Roland Kwitt
Comparing object recognition in humans and deep convolutional neural networks -- An eye tracking study
Jul 30, 2021
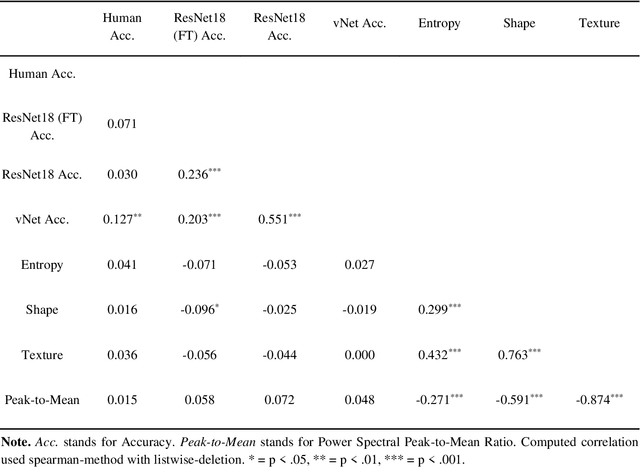
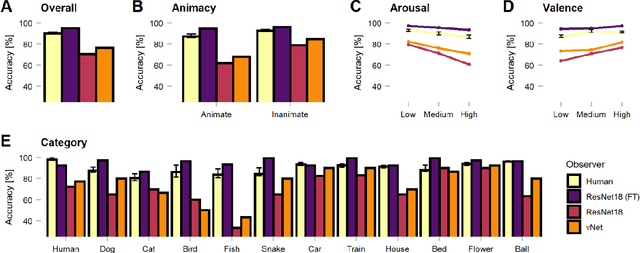
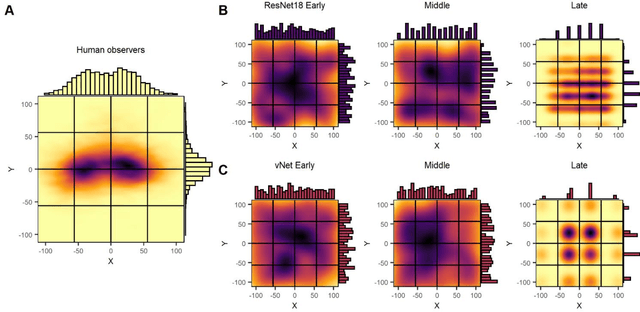
Abstract:Deep convolutional neural networks (DCNNs) and the ventral visual pathway share vast architectural and functional similarities in visual challenges such as object recognition. Recent insights have demonstrated that both hierarchical cascades can be compared in terms of both exerted behavior and underlying activation. However, these approaches ignore key differences in spatial priorities of information processing. In this proof-of-concept study, we demonstrate a comparison of human observers (N = 45) and three feedforward DCNNs through eye tracking and saliency maps. The results reveal fundamentally different resolutions in both visualization methods that need to be considered for an insightful comparison. Moreover, we provide evidence that a DCNN with biologically plausible receptive field sizes called vNet reveals higher agreement with human viewing behavior as contrasted with a standard ResNet architecture. We find that image-specific factors such as category, animacy, arousal, and valence have a direct link to the agreement of spatial object recognition priorities in humans and DCNNs, while other measures such as difficulty and general image properties do not. With this approach, we try to open up new perspectives at the intersection of biological and computer vision research.
Topological Attention for Time Series Forecasting
Jul 19, 2021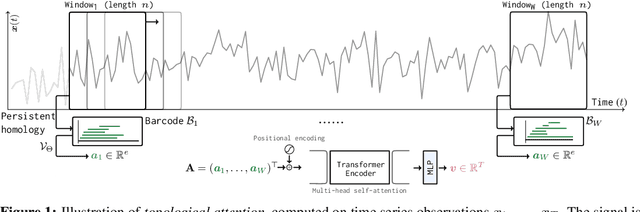
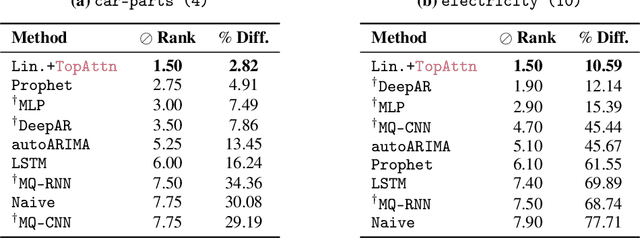
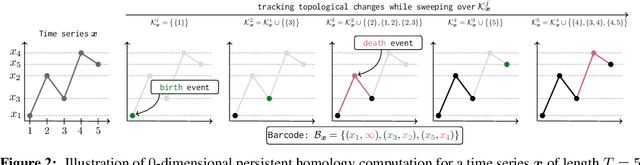
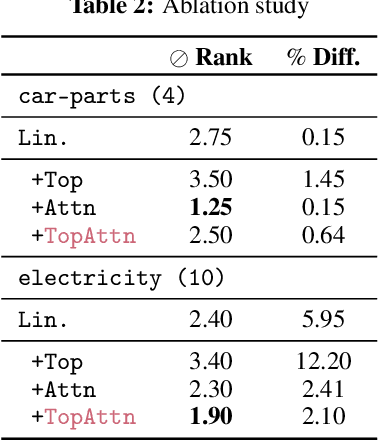
Abstract:The problem of (point) forecasting $ \textit{univariate} $ time series is considered. Most approaches, ranging from traditional statistical methods to recent learning-based techniques with neural networks, directly operate on raw time series observations. As an extension, we study whether $\textit{local topological properties}$, as captured via persistent homology, can serve as a reliable signal that provides complementary information for learning to forecast. To this end, we propose $\textit{topological attention}$, which allows attending to local topological features within a time horizon of historical data. Our approach easily integrates into existing end-to-end trainable forecasting models, such as $\texttt{N-BEATS}$, and in combination with the latter exhibits state-of-the-art performance on the large-scale M4 benchmark dataset of 100,000 diverse time series from different domains. Ablation experiments, as well as a comparison to a broad range of forecasting methods in a setting where only a single time series is available for training, corroborate the beneficial nature of including local topological information through an attention mechanism.
ICON: Learning Regular Maps Through Inverse Consistency
May 21, 2021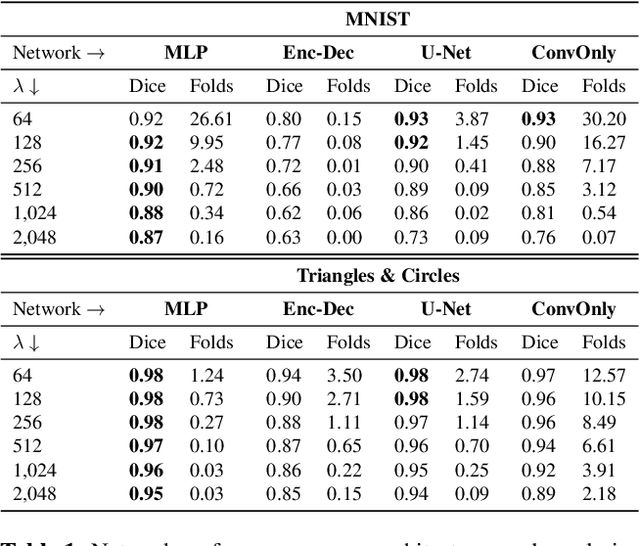
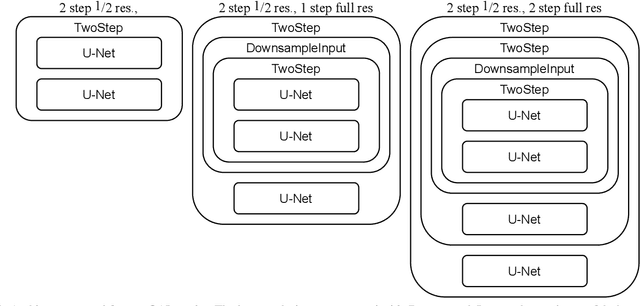
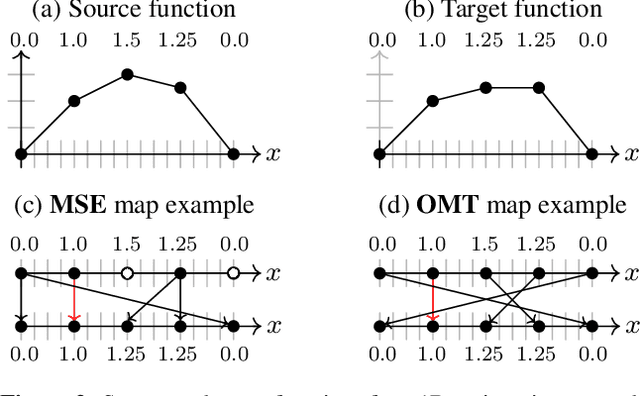
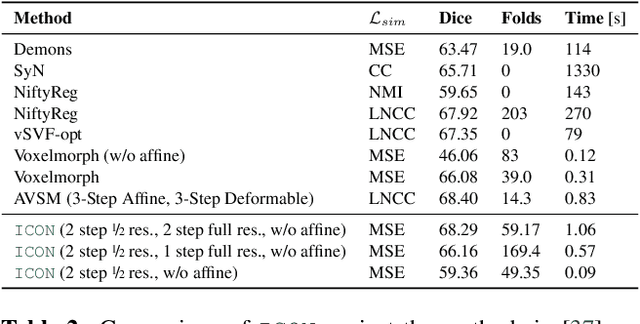
Abstract:Learning maps between data samples is fundamental. Applications range from representation learning, image translation and generative modeling, to the estimation of spatial deformations. Such maps relate feature vectors, or map between feature spaces. Well-behaved maps should be regular, which can be imposed explicitly or may emanate from the data itself. We explore what induces regularity for spatial transformations, e.g., when computing image registrations. Classical optimization-based models compute maps between pairs of samples and rely on an appropriate regularizer for well-posedness. Recent deep learning approaches have attempted to avoid using such regularizers altogether by relying on the sample population instead. We explore if it is possible to obtain spatial regularity using an inverse consistency loss only and elucidate what explains map regularity in such a context. We find that deep networks combined with an inverse consistency loss and randomized off-grid interpolation yield well behaved, approximately diffeomorphic, spatial transformations. Despite the simplicity of this approach, our experiments present compelling evidence, on both synthetic and real data, that regular maps can be obtained without carefully tuned explicit regularizers, while achieving competitive registration performance.
Sparse Pose Trajectory Completion
May 01, 2021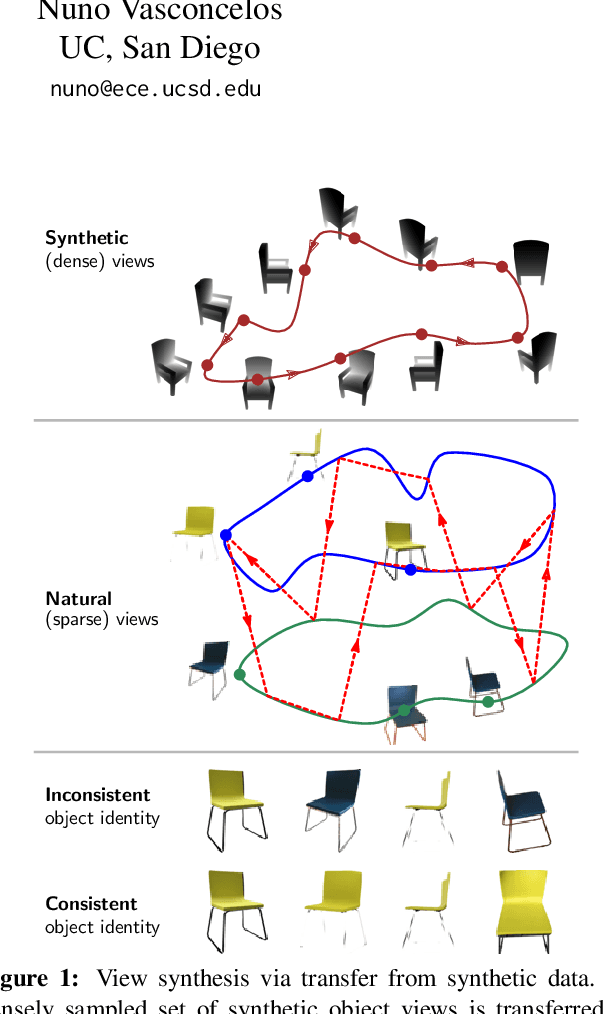
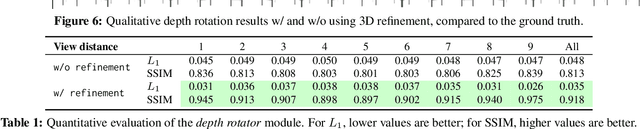
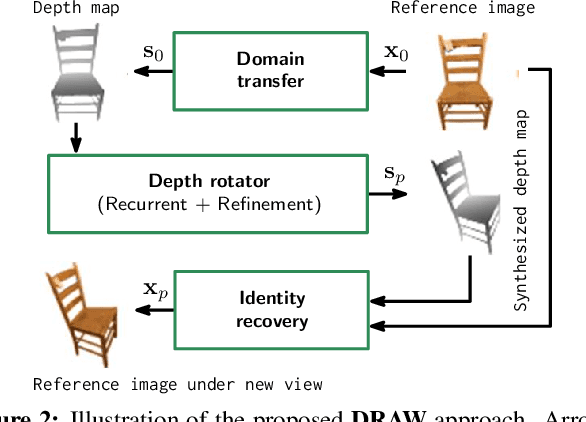
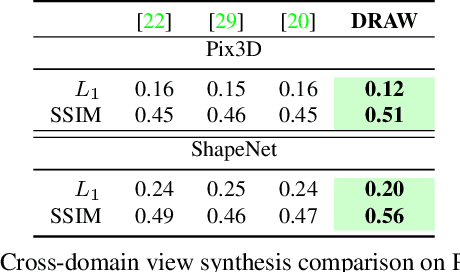
Abstract:We propose a method to learn, even using a dataset where objects appear only in sparsely sampled views (e.g. Pix3D), the ability to synthesize a pose trajectory for an arbitrary reference image. This is achieved with a cross-modal pose trajectory transfer mechanism. First, a domain transfer function is trained to predict, from an RGB image of the object, its 2D depth map. Then, a set of image views is generated by learning to simulate object rotation in the depth space. Finally, the generated poses are mapped from this latent space into a set of corresponding RGB images using a learned identity preserving transform. This results in a dense pose trajectory of the object in image space. For each object type (e.g., a specific Ikea chair model), a 3D CAD model is used to render a full pose trajectory of 2D depth maps. In the absence of dense pose sampling in image space, these latent space trajectories provide cross-modal guidance for learning. The learned pose trajectories can be transferred to unseen examples, effectively synthesizing all object views in image space. Our method is evaluated on the Pix3D and ShapeNet datasets, in the setting of novel view synthesis under sparse pose supervision, demonstrating substantial improvements over recent art.
Dissecting Supervised Constrastive Learning
Feb 17, 2021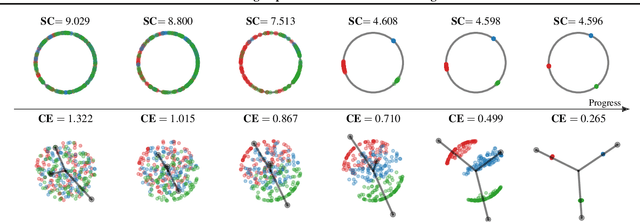
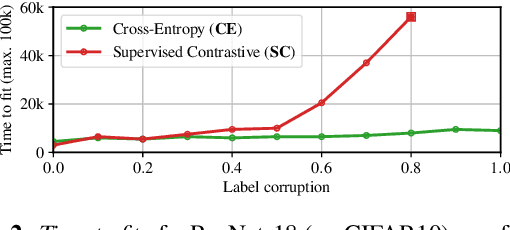
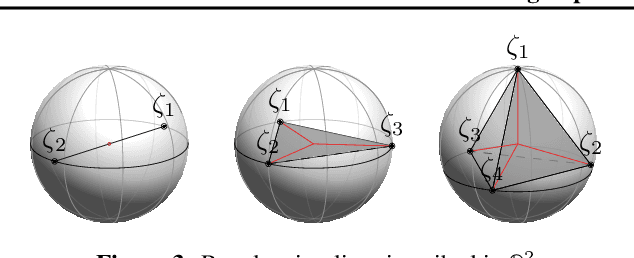
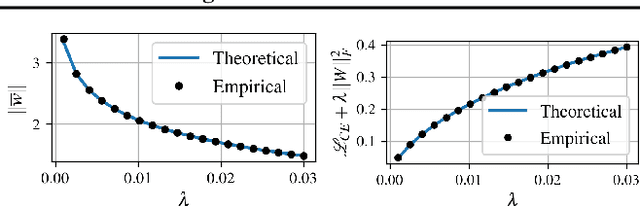
Abstract:Minimizing cross-entropy over the softmax scores of a linear map composed with a high-capacity encoder is arguably the most popular choice for training neural networks on supervised learning tasks. However, recent works show that one can directly optimize the encoder instead, to obtain equally (or even more) discriminative representations via a supervised variant of a contrastive objective. In this work, we address the question whether there are fundamental differences in the sought-for representation geometry in the output space of the encoder at minimal loss. Specifically, we prove, under mild assumptions, that both losses attain their minimum once the representations of each class collapse to the vertices of a regular simplex, inscribed in a hypersphere. We provide empirical evidence that this configuration is attained in practice and that reaching a close-to-optimal state typically indicates good generalization performance. Yet, the two losses show remarkably different optimization behavior. The number of iterations required to perfectly fit to data scales superlinearly with the amount of randomly flipped labels for the supervised contrastive loss. This is in contrast to the approximately linear scaling previously reported for networks trained with cross-entropy.
A Shooting Formulation of Deep Learning
Jun 18, 2020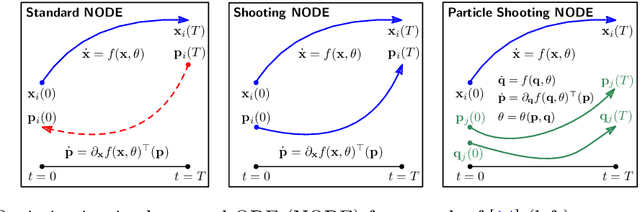
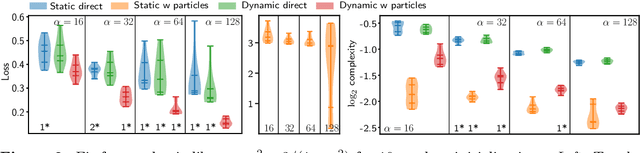
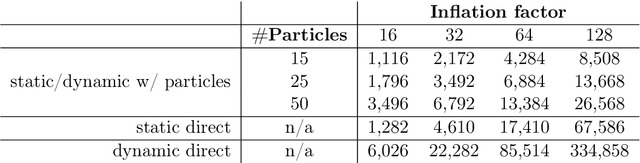
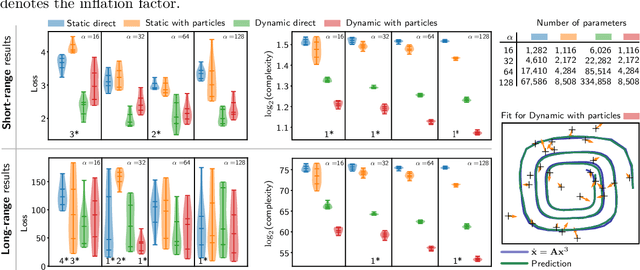
Abstract:Continuous-depth neural networks can be viewed as deep limits of discrete neural networks whose dynamics resemble a discretization of an ordinary differential equation (ODE). Although important steps have been taken to realize the advantages of such continuous formulations, most current techniques are not truly continuous-depth as they assume identical layers. Indeed, existing works throw into relief the myriad difficulties presented by an infinite-dimensional parameter space in learning a continuous-depth neural ODE. To this end, we introduce a shooting formulation which shifts the perspective from parameterizing a network layer-by-layer to parameterizing over optimal networks described only by a set of initial conditions. For scalability, we propose a novel particle-ensemble parametrization which fully specifies the optimal weight trajectory of the continuous-depth neural network. Our experiments show that our particle-ensemble shooting formulation can achieve competitive performance, especially on long-range forecasting tasks. Finally, though the current work is inspired by continuous-depth neural networks, the particle-ensemble shooting formulation also applies to discrete-time networks and may lead to a new fertile area of research in deep learning parametrization.
Topologically Densified Distributions
Feb 12, 2020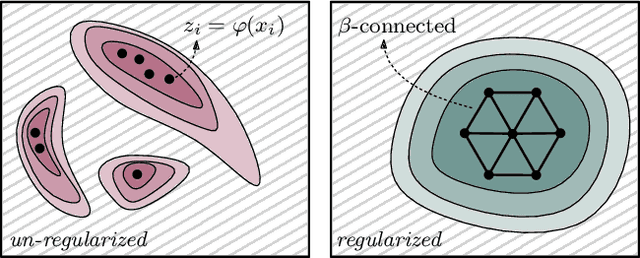
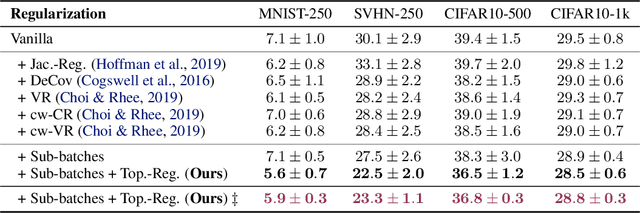
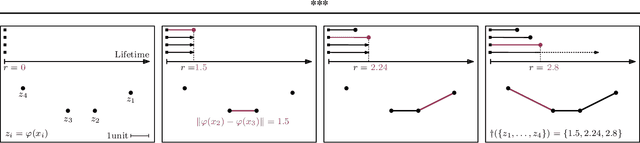
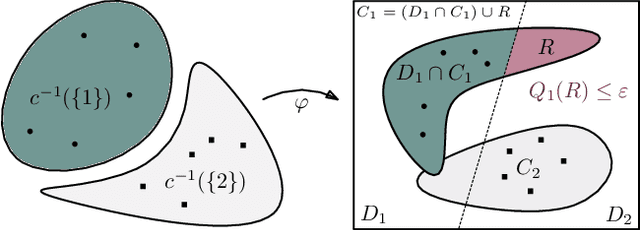
Abstract:We study regularization in the context of small sample-size learning with over-parameterized neural networks. Specifically, we shift focus from architectural properties, such as norms on the network weights, to properties of the internal representations before a linear classifier. Specifically, we impose a topological constraint on samples drawn from the probability measure induced in that space. This provably leads to mass concentration effects around the representations of training instances, i.e., a property beneficial for generalization. By leveraging previous work to impose topological constraints in a neural network setting, we provide empirical evidence (across various vision benchmarks) to support our claim for better generalization.
Deep Multi-View Learning via Task-Optimal CCA
Jul 17, 2019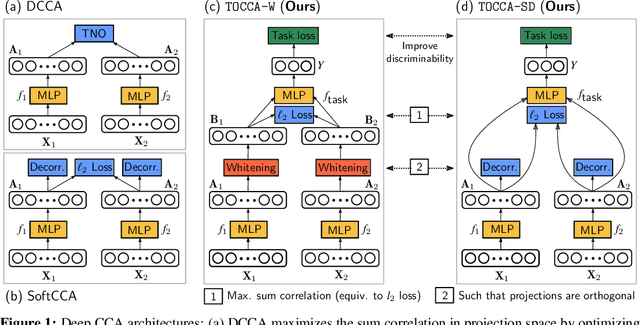
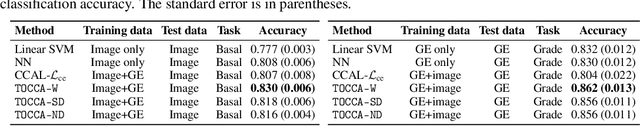
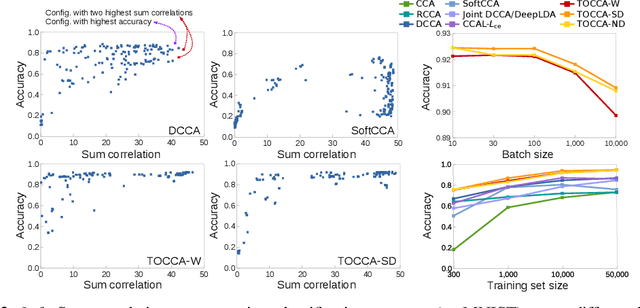
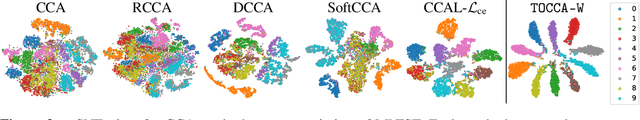
Abstract:Canonical Correlation Analysis (CCA) is widely used for multimodal data analysis and, more recently, for discriminative tasks such as multi-view learning; however, it makes no use of class labels. Recent CCA methods have started to address this weakness but are limited in that they do not simultaneously optimize the CCA projection for discrimination and the CCA projection itself, or they are linear only. We address these deficiencies by simultaneously optimizing a CCA-based and a task objective in an end-to-end manner. Together, these two objectives learn a non-linear CCA projection to a shared latent space that is highly correlated and discriminative. Our method shows a significant improvement over previous state-of-the-art (including deep supervised approaches) for cross-view classification, regularization with a second view, and semi-supervised learning on real data.
Connectivity-Optimized Representation Learning via Persistent Homology
Jun 21, 2019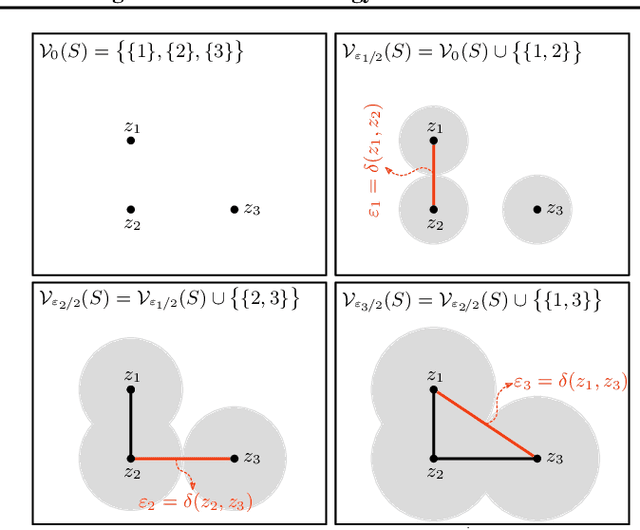
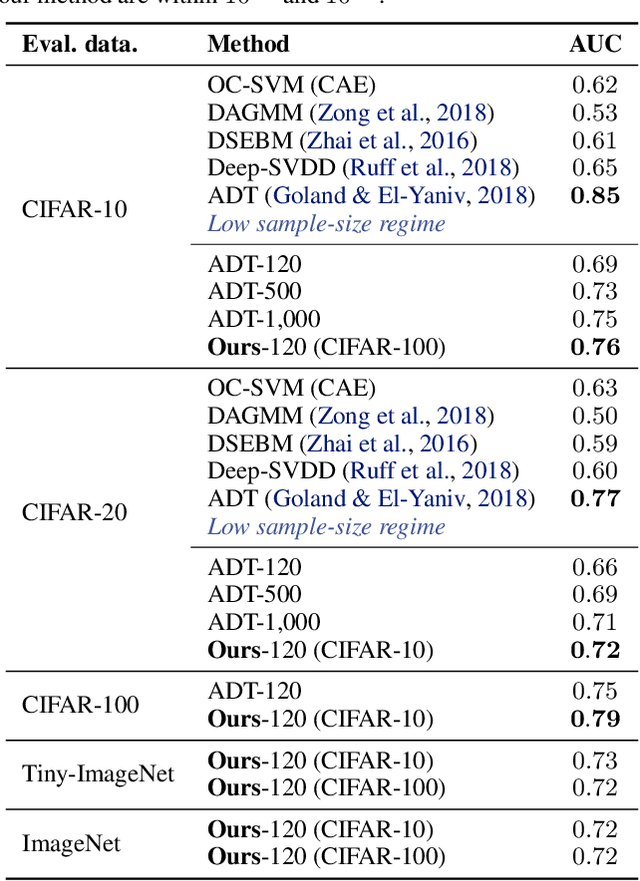
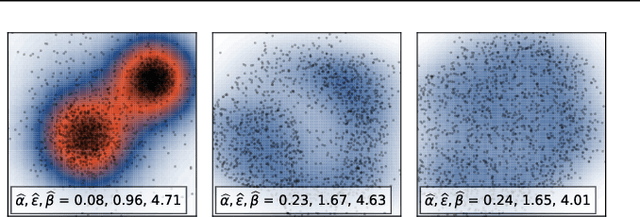
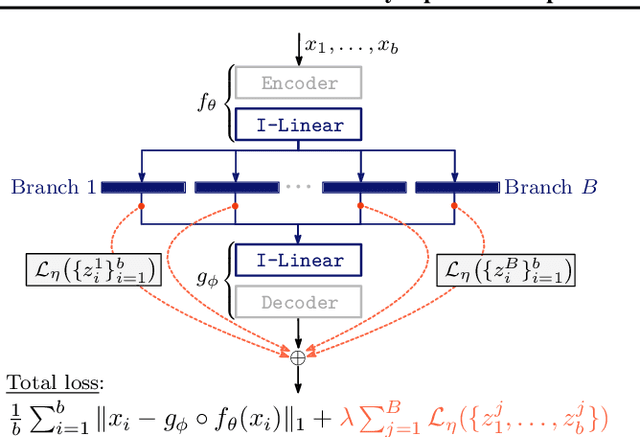
Abstract:We study the problem of learning representations with controllable connectivity properties. This is beneficial in situations when the imposed structure can be leveraged upstream. In particular, we control the connectivity of an autoencoder's latent space via a novel type of loss, operating on information from persistent homology. Under mild conditions, this loss is differentiable and we present a theoretical analysis of the properties induced by the loss. We choose one-class learning as our upstream task and demonstrate that the imposed structure enables informed parameter selection for modeling the in-class distribution via kernel density estimators. Evaluated on computer vision data, these one-class models exhibit competitive performance and, in a low sample size regime, outperform other methods by a large margin. Notably, our results indicate that a single autoencoder, trained on auxiliary (unlabeled) data, yields a mapping into latent space that can be reused across datasets for one-class learning.
Graph Filtration Learning
May 27, 2019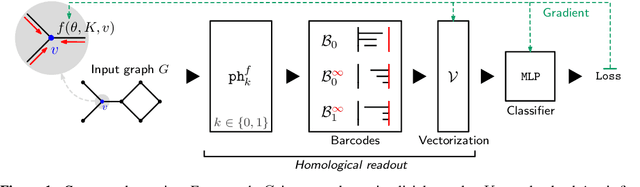
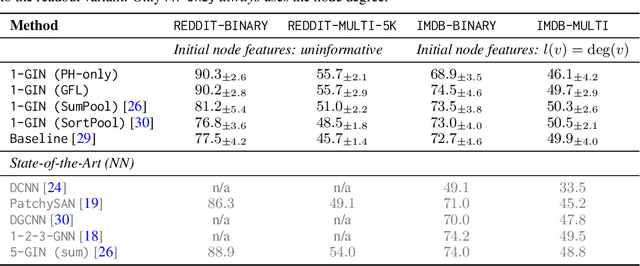
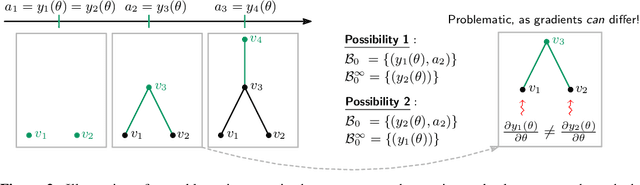
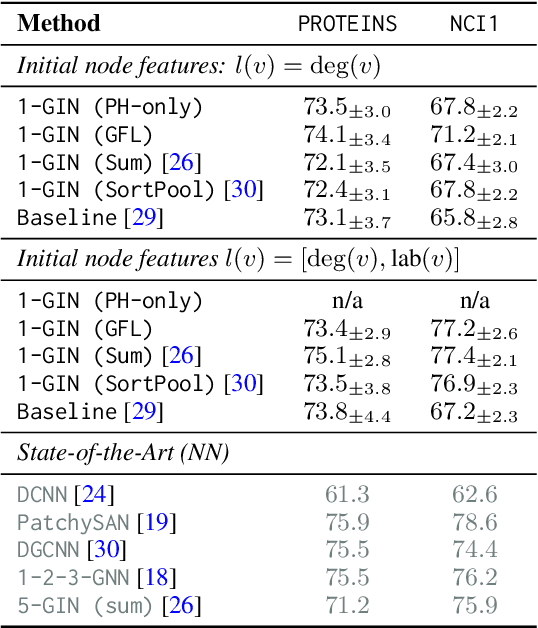
Abstract:We propose an approach to learning with graph-structured data in the problem domain of graph classification. In particular, we present a novel type of readout operation to aggregate node features into a graph-level representation. To this end, we leverage persistent homology computed via a real-valued, learnable, filter function. We establish the theoretical foundation for differentiating through the persistent homology computation. Empirically, we show that this type of readout operation compares favorably to previous techniques, especially when the graph connectivity structure is informative for the learning problem.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge