in high dimensions. Dimensionality reduction can help, but if the dimension reduction mapping is learned from the data, then it needs to be differentially private too, which can carry a significant privacy cost. Furthermore, the selection of any hyperparameters (such as the target dimensionality) needs to also avoid leaking private information. Results: We study an approach that uses a large public dataset of similar type to learn a compact representation for differentially private learning. We compare three representation learning methods: variational autoencoders, PCA and random projection. We solve two machine learning tasks on gene expression of cancer cell lines: cancer type classification, and drug sensitivity prediction. The experiments demonstrate significant benefit from all representation learning methods with variational autoencoders providing the most accurate predictions most often. Our results significantly improve over previous state-of-the-art in accuracy of differentially private drug sensitivity prediction.
Recovering Pairwise Interactions Using Neural Networks
Jan 24, 2019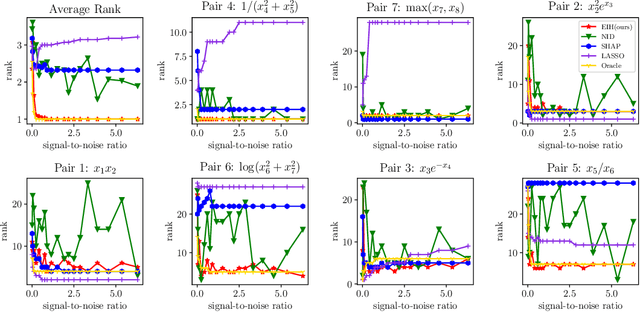
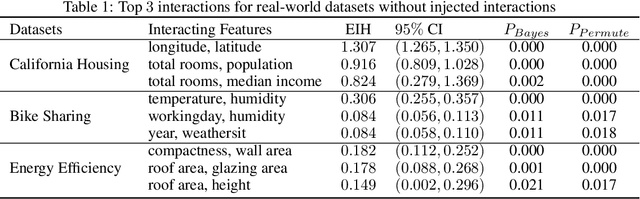
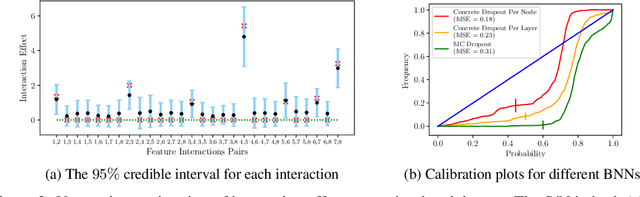
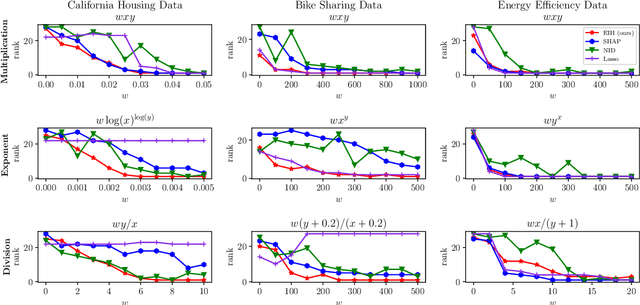
Abstract:Recovering pairwise interactions, i.e. pairs of input features whose joint effect on an output is different from the sum of their marginal effects, is central in many scientific applications. We conceptualize a solution to this problem as a two-stage procedure: first, we model the relationship between the features and the output using a flexible hybrid neural network; second, we detect feature interactions from the trained model. For the second step we propose a simple and intuitive interaction measure (IM), which has no specific requirements on the machine learning model used in the first step, only that it defines a mapping from an input to an output. And in a special case it reduces to the averaged Hessian of the input-output mapping. Importantly, our method upper bounds the interaction recovery error with the error of the learning model, which ensures that we can improve the recovered interactions by training a more accurate model. We present analyses of simulated and real-world data which demonstrate the benefits of our method compared to available alternatives, and theoretically analyse its properties and relation to other methods.
Neural Non-Stationary Spectral Kernel
Nov 27, 2018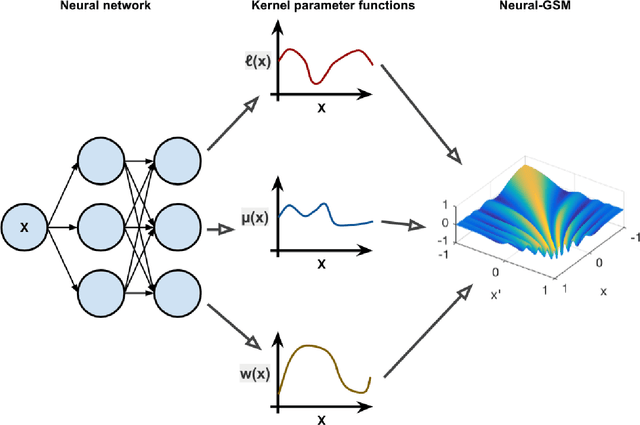
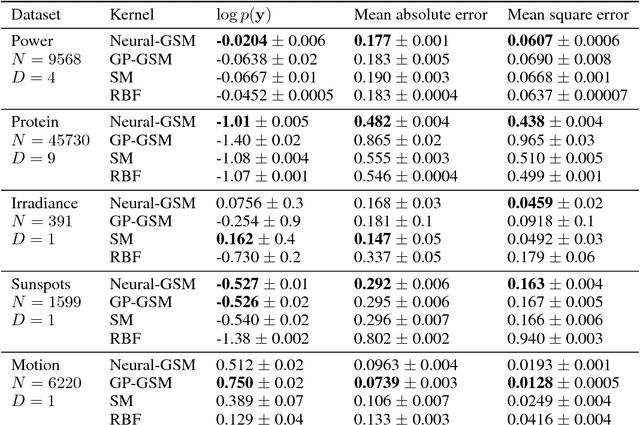
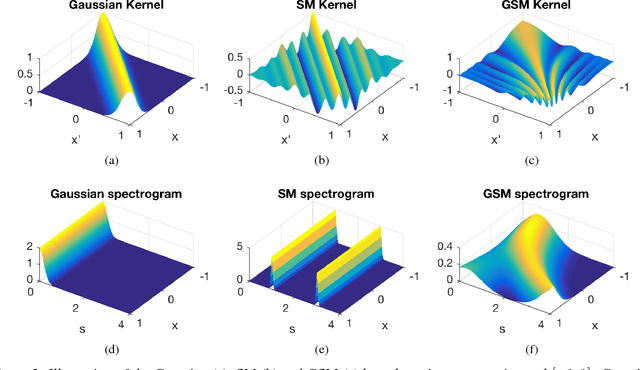
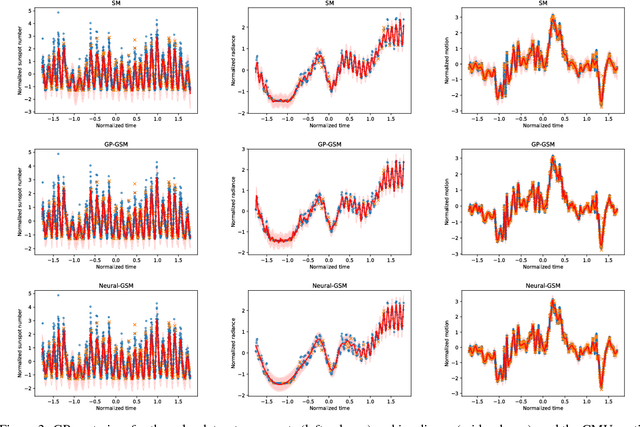
Abstract:Standard kernels such as Mat\'ern or RBF kernels only encode simple monotonic dependencies within the input space. Spectral mixture kernels have been proposed as general-purpose, flexible kernels for learning and discovering more complicated patterns in the data. Spectral mixture kernels have recently been generalized into non-stationary kernels by replacing the mixture weights, frequency means and variances by input-dependent functions. These functions have also been modelled as Gaussian processes on their own. In this paper we propose modelling the hyperparameter functions with neural networks, and provide an experimental comparison between the stationary spectral mixture and the two non-stationary spectral mixtures. Scalable Gaussian process inference is implemented within the sparse variational framework for all the kernels considered. We show that the neural variant of the kernel is able to achieve the best performance, among alternatives, on several benchmark datasets.
Approximate Bayesian Computation via Population Monte Carlo and Classification
Oct 29, 2018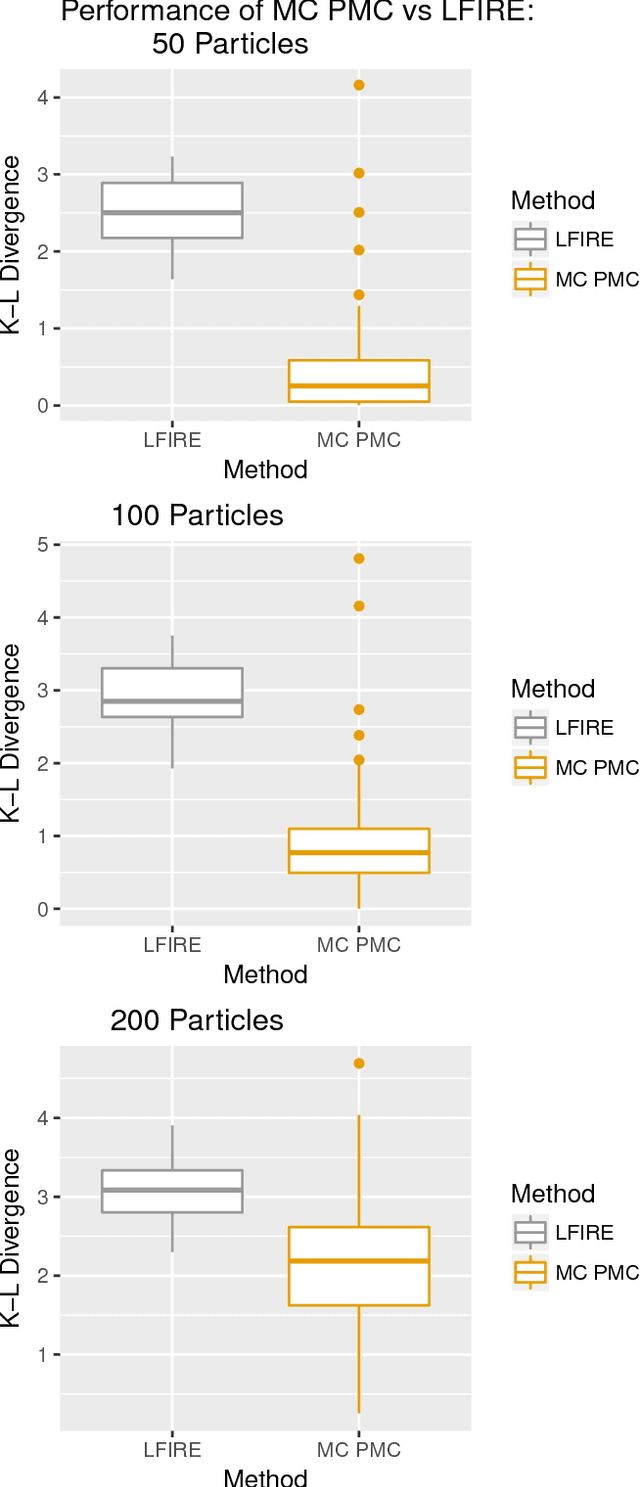
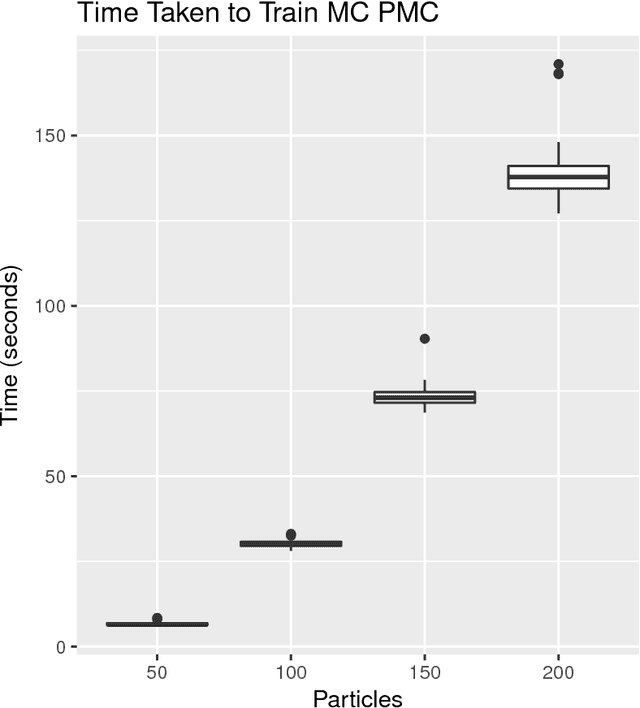
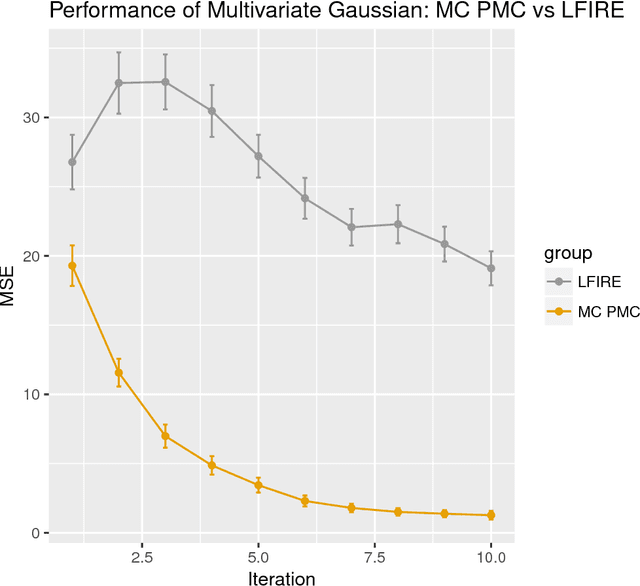
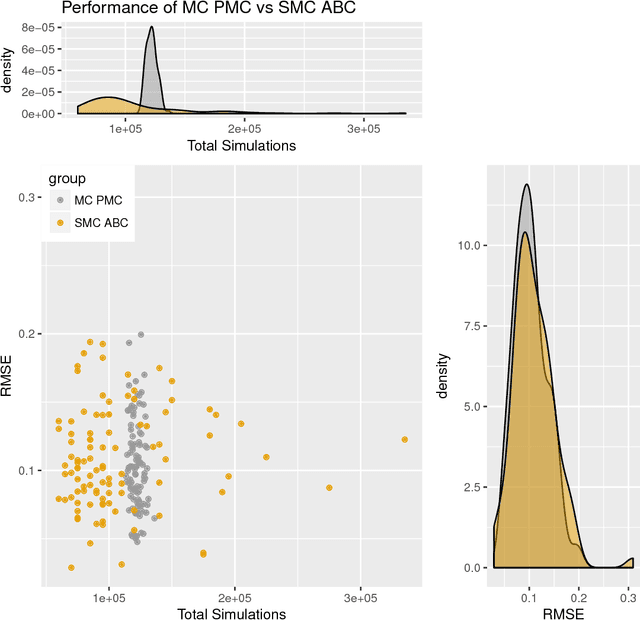
Abstract:Approximate Bayesian computation (ABC) methods can be used to sample from posterior distributions when the likelihood function is unavailable or intractable, as is often the case in biological systems. Sequential Monte Carlo (SMC) methods have been combined with ABC to improve efficiency, however these approaches require many simulations from the likelihood. We propose a classification approach within a population Monte Carlo (PMC) framework, where model class probabilities are used to update the particle weights. Our proposed approach outperforms state-of-the-art ratio estimation methods while retaining the automatic selection of summary statistics, and performs competitively with SMC ABC.
Deep learning with differential Gaussian process flows
Oct 15, 2018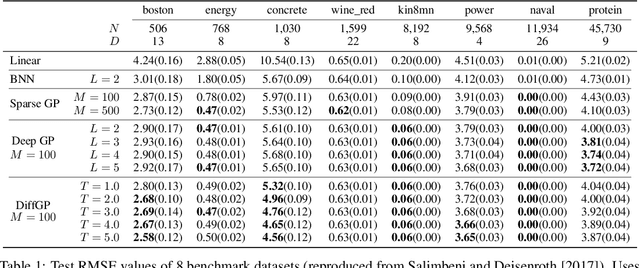
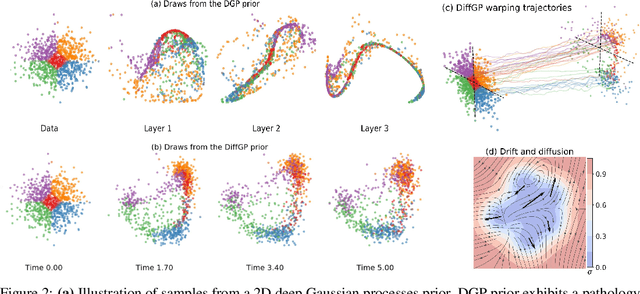
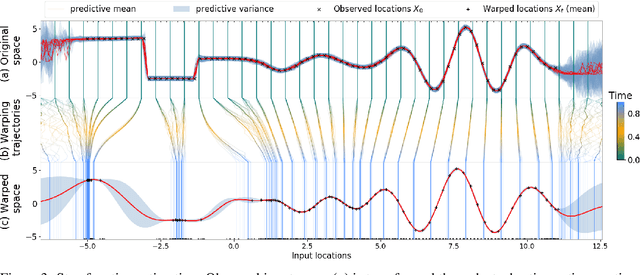
Abstract:We propose a novel deep learning paradigm of differential flows that learn a stochastic differential equation transformations of inputs prior to a standard classification or regression function. The key property of differential Gaussian processes is the warping of inputs through infinitely deep, but infinitesimal, differential fields, that generalise discrete layers into a dynamical system. We demonstrate state-of-the-art results that exceed the performance of deep Gaussian processes and neural networks
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge