Regina Barzilay
Consistent Accelerated Inference via Confident Adaptive Transformers
Apr 18, 2021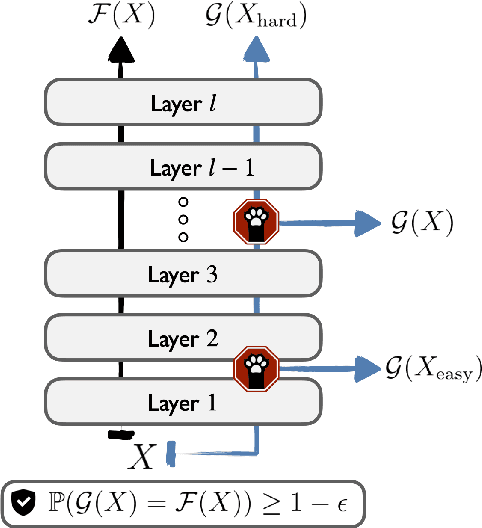
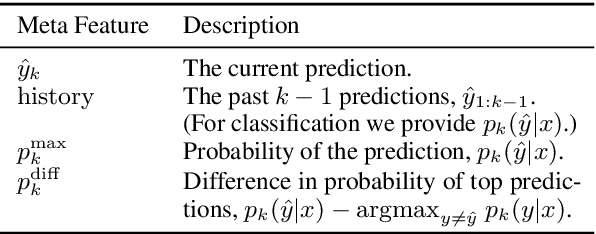

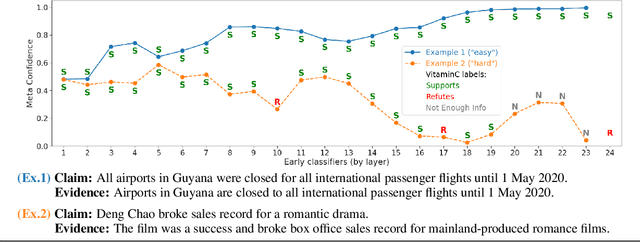
Abstract:We develop a novel approach for confidently accelerating inference in the large and expensive multilayer Transformers that are now ubiquitous in natural language processing (NLP). Amortized or approximate computational methods increase efficiency, but can come with unpredictable performance costs. In this work, we present CATs -- Confident Adaptive Transformers -- in which we simultaneously increase computational efficiency, while guaranteeing a specifiable degree of consistency with the original model with high confidence. Our method trains additional prediction heads on top of intermediate layers, and dynamically decides when to stop allocating computational effort to each input using a meta consistency classifier. To calibrate our early prediction stopping rule, we formulate a unique extension of conformal prediction. We demonstrate the effectiveness of this approach on four classification and regression tasks.
Generating Related Work
Apr 18, 2021


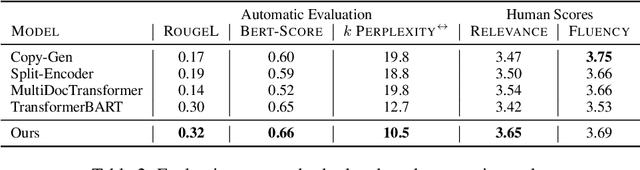
Abstract:Communicating new research ideas involves highlighting similarities and differences with past work. Authors write fluent, often long sections to survey the distinction of a new paper with related work. In this work we model generating related work sections while being cognisant of the motivation behind citing papers. Our content planning model generates a tree of cited papers before a surface realization model lexicalizes this skeleton. Our model outperforms several strong state-of-the-art summarization and multi-document summarization models on generating related work on an ACL Anthology (AA) based dataset which we contribute.
Nutribullets Hybrid: Multi-document Health Summarization
Apr 08, 2021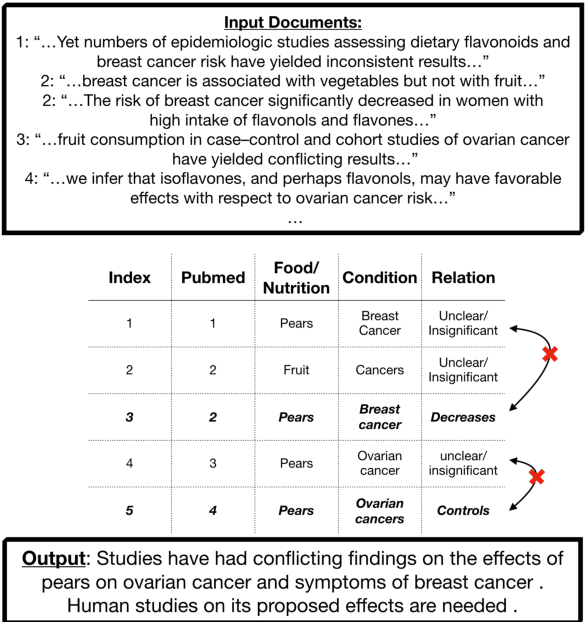
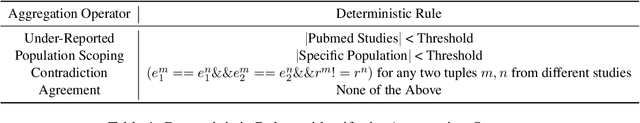

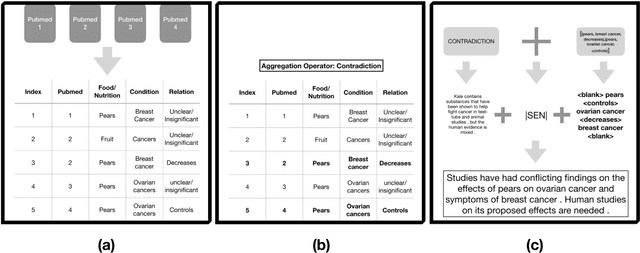

Abstract:We present a method for generating comparative summaries that highlights similarities and contradictions in input documents. The key challenge in creating such summaries is the lack of large parallel training data required for training typical summarization systems. To this end, we introduce a hybrid generation approach inspired by traditional concept-to-text systems. To enable accurate comparison between different sources, the model first learns to extract pertinent relations from input documents. The content planning component uses deterministic operators to aggregate these relations after identifying a subset for inclusion into a summary. The surface realization component lexicalizes this information using a text-infilling language model. By separately modeling content selection and realization, we can effectively train them with limited annotations. We implemented and tested the model in the domain of nutrition and health -- rife with inconsistencies. Compared to conventional methods, our framework leads to more faithful, relevant and aggregation-sensitive summarization -- while being equally fluent.
Nutri-bullets: Summarizing Health Studies by Composing Segments
Mar 22, 2021


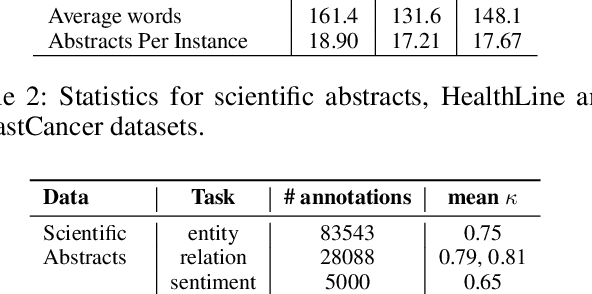
Abstract:We introduce \emph{Nutri-bullets}, a multi-document summarization task for health and nutrition. First, we present two datasets of food and health summaries from multiple scientific studies. Furthermore, we propose a novel \emph{extract-compose} model to solve the problem in the regime of limited parallel data. We explicitly select key spans from several abstracts using a policy network, followed by composing the selected spans to present a summary via a task specific language model. Compared to state-of-the-art methods, our approach leads to more faithful, relevant and diverse summarization -- properties imperative to this application. For instance, on the BreastCancer dataset our approach gets a more than 50\% improvement on relevance and faithfulness.\footnote{Our code and data is available at \url{https://github.com/darsh10/Nutribullets.}}
* 12 pages
Get Your Vitamin C! Robust Fact Verification with Contrastive Evidence
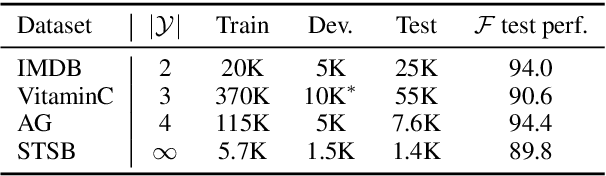
Mar 15, 2021Abstract:Typical fact verification models use retrieved written evidence to verify claims. Evidence sources, however, often change over time as more information is gathered and revised. In order to adapt, models must be sensitive to subtle differences in supporting evidence. We present VitaminC, a benchmark infused with challenging cases that require fact verification models to discern and adjust to slight factual changes. We collect over 100,000 Wikipedia revisions that modify an underlying fact, and leverage these revisions, together with additional synthetically constructed ones, to create a total of over 400,000 claim-evidence pairs. Unlike previous resources, the examples in VitaminC are contrastive, i.e., they contain evidence pairs that are nearly identical in language and content, with the exception that one supports a given claim while the other does not. We show that training using this design increases robustness -- improving accuracy by 10% on adversarial fact verification and 6% on adversarial natural language inference (NLI). Moreover, the structure of VitaminC leads us to define additional tasks for fact-checking resources: tagging relevant words in the evidence for verifying the claim, identifying factual revisions, and providing automatic edits via factually consistent text generation.
Few-shot Conformal Prediction with Auxiliary Tasks
Feb 17, 2021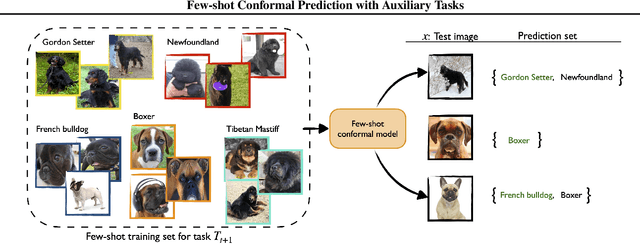
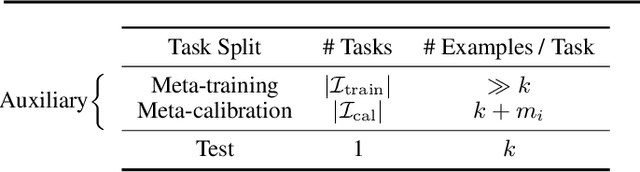
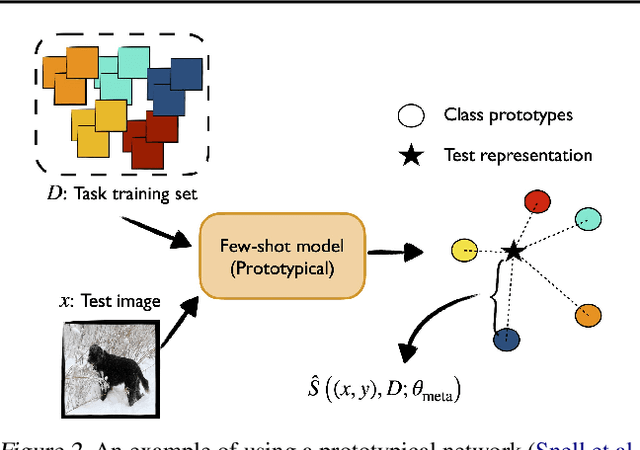
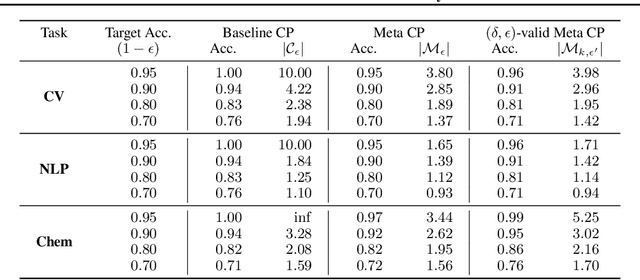
Abstract:We develop a novel approach to conformal prediction when the target task has limited data available for training. Conformal prediction identifies a small set of promising output candidates in place of a single prediction, with guarantees that the set contains the correct answer with high probability. When training data is limited, however, the predicted set can easily become unusably large. In this work, we obtain substantially tighter prediction sets while maintaining desirable marginal guarantees by casting conformal prediction as a meta-learning paradigm over exchangeable collections of auxiliary tasks. Our conformalization algorithm is simple, fast, and agnostic to the choice of underlying model, learning algorithm, or dataset. We demonstrate the effectiveness of this approach across a number of few-shot classification and regression tasks in natural language processing, computer vision, and computational chemistry for drug discovery.
CapWAP: Captioning with a Purpose
Nov 09, 2020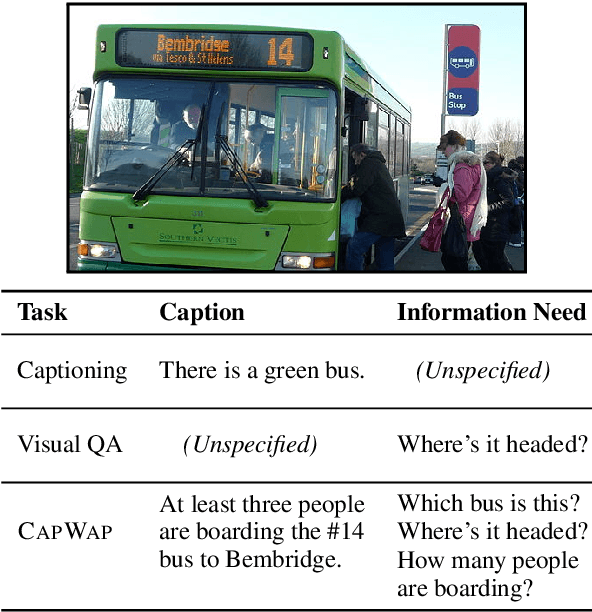
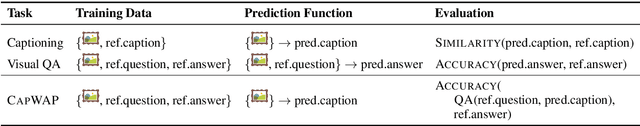
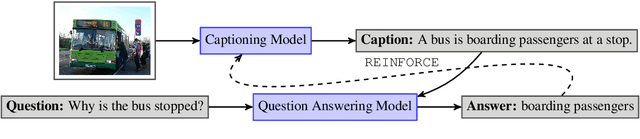

Abstract:The traditional image captioning task uses generic reference captions to provide textual information about images. Different user populations, however, will care about different visual aspects of images. In this paper, we propose a new task, Captioning with a Purpose (CapWAP). Our goal is to develop systems that can be tailored to be useful for the information needs of an intended population, rather than merely provide generic information about an image. In this task, we use question-answer (QA) pairs---a natural expression of information need---from users, instead of reference captions, for both training and post-inference evaluation. We show that it is possible to use reinforcement learning to directly optimize for the intended information need, by rewarding outputs that allow a question answering model to provide correct answers to sampled user questions. We convert several visual question answering datasets into CapWAP datasets, and demonstrate that under a variety of scenarios our purposeful captioning system learns to anticipate and fulfill specific information needs better than its generic counterparts, as measured by QA performance on user questions from unseen images, when using the caption alone as context.
Modeling Drug Combinations based on Molecular Structures and Biological Targets
Nov 09, 2020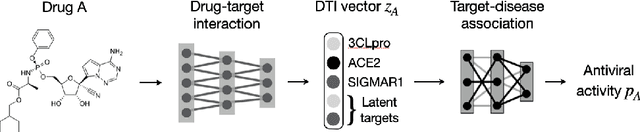
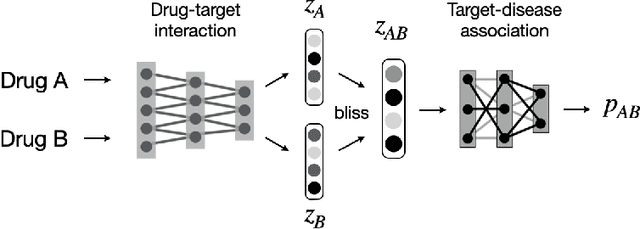
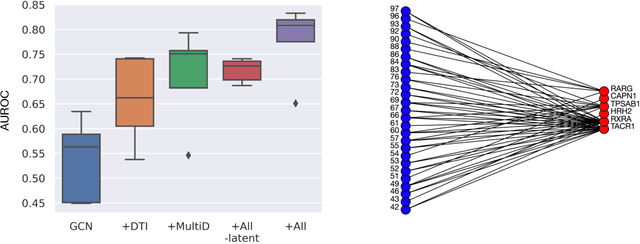
Abstract:Drug combinations play an important role in therapeutics due to its better efficacy and reduced toxicity. Since validating drug combinations via direct screening is prohibitively expensive due to combinatorial explosion, recent approaches have applied machine learning to identify synergistic combinations for cancer. However, these approaches is not readily applicable to many diseases with limited combination data. Motivated by the fact that drug synergy is closely tied with biological targets, we propose a model that jointly learns drug-target interaction and drug synergy. The model, parametrized as a graph convolutional network, consists of two parts: a drug-target interaction and target-disease association module. These modules are trained together on drug combination screen as well as abundant drug-target interaction data. Our model is trained and evaluated on two SARS-CoV-2 drug combination screens and achieves 0.777 test AUC, which is 10% higher than the model trained without drug-target interaction.
Deciphering Undersegmented Ancient Scripts Using Phonetic Prior
Oct 21, 2020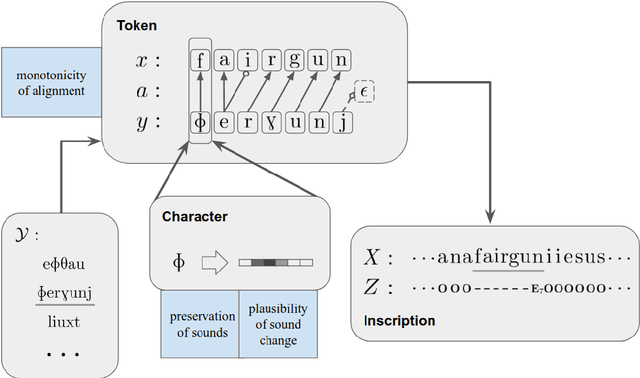

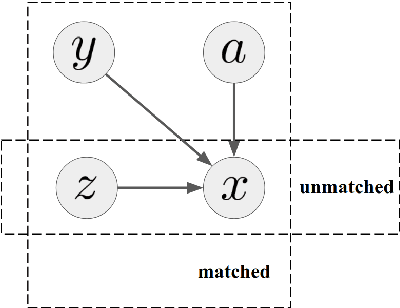
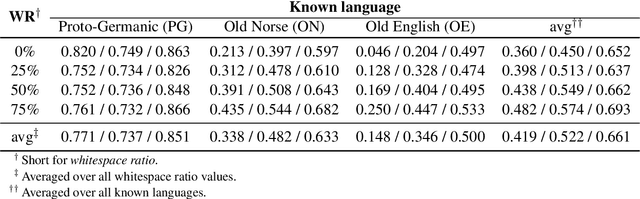
Abstract:Most undeciphered lost languages exhibit two characteristics that pose significant decipherment challenges: (1) the scripts are not fully segmented into words; (2) the closest known language is not determined. We propose a decipherment model that handles both of these challenges by building on rich linguistic constraints reflecting consistent patterns in historical sound change. We capture the natural phonological geometry by learning character embeddings based on the International Phonetic Alphabet (IPA). The resulting generative framework jointly models word segmentation and cognate alignment, informed by phonological constraints. We evaluate the model on both deciphered languages (Gothic, Ugaritic) and an undeciphered one (Iberian). The experiments show that incorporating phonetic geometry leads to clear and consistent gains. Additionally, we propose a measure for language closeness which correctly identifies related languages for Gothic and Ugaritic. For Iberian, the method does not show strong evidence supporting Basque as a related language, concurring with the favored position by the current scholarship.
Relaxed Conformal Prediction Cascades for Efficient Inference Over Many Labels
Jul 08, 2020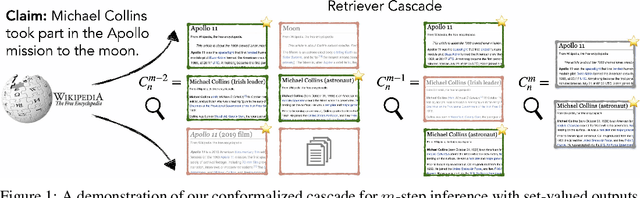
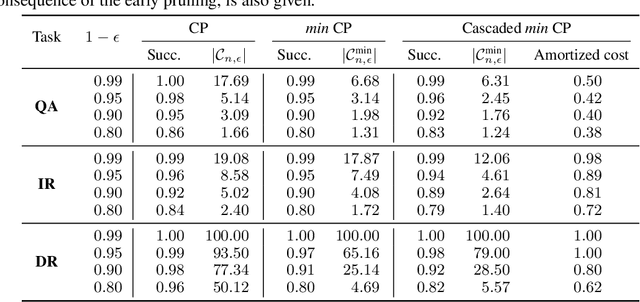
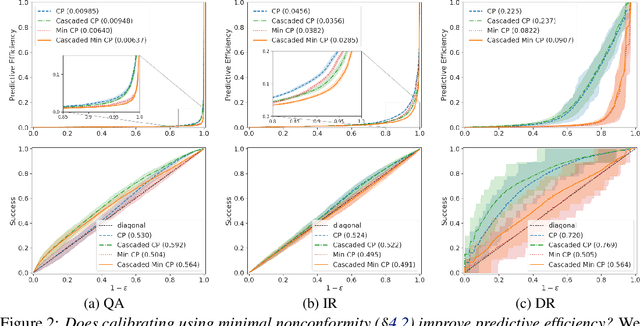
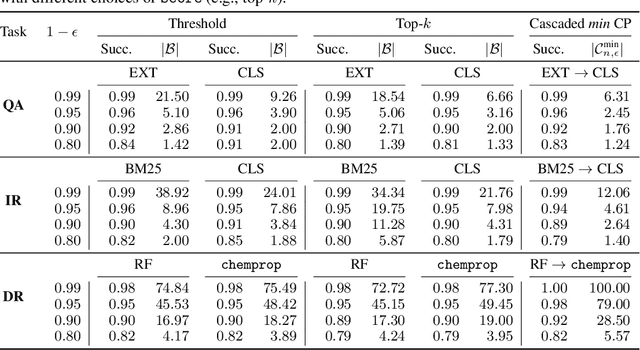
Abstract:Providing a small set of promising candidates in place of a single prediction is well-suited for many open-ended classification tasks. Conformal Prediction (CP) is a technique for creating classifiers that produce a valid set of predictions that contains the true answer with arbitrarily high probability. In practice, however, standard CP can suffer from both low predictive and computational efficiency during inference---i.e., the predicted set is both unusably large, and costly to obtain. This is particularly pervasive in the considered setting, where the correct answer is not unique and the number of total possible answers is high. In this work, we develop two simple and complementary techniques for improving both types of efficiencies. First, we relax CP validity to arbitrary criterions of success---allowing our framework to make more efficient predictions while remaining "equivalently correct." Second, we amortize cost by conformalizing prediction cascades, in which we aggressively prune implausible labels early on by using progressively stronger classifiers---while still guaranteeing marginal coverage. We demonstrate the empirical effectiveness of our approach for multiple applications in natural language processing and computational chemistry for drug discovery.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge