Raphael Meier
Enhancing clinical MRI Perfusion maps with data-driven maps of complementary nature for lesion outcome prediction
Jun 12, 2018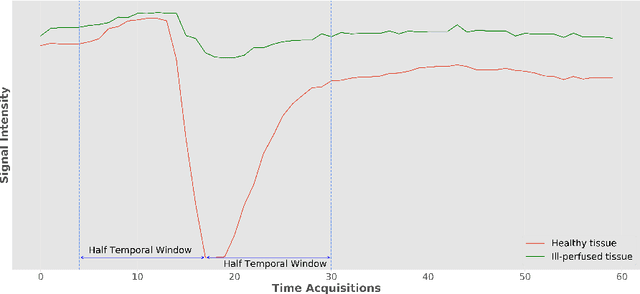
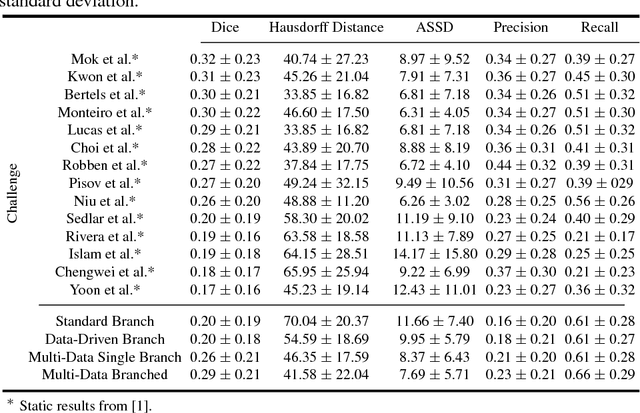
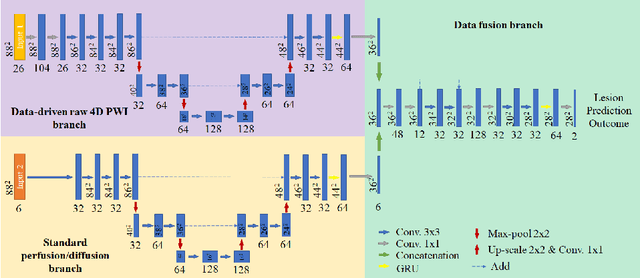
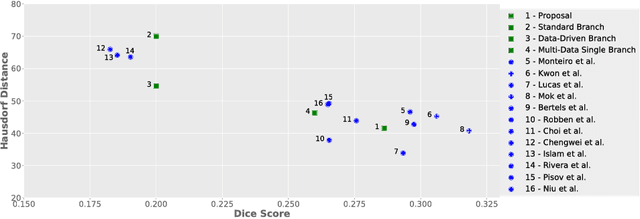
Abstract:Stroke is the second most common cause of death in developed countries, where rapid clinical intervention can have a major impact on a patient's life. To perform the revascularization procedure, the decision making of physicians considers its risks and benefits based on multi-modal MRI and clinical experience. Therefore, automatic prediction of the ischemic stroke lesion outcome has the potential to assist the physician towards a better stroke assessment and information about tissue outcome. Typically, automatic methods consider the information of the standard kinetic models of diffusion and perfusion MRI (e.g. Tmax, TTP, MTT, rCBF, rCBV) to perform lesion outcome prediction. In this work, we propose a deep learning method to fuse this information with an automated data selection of the raw 4D PWI image information, followed by a data-driven deep-learning modeling of the underlying blood flow hemodynamics. We demonstrate the ability of the proposed approach to improve prediction of tissue at risk before therapy, as compared to only using the standard clinical perfusion maps, hence suggesting on the potential benefits of the proposed data-driven raw perfusion data modelling approach.
Synthetic Perfusion Maps: Imaging Perfusion Deficits in DSC-MRI with Deep Learning
Jun 11, 2018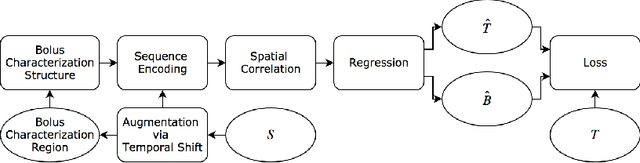
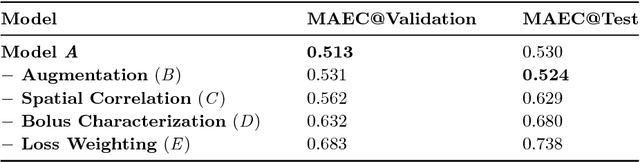
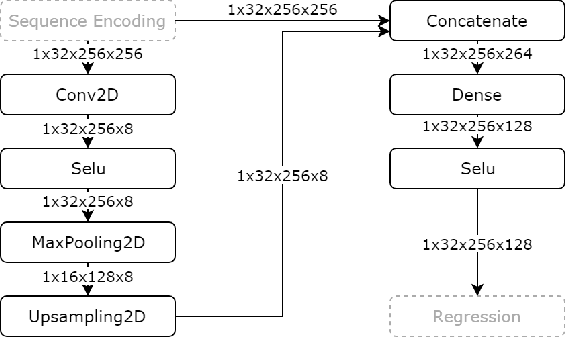
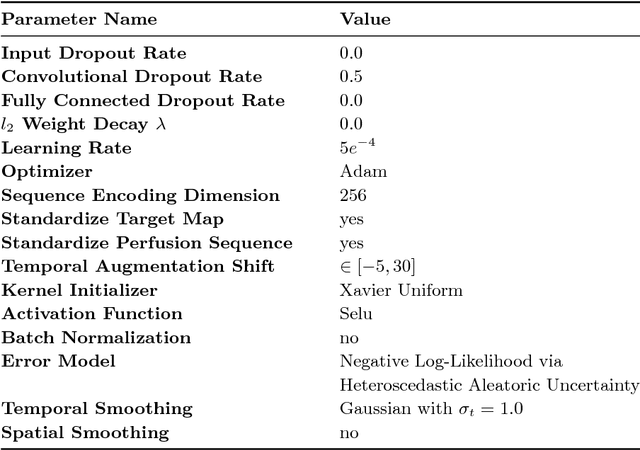
Abstract:In this work, we present a novel convolutional neural net- work based method for perfusion map generation in dynamic suscepti- bility contrast-enhanced perfusion imaging. The proposed architecture is trained end-to-end and solely relies on raw perfusion data for inference. We used a dataset of 151 acute ischemic stroke cases for evaluation. Our method generates perfusion maps that are comparable to the target maps used for clinical routine, while being model-free, fast, and less noisy.
Uncertainty-driven Sanity Check: Application to Postoperative Brain Tumor Cavity Segmentation
Jun 08, 2018
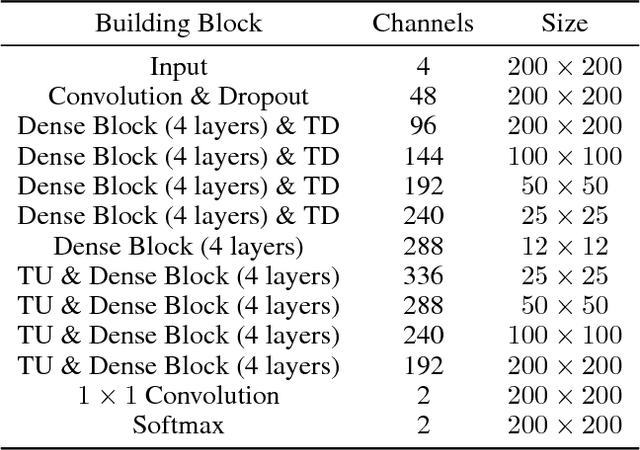
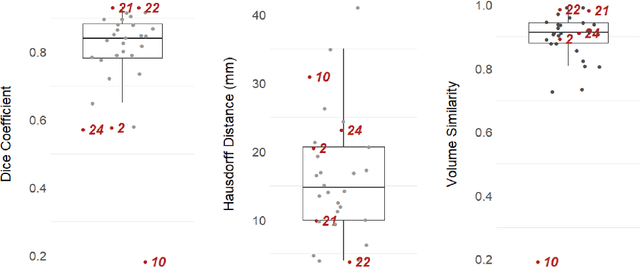
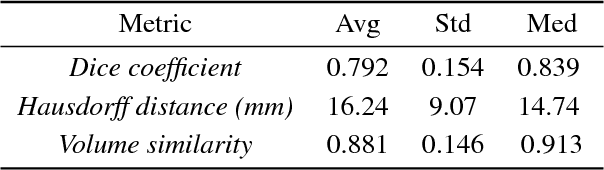
Abstract:Uncertainty estimates of modern neuronal networks provide additional information next to the computed predictions and are thus expected to improve the understanding of the underlying model. Reliable uncertainties are particularly interesting for safety-critical computer-assisted applications in medicine, e.g., neurosurgical interventions and radiotherapy planning. We propose an uncertainty-driven sanity check for the identification of segmentation results that need particular expert review. Our method uses a fully-convolutional neural network and computes uncertainty estimates by the principle of Monte Carlo dropout. We evaluate the performance of the proposed method on a clinical dataset with 30 postoperative brain tumor images. The method can segment the highly inhomogeneous resection cavities accurately (Dice coefficients 0.792 $\pm$ 0.154). Furthermore, the proposed sanity check is able to detect the worst segmentation and three out of the four outliers. The results highlight the potential of using the additional information from the model's parameter uncertainty to validate the segmentation performance of a deep learning model.
On the Effect of Inter-observer Variability for a Reliable Estimation of Uncertainty of Medical Image Segmentation
Jun 07, 2018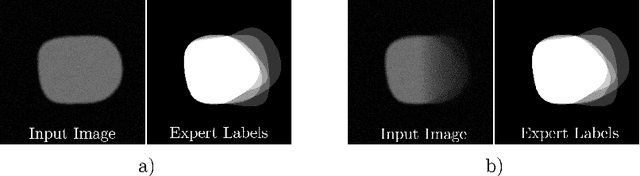
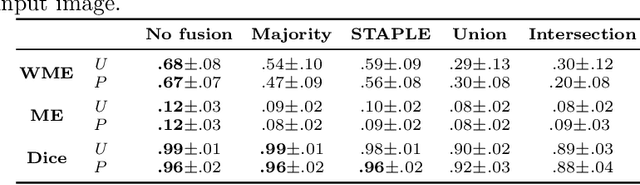
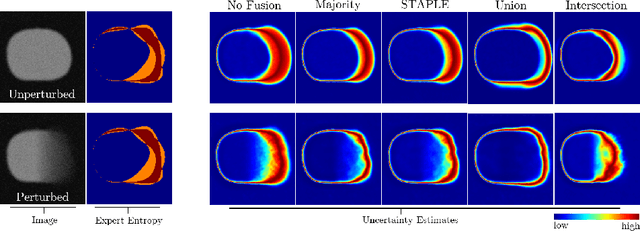
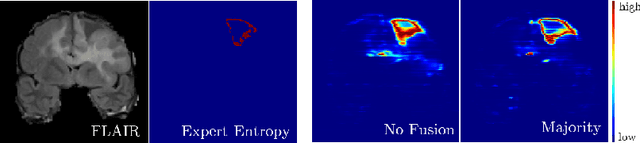
Abstract:Uncertainty estimation methods are expected to improve the understanding and quality of computer-assisted methods used in medical applications (e.g., neurosurgical interventions, radiotherapy planning), where automated medical image segmentation is crucial. In supervised machine learning, a common practice to generate ground truth label data is to merge observer annotations. However, as many medical image tasks show a high inter-observer variability resulting from factors such as image quality, different levels of user expertise and domain knowledge, little is known as to how inter-observer variability and commonly used fusion methods affect the estimation of uncertainty of automated image segmentation. In this paper we analyze the effect of common image label fusion techniques on uncertainty estimation, and propose to learn the uncertainty among observers. The results highlight the negative effect of fusion methods applied in deep learning, to obtain reliable estimates of segmentation uncertainty. Additionally, we show that the learned observers' uncertainty can be combined with current standard Monte Carlo dropout Bayesian neural networks to characterize uncertainty of model's parameters.
Perturb-and-MPM: Quantifying Segmentation Uncertainty in Dense Multi-Label CRFs
Mar 02, 2017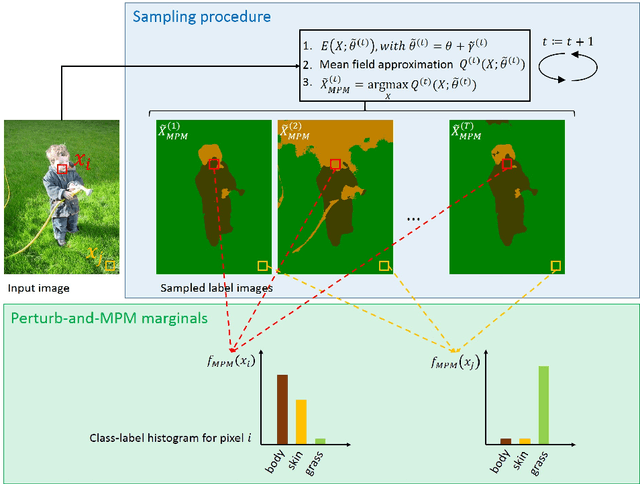
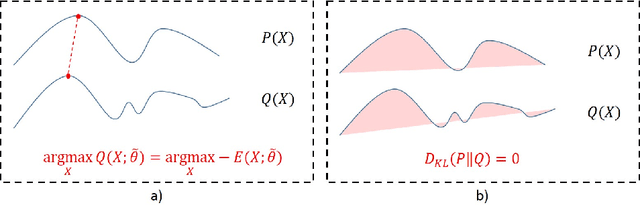
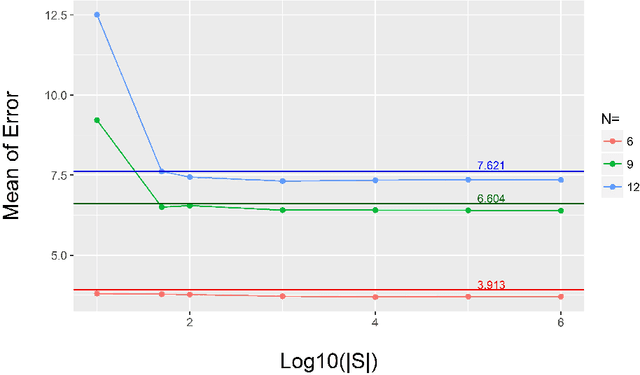
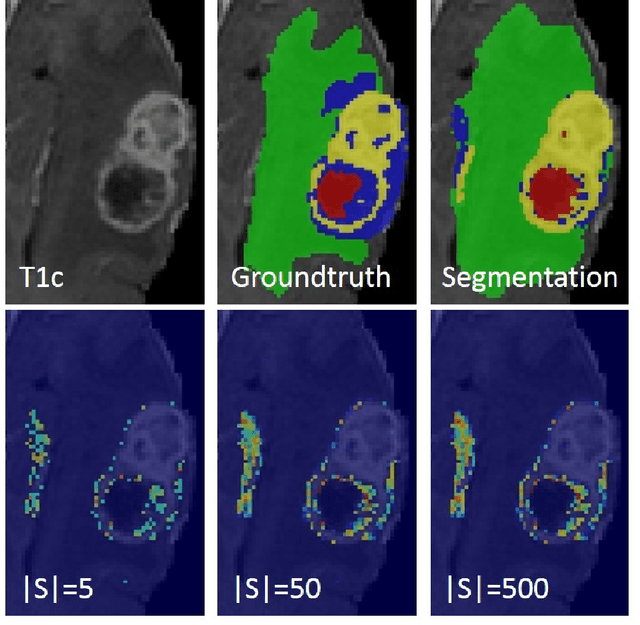
Abstract:This paper proposes a novel approach for uncertainty quantification in dense Conditional Random Fields (CRFs). The presented approach, called Perturb-and-MPM, enables efficient, approximate sampling from dense multi-label CRFs via random perturbations. An analytic error analysis was performed which identified the main cause of approximation error as well as showed that the error is bounded. Spatial uncertainty maps can be derived from the Perturb-and-MPM model, which can be used to visualize uncertainty in image segmentation results. The method is validated on synthetic and clinical Magnetic Resonance Imaging data. The effectiveness of the approach is demonstrated on the challenging problem of segmenting the tumor core in glioblastoma. We found that areas of high uncertainty correspond well to wrongly segmented image regions. Furthermore, we demonstrate the potential use of uncertainty maps to refine imaging biomarkers in the case of extent of resection and residual tumor volume in brain tumor patients.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge