Rajesh Ranganath
New York University
Adversarial Examples for Electrocardiograms
Jun 04, 2019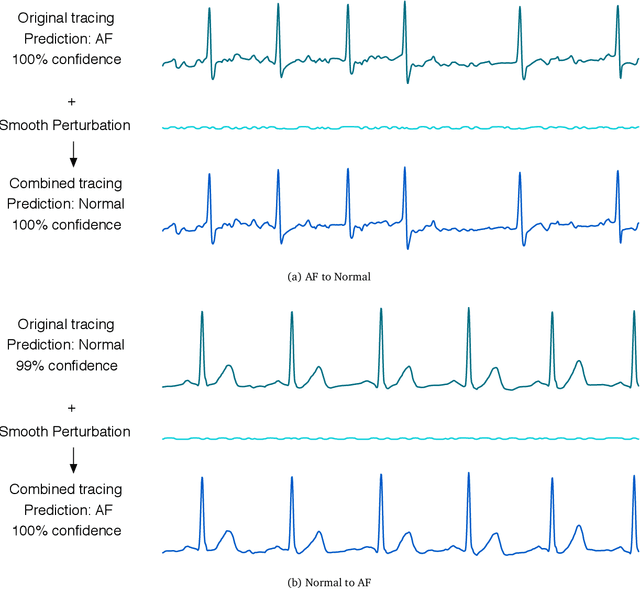
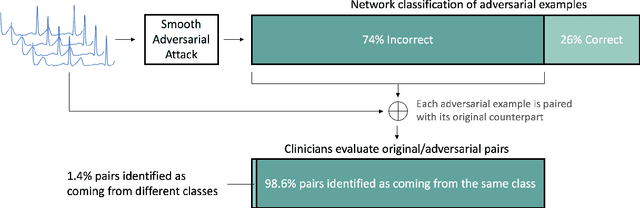
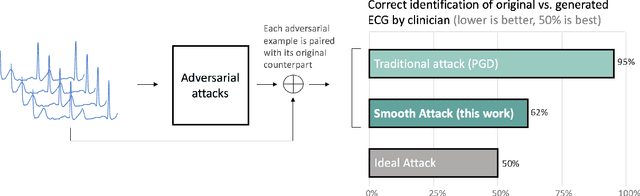
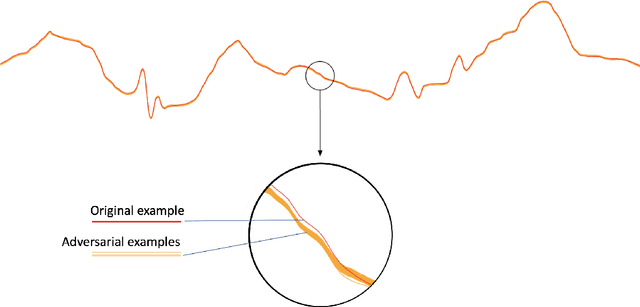
Abstract:In recent years, the electrocardiogram (ECG) has seen a large diffusion in both medical and commercial applications, fueled by the rise of single-lead versions. Single-lead ECG can be embedded in medical devices and wearable products such as the injectable Medtronic Linq monitor, the iRhythm Ziopatch wearable monitor, and the Apple Watch Series 4. Recently, deep neural networks have been used to automatically analyze ECG tracings, outperforming even physicians specialized in cardiac electrophysiology in detecting certain rhythm irregularities. However, deep learning classifiers have been shown to be brittle to adversarial examples, which are examples created to look incontrovertibly belonging to a certain class to a human eye but contain subtle features that fool the classifier into misclassifying them into the wrong class. Very recently, adversarial examples have also been created for medical-related tasks. Yet, traditional attack methods to create adversarial examples, such as projected gradient descent (PGD) do not extend directly to ECG signals, as they generate examples that introduce square wave artifacts that are not physiologically plausible. Here, we developed a method to construct smoothed adversarial examples for single-lead ECG. First, we implemented a neural network model achieving state-of-the-art performance on the data from the 2017 PhysioNet/Computing-in-Cardiology Challenge for arrhythmia detection from single lead ECG classification. For this model, we utilized a new technique to generate smoothed examples to produce signals that are 1) indistinguishable to cardiologists from the original examples and 2) incorrectly classified by the neural network. Finally, we show that adversarial examples are not unique and provide a general technique to collate and perturb known adversarial examples to create new ones.
Kernelized Complete Conditional Stein Discrepancy
Apr 13, 2019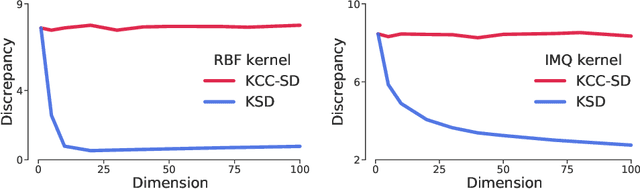
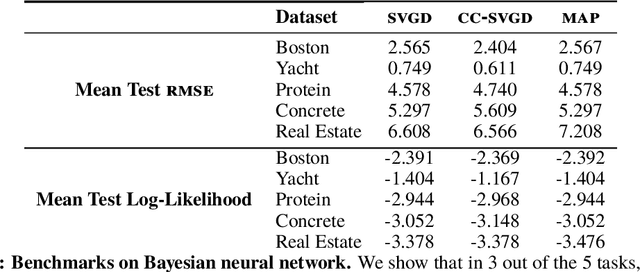
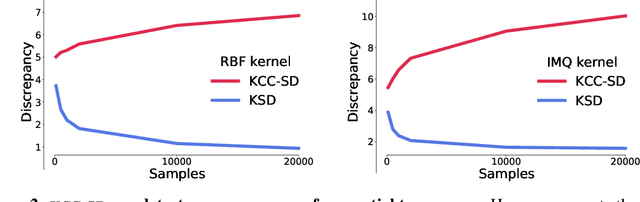
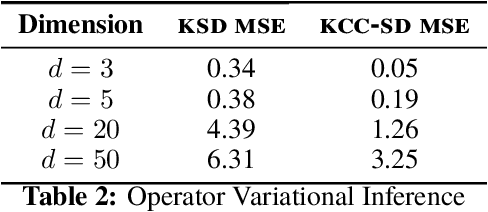
Abstract:Much of machine learning relies on comparing distributions with discrepancy measures. Stein's method creates discrepancy measures between two distributions that require only the unnormalized density of one and samples from the other. Stein discrepancies can be combined with kernels to define the kernelized Stein discrepancies (KSDs).While kernels make Stein discrepancies tractable, they pose several challenges in high dimensions. We introduce kernelized complete conditional Stein discrepancies (KCC-SDs). Complete conditionals turn a multivariate distribution into multiple univariate distributions. We prove that KCC-SDs detect convergence and non-convergence, and that they upper-bound KSDs. We empirically show that KCC-SDs detect non-convergence where KSDs fail. Our experiments illustrate the difference between KCC-SDs and KSDs when comparing high-dimensional distributions and performing variational inference.
ClinicalBERT: Modeling Clinical Notes and Predicting Hospital Readmission
Apr 11, 2019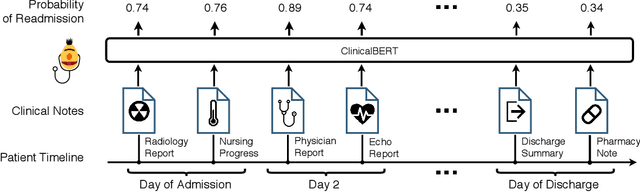
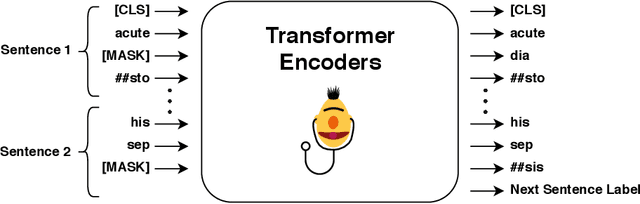



Abstract:Clinical notes contain information about patients that goes beyond structured data like lab values and medications. However, clinical notes have been underused relative to structured data, because notes are high-dimensional and sparse. This work develops and evaluates representations of clinical notes using bidirectional transformers (ClinicalBERT). ClinicalBERT uncovers high-quality relationships between medical concepts as judged by humans. ClinicalBert outperforms baselines on 30-day hospital readmission prediction using both discharge summaries and the first few days of notes in the intensive care unit. Code and model parameters are available.
The Random Conditional Distribution for Higher-Order Probabilistic Inference
Mar 25, 2019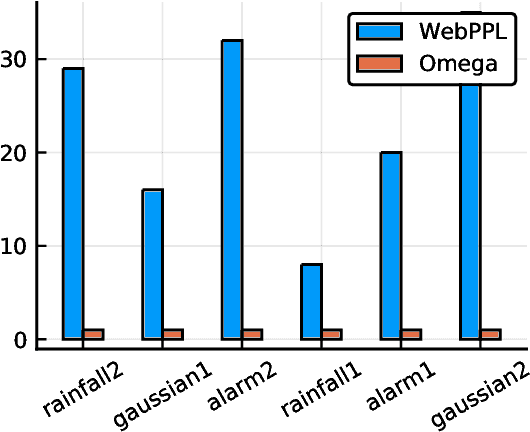


Abstract:The need to condition distributional properties such as expectation, variance, and entropy arises in algorithmic fairness, model simplification, robustness and many other areas. At face value however, distributional properties are not random variables, and hence conditioning them is a semantic error and type error in probabilistic programming languages. On the other hand, distributional properties are contingent on other variables in the model, change in value when we observe more information, and hence in a precise sense are random variables too. In order to capture the uncertain over distributional properties, we introduce a probability construct -- the random conditional distribution -- and incorporate it into a probabilistic programming language Omega. A random conditional distribution is a higher-order random variable whose realizations are themselves conditional random variables. In Omega we extend distributional properties of random variables to random conditional distributions, such that for example while the expectation a real valued random variable is a real value, the expectation of a random conditional distribution is a distribution over expectations. As a consequence, it requires minimal syntax to encode inference problems over distributional properties, which so far have evaded treatment within probabilistic programming systems and probabilistic modeling in general. We demonstrate our approach case studies in algorithmic fairness and robustness.
Support and Invertibility in Domain-Invariant Representations
Mar 21, 2019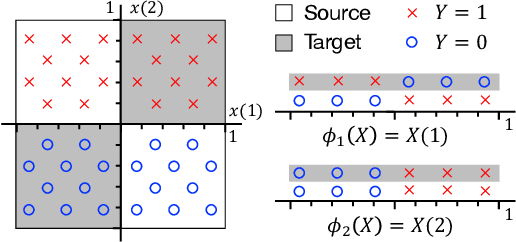
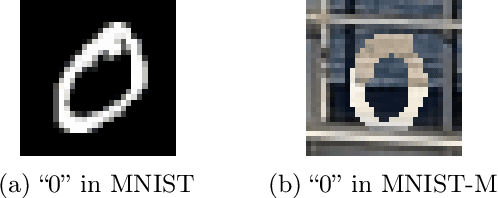
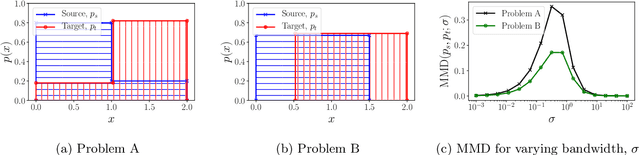

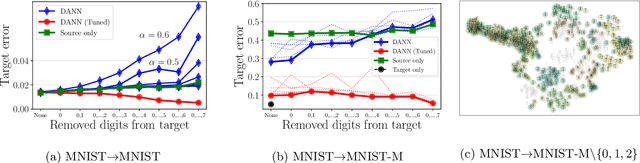
Abstract:Learning domain-invariant representations has become a popular approach to unsupervised domain adaptation and is often justified by invoking a particular suite of theoretical results. We argue that there are two significant flaws in such arguments. First, the results in question hold only for a fixed representation and do not account for information lost in non-invertible transformations. Second, domain invariance is often a far too strict requirement and does not always lead to consistent estimation, even under strong and favorable assumptions. In this work, we give generalization bounds for unsupervised domain adaptation that hold for any representation function by acknowledging the cost of non-invertibility. In addition, we show that penalizing distance between densities is often wasteful and propose a bound based on measuring the extent to which the support of the source domain covers the target domain. We perform experiments on well-known benchmarks that illustrate the short-comings of current standard practice.
The Variational Predictive Natural Gradient
Mar 07, 2019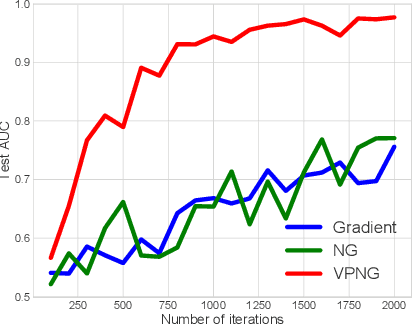
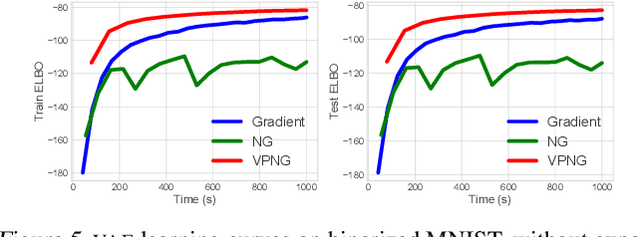
Abstract:Variational inference transforms posterior inference into parametric optimization thereby enabling the use of latent variable models where otherwise impractical. However, variational inference can be finicky when different variational parameters control variables that are strongly correlated under the model. Traditional natural gradients based on the variational approximation fail to correct for correlations when the approximation is not the true posterior. To address this, we construct a new natural gradient called the variational predictive natural gradient. It is constructed as an average of the Fisher information of the reparameterized predictive model distribution. Unlike traditional natural gradients for variational inference, this natural gradient accounts for the relationship between model parameters and variational parameters. We also show the variational predictive natural gradient relates to the negative Hessian of the expected log-likelihood. A simple example shows the insight. We demonstrate the empirical value of our method on a classification task, a deep generative model of images, and probabilistic matrix factorization for recommendation.
Soft Constraints for Inference with Declarative Knowledge
Jan 16, 2019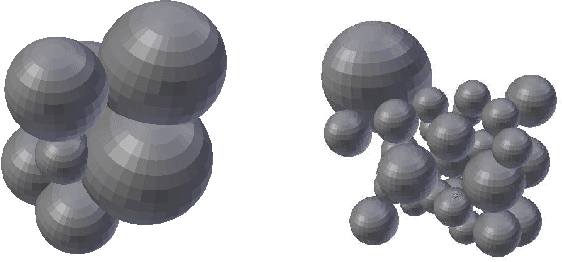
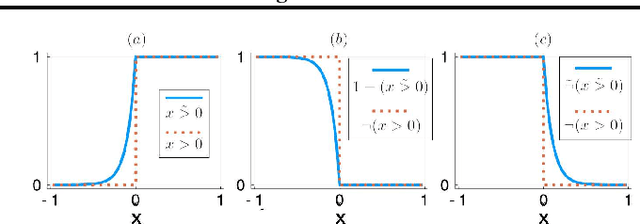
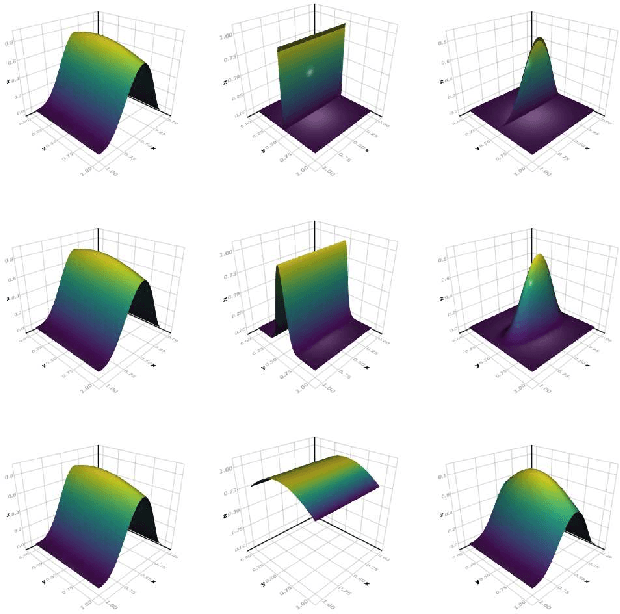
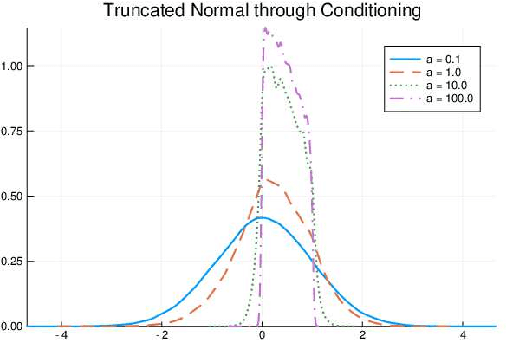
Abstract:We develop a likelihood free inference procedure for conditioning a probabilistic model on a predicate. A predicate is a Boolean valued function which expresses a yes/no question about a domain. Our contribution, which we call predicate exchange, constructs a softened predicate which takes value in the unit interval [0, 1] as opposed to a simply true or false. Intuitively, 1 corresponds to true, and a high value (such as 0.999) corresponds to "nearly true" as determined by a distance metric. We define Boolean algebra for soft predicates, such that they can be negated, conjoined and disjoined arbitrarily. A softened predicate can serve as a tractable proxy to a likelihood function for approximate posterior inference. However, to target exact inference, we temper the relaxation by a temperature parameter, and add a accept/reject phase use to replica exchange Markov Chain Mont Carlo, which exchanges states between a sequence of models conditioned on predicates at varying temperatures. We describe a lightweight implementation of predicate exchange that it provides a language independent layer that can be implemented on top of existingn modeling formalisms.
Multiple Causal Inference with Latent Confounding
Aug 07, 2018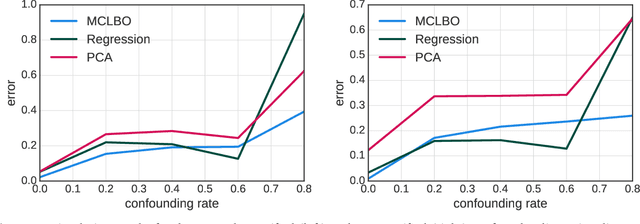
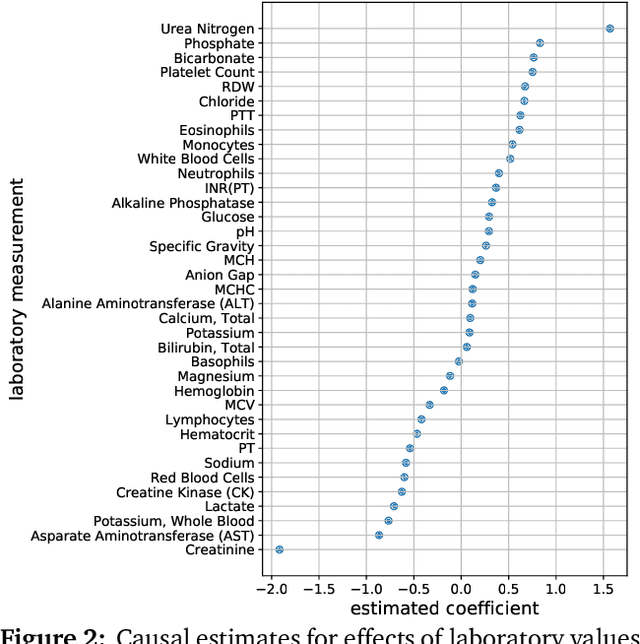
Abstract:Causal inference from observational data requires assumptions. These assumptions range from measuring confounders to identifying instruments. Traditionally, these assumptions have focused on estimation in a single causal problem. In this work, we develop techniques for causal estimation in causal problems with multiple treatments. We develop two assumptions based on shared confounding between treatments and independence of treatments given the confounder. Together these assumptions lead to a confounder estimator regularized by mutual information. For this estimator, we develop a tractable lower bound. To fit the outcome model, we use the residual information in the treatments given the confounder. We validate on simulations and an example from clinical medicine.
Noisin: Unbiased Regularization for Recurrent Neural Networks
Jul 13, 2018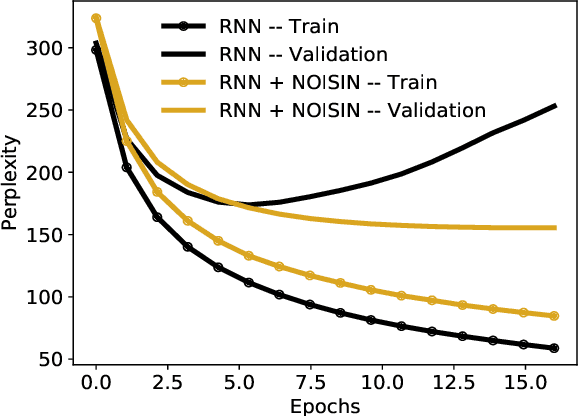
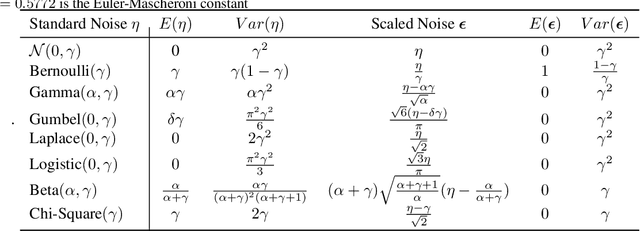

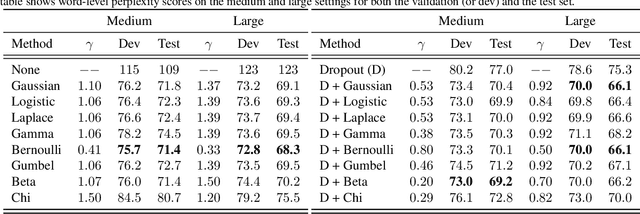
Abstract:Recurrent neural networks (RNNs) are powerful models of sequential data. They have been successfully used in domains such as text and speech. However, RNNs are susceptible to overfitting; regularization is important. In this paper we develop Noisin, a new method for regularizing RNNs. Noisin injects random noise into the hidden states of the RNN and then maximizes the corresponding marginal likelihood of the data. We show how Noisin applies to any RNN and we study many different types of noise. Noisin is unbiased--it preserves the underlying RNN on average. We characterize how Noisin regularizes its RNN both theoretically and empirically. On language modeling benchmarks, Noisin improves over dropout by as much as 12.2% on the Penn Treebank and 9.4% on the Wikitext-2 dataset. We also compared the state-of-the-art language model of Yang et al. 2017, both with and without Noisin. On the Penn Treebank, the method with Noisin more quickly reaches state-of-the-art performance.
Opportunities in Machine Learning for Healthcare
Jun 05, 2018Abstract:Healthcare is a natural arena for the application of machine learning, especially as modern electronic health records (EHRs) provide increasingly large amounts of data to answer clinically meaningful questions. However, clinical data and practice present unique challenges that complicate the use of common methodologies. This article serves as a primer on addressing these challenges and highlights opportunities for members of the machine learning and data science communities to contribute to this growing domain.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge