Ram Srivatsav Ghorakavi
Dense Depth Estimation of a Complex Dynamic Scene without Explicit 3D Motion Estimation
Mar 23, 2019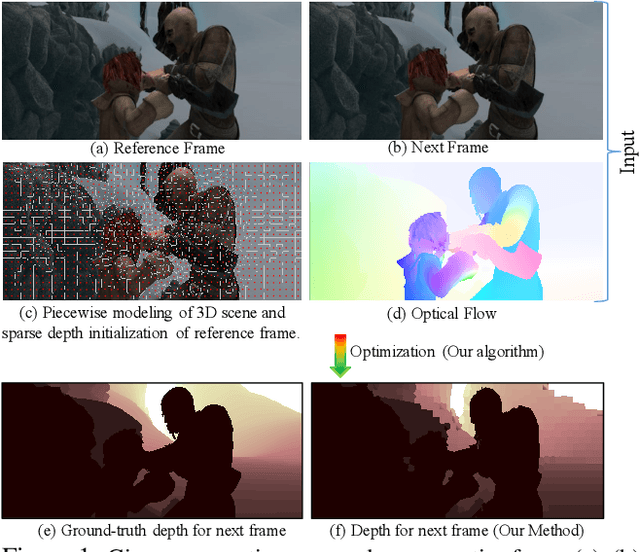
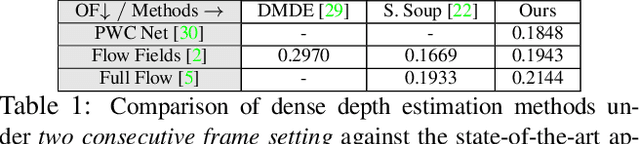
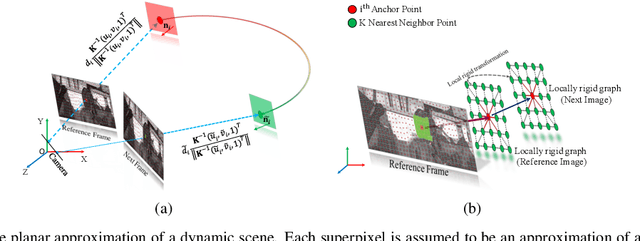
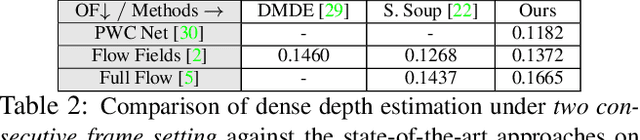
Abstract:Recent geometric methods need reliable estimates of 3D motion parameters to procure accurate dense depth map of a complex dynamic scene from monocular images \cite{kumar2017monocular, ranftl2016dense}. Generally, to estimate \textbf{precise} measurements of relative 3D motion parameters and to validate its accuracy using image data is a challenging task. In this work, we propose an alternative approach that circumvents the 3D motion estimation requirement to obtain a dense depth map of a dynamic scene. Given per-pixel optical flow correspondences between two consecutive frames and, the sparse depth prior for the reference frame, we show that, we can effectively recover the dense depth map for the successive frames without solving for 3D motion parameters. Our method assumes a piece-wise planar model of a dynamic scene, which undergoes rigid transformation locally, and as-rigid-as-possible transformation globally between two successive frames. Under our assumption, we can avoid the explicit estimation of 3D rotation and translation to estimate scene depth. In essence, our formulation provides an unconventional way to think and recover the dense depth map of a complex dynamic scene which is incremental and motion free in nature. Our proposed method does not make object level or any other high-level prior assumption about the dynamic scene, as a result, it is applicable to a wide range of scenarios. Experimental results on the benchmarks dataset show the competence of our approach for multiple frames.
TBNet:Pulmonary Tuberculosis Diagnosing System using Deep Neural Networks
Feb 24, 2019



Abstract:Tuberculosis is a deadly infectious disease prevalent around the world. Due to the lack of proper technology in place, the early detection of this disease is unattainable. Also, the available methods to detect Tuberculosis is not up-to a commendable standards due to their dependency on unnecessary features, this make such technology obsolete for a reliable health-care technology. In this paper, I propose a deep-learning based system which diagnoses tuberculosis based on the important features in Chest X-rays along with original chest X-rays. Employing our system will accelerate the process of tuberculosis diagnosis by overcoming the need to perform the time-consuming sputum-based testing method (Diagnostic Microbiology). In contrast to the previous methods \cite{kant2018towards, melendez2016automated}, our work utilizes the state-of-the-art ResNet \cite{he2016deep} with proper data augmentation using traditional robust features like Haar \cite{viola2005detecting,viola2001rapid} and LBP \cite{ojala1994performance,ojala1996comparative}. I observed that such a procedure enhances the rate of tuberculosis detection to a highly satisfactory level. Our work uses the publicly available pulmonary chest X-ray dataset to train our network \cite{jaeger2014two}. Nevertheless, the publicly available dataset is very small and is inadequate to achieve the best accuracy. To overcome this issue I have devised an intuitive feature based data augmentation pipeline. Our approach shall help the deep neural network \cite{lecun2015deep,he2016deep,krizhevsky2012imagenet} to focus its training on tuberculosis affected regions making it more robust and accurate, when compared to other conventional methods that use procedures like mirroring and rotation. By using our simple yet powerful techniques, I observed a 10\% boost in performance accuracy.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge