Polina Golland
Symmetric Volume Maps
Feb 05, 2022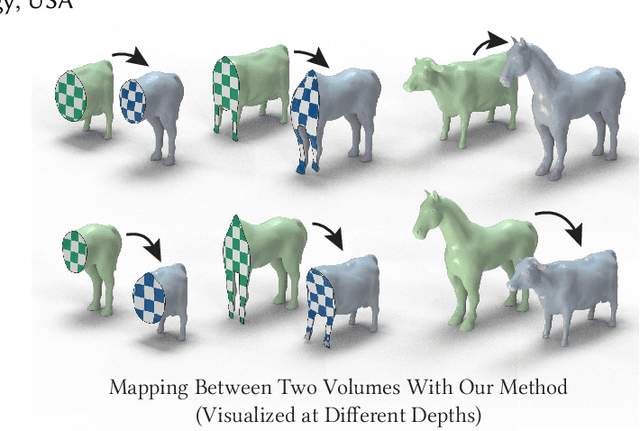
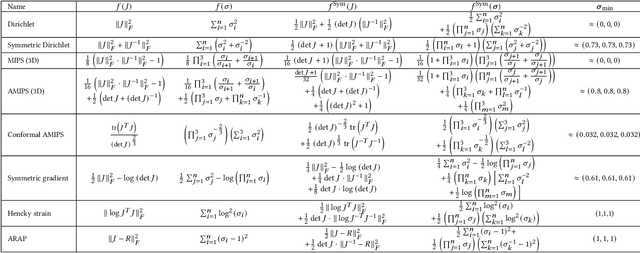

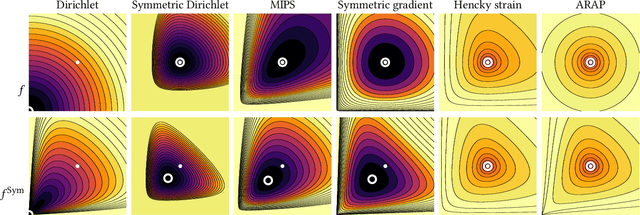
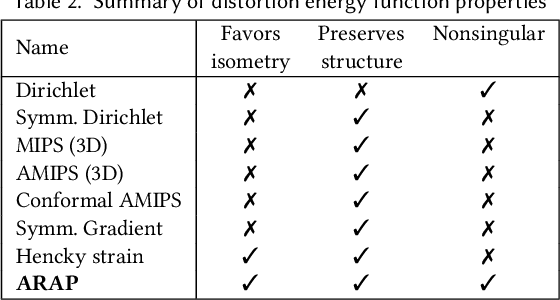
Abstract:Although shape correspondence is a central problem in geometry processing, most methods for this task apply only to two-dimensional surfaces. The neglected task of volumetric correspondence--a natural extension relevant to shapes extracted from simulation, medical imaging, volume rendering, and even improving surface maps of boundary representations--presents unique challenges that do not appear in the two-dimensional case. In this work, we propose a method for mapping between volumes represented as tetrahedral meshes. Our formulation minimizes a distortion energy designed to extract maps symmetrically, i.e., without dependence on the ordering of the source and target domains. We accompany our method with theoretical discussion describing the consequences of this symmetry assumption, leading us to select a symmetrized ARAP energy that favors isometric correspondences. Our final formulation optimizes for near-isometry while matching the boundary. We demonstrate our method on a diverse geometric dataset, producing low-distortion matchings that align to the boundary.
Hypernet-Ensemble Learning of Segmentation Probability for Medical Image Segmentation with Ambiguous Labels
Dec 13, 2021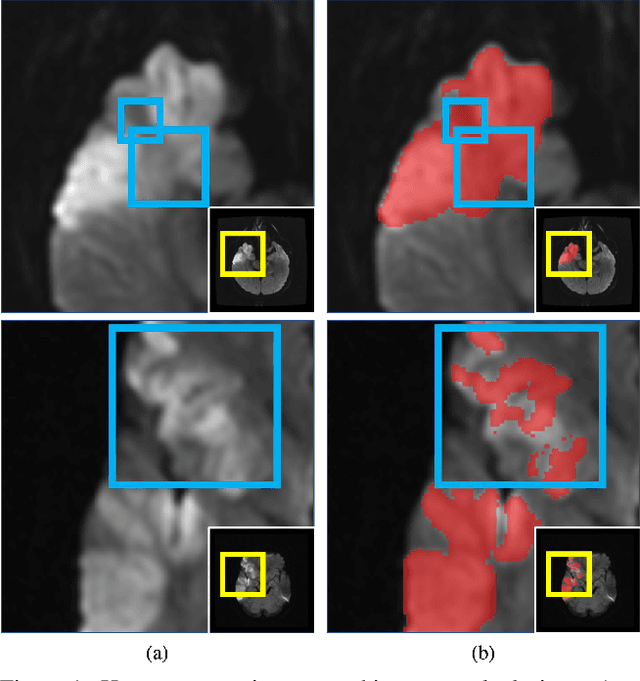
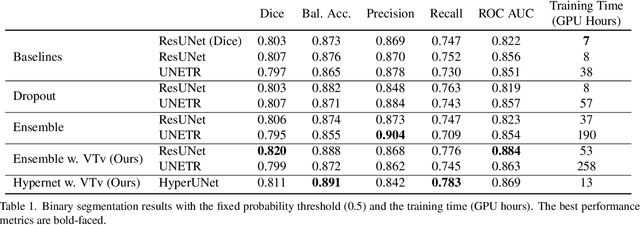
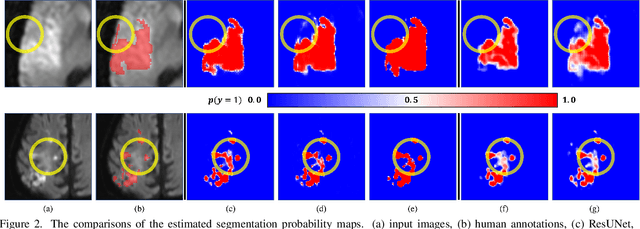
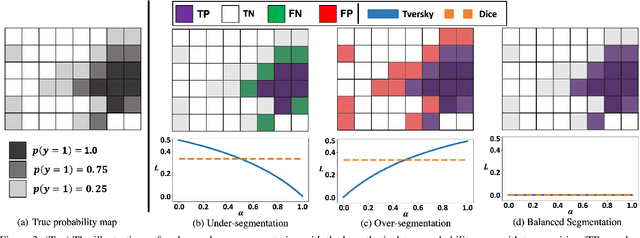
Abstract:Despite the superior performance of Deep Learning (DL) on numerous segmentation tasks, the DL-based approaches are notoriously overconfident about their prediction with highly polarized label probability. This is often not desirable for many applications with the inherent label ambiguity even in human annotations. This challenge has been addressed by leveraging multiple annotations per image and the segmentation uncertainty. However, multiple per-image annotations are often not available in a real-world application and the uncertainty does not provide full control on segmentation results to users. In this paper, we propose novel methods to improve the segmentation probability estimation without sacrificing performance in a real-world scenario that we have only one ambiguous annotation per image. We marginalize the estimated segmentation probability maps of networks that are encouraged to under-/over-segment with the varying Tversky loss without penalizing balanced segmentation. Moreover, we propose a unified hypernetwork ensemble method to alleviate the computational burden of training multiple networks. Our approaches successfully estimated the segmentation probability maps that reflected the underlying structures and provided the intuitive control on segmentation for the challenging 3D medical image segmentation. Although the main focus of our proposed methods is not to improve the binary segmentation performance, our approaches marginally outperformed the state-of-the-arts. The codes are available at \url{https://github.com/sh4174/HypernetEnsemble}.
Volumetric Parameterization of the Placenta to a Flattened Template
Nov 15, 2021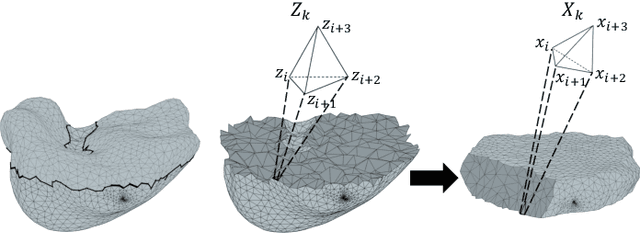
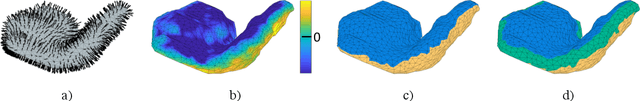
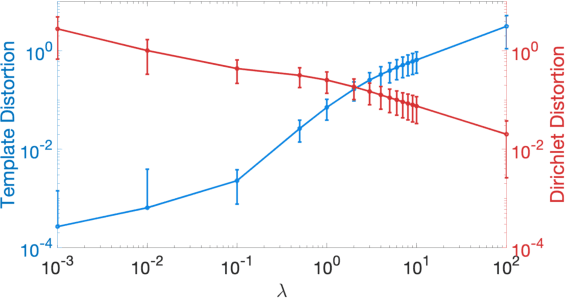
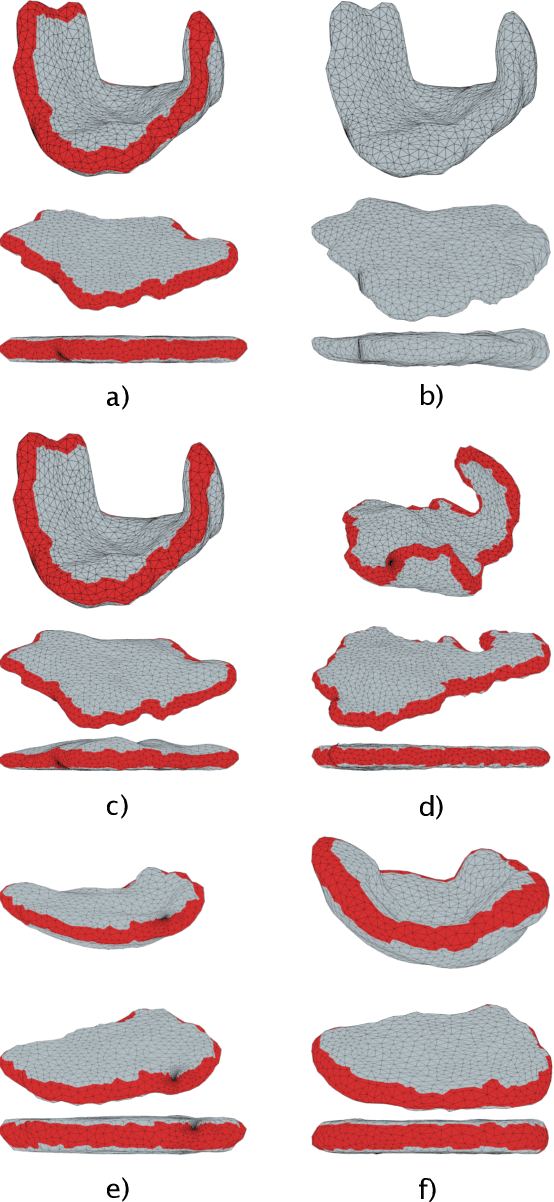
Abstract:We present a volumetric mesh-based algorithm for parameterizing the placenta to a flattened template to enable effective visualization of local anatomy and function. MRI shows potential as a research tool as it provides signals directly related to placental function. However, due to the curved and highly variable in vivo shape of the placenta, interpreting and visualizing these images is difficult. We address interpretation challenges by mapping the placenta so that it resembles the familiar ex vivo shape. We formulate the parameterization as an optimization problem for mapping the placental shape represented by a volumetric mesh to a flattened template. We employ the symmetric Dirichlet energy to control local distortion throughout the volume. Local injectivity in the mapping is enforced by a constrained line search during the gradient descent optimization. We validate our method using a research study of 111 placental shapes extracted from BOLD MRI images. Our mapping achieves sub-voxel accuracy in matching the template while maintaining low distortion throughout the volume. We demonstrate how the resulting flattening of the placenta improves visualization of anatomy and function. Our code is freely available at https://github.com/mabulnaga/placenta-flattening .
Image Classification with Consistent Supporting Evidence
Nov 13, 2021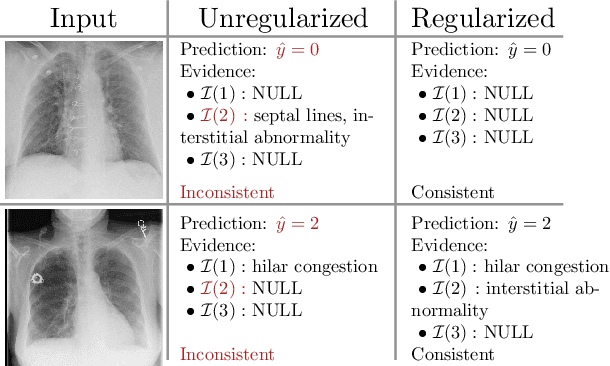
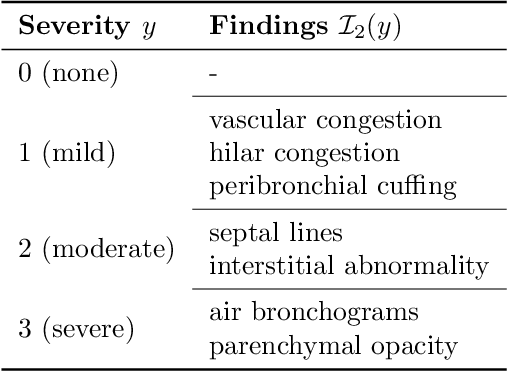
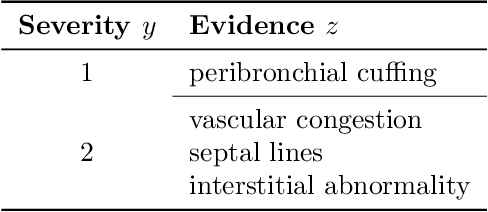


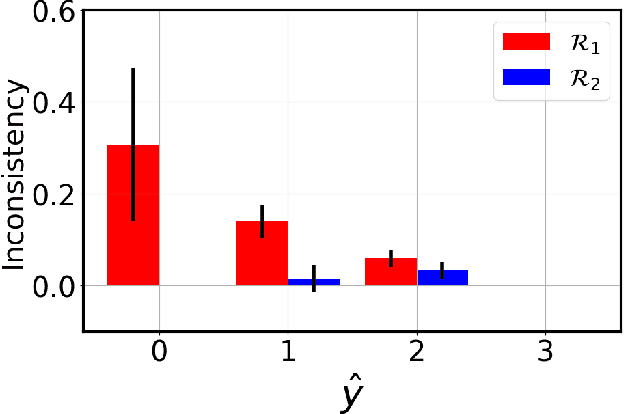
Abstract:Adoption of machine learning models in healthcare requires end users' trust in the system. Models that provide additional supportive evidence for their predictions promise to facilitate adoption. We define consistent evidence to be both compatible and sufficient with respect to model predictions. We propose measures of model inconsistency and regularizers that promote more consistent evidence. We demonstrate our ideas in the context of edema severity grading from chest radiographs. We demonstrate empirically that consistent models provide competitive performance while supporting interpretation.
3D-StyleGAN: A Style-Based Generative Adversarial Network for Generative Modeling of Three-Dimensional Medical Images
Jul 20, 2021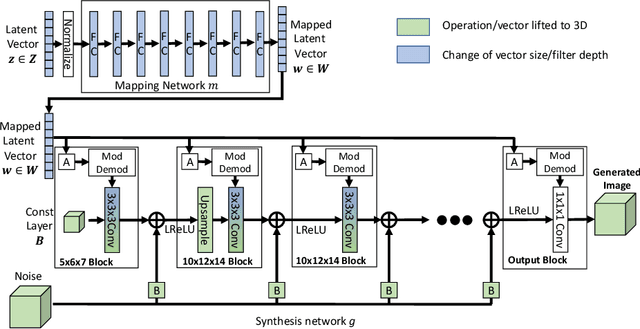
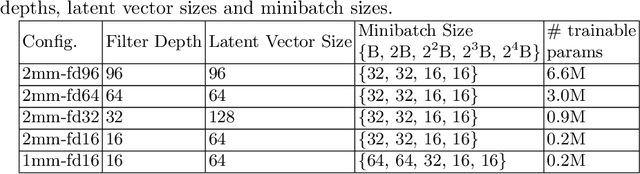
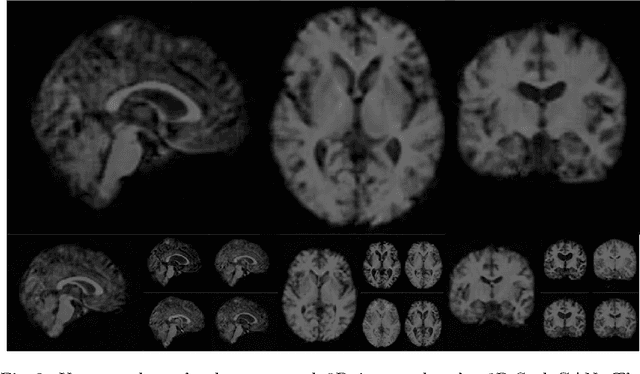
Abstract:Image synthesis via Generative Adversarial Networks (GANs) of three-dimensional (3D) medical images has great potential that can be extended to many medical applications, such as, image enhancement and disease progression modeling. However, current GAN technologies for 3D medical image synthesis need to be significantly improved to be readily adapted to real-world medical problems. In this paper, we extend the state-of-the-art StyleGAN2 model, which natively works with two-dimensional images, to enable 3D image synthesis. In addition to the image synthesis, we investigate the controllability and interpretability of the 3D-StyleGAN via style vectors inherited form the original StyleGAN2 that are highly suitable for medical applications: (i) the latent space projection and reconstruction of unseen real images, and (ii) style mixing. We demonstrate the 3D-StyleGAN's performance and feasibility with ~12,000 three-dimensional full brain MR T1 images, although it can be applied to any 3D volumetric images. Furthermore, we explore different configurations of hyperparameters to investigate potential improvement of the image synthesis with larger networks. The codes and pre-trained networks are available online: https://github.com/sh4174/3DStyleGAN.
STRESS: Super-Resolution for Dynamic Fetal MRI using Self-Supervised Learning
Jun 30, 2021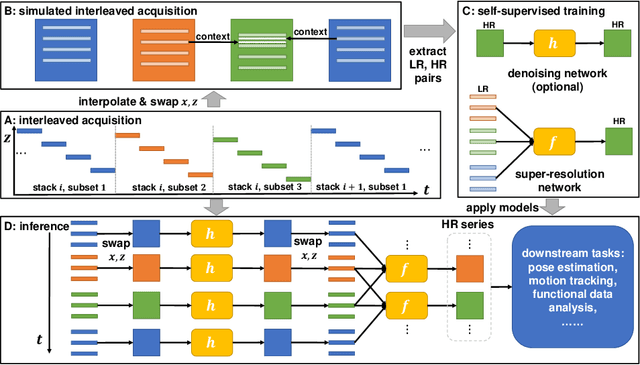
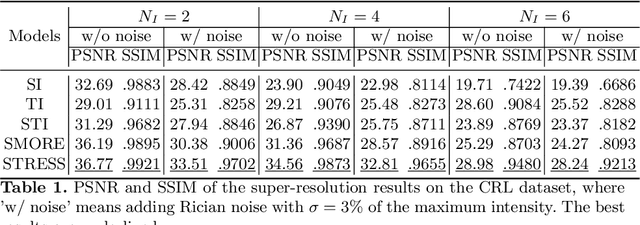
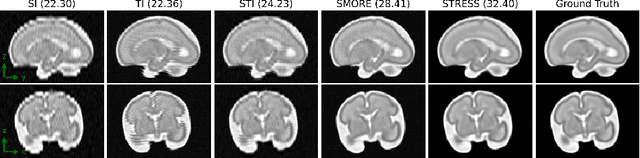

Abstract:Fetal motion is unpredictable and rapid on the scale of conventional MR scan times. Therefore, dynamic fetal MRI, which aims at capturing fetal motion and dynamics of fetal function, is limited to fast imaging techniques with compromises in image quality and resolution. Super-resolution for dynamic fetal MRI is still a challenge, especially when multi-oriented stacks of image slices for oversampling are not available and high temporal resolution for recording the dynamics of the fetus or placenta is desired. Further, fetal motion makes it difficult to acquire high-resolution images for supervised learning methods. To address this problem, in this work, we propose STRESS (Spatio-Temporal Resolution Enhancement with Simulated Scans), a self-supervised super-resolution framework for dynamic fetal MRI with interleaved slice acquisitions. Our proposed method simulates an interleaved slice acquisition along the high-resolution axis on the originally acquired data to generate pairs of low- and high-resolution images. Then, it trains a super-resolution network by exploiting both spatial and temporal correlations in the MR time series, which is used to enhance the resolution of the original data. Evaluations on both simulated and in utero data show that our proposed method outperforms other self-supervised super-resolution methods and improves image quality, which is beneficial to other downstream tasks and evaluations.
Segmentation of Anatomical Layers and Artifacts in Intravascular Polarization Sensitive Optical Coherence Tomography Using Attending Physician and Boundary Cardinality Lost Terms
May 11, 2021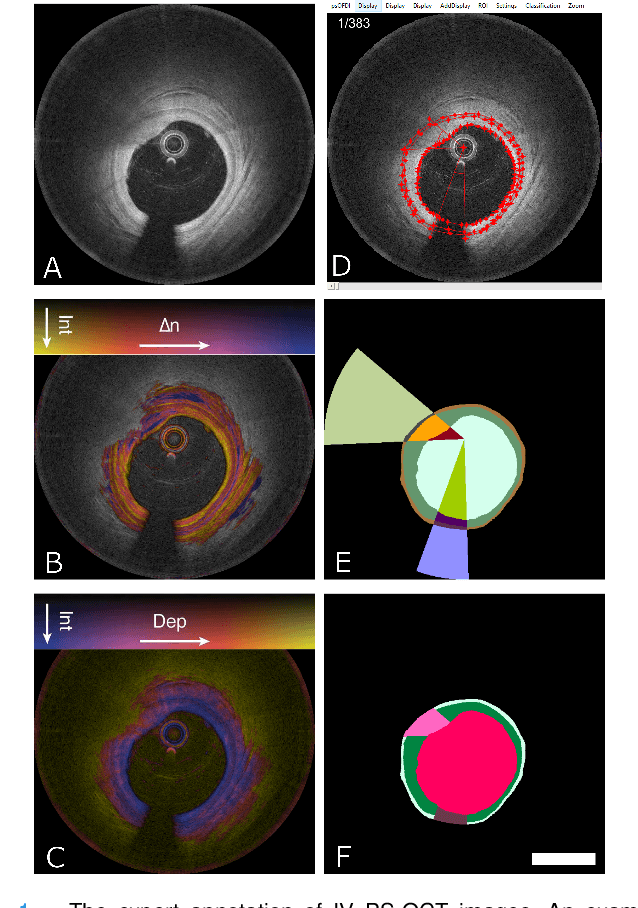
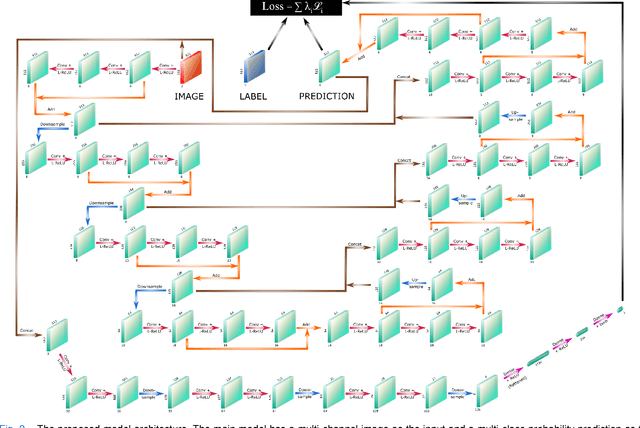
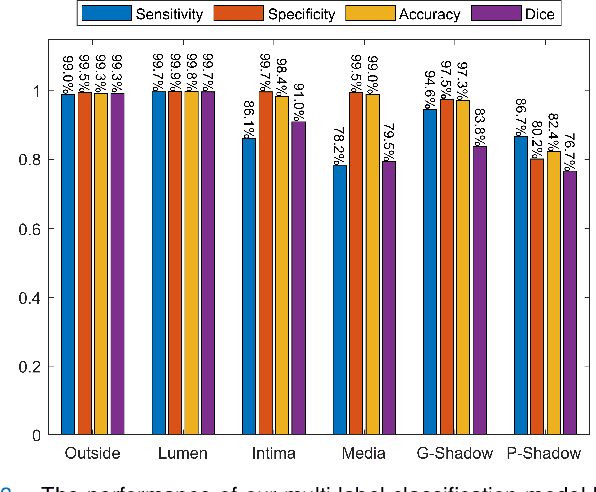

Abstract:Cardiovascular diseases are the leading cause of death and require a spectrum of diagnostic procedures as well as invasive interventions. Medical imaging is a vital part of the healthcare system, facilitating both diagnosis and guidance for intervention. Intravascular ultrasound and optical coherence tomography are widely available for characterizing coronary stenoses and provide critical vessel parameters to optimize percutaneous intervention. Intravascular polarization-sensitive optical coherence tomography (PS-OCT) can simultaneously provide high-resolution cross-sectional images of vascular structures while also revealing preponderant tissue components such as collagen and smooth muscle and thereby enhance plaque characterization. Automated interpretation of these features would facilitate the objective clinical investigation of the natural history and significance of coronary atheromas. Here, we propose a convolutional neural network model and optimize its performance using a new multi-term loss function to classify the lumen, intima, and media layers in addition to the guidewire and plaque artifacts. Our multi-class classification model outperforms the state-of-the-art methods in detecting the anatomical layers based on accuracy, Dice coefficient, and average boundary error. Furthermore, the proposed model segments two classes of major artifacts and detects the anatomical layers within the thickened vessel wall regions, which were excluded from analysis by other studies. The source code and the trained model are publicly available at https://github.com/mhaft/OCTseg .
Equivariant Filters for Efficient Tracking in 3D Imaging
Mar 18, 2021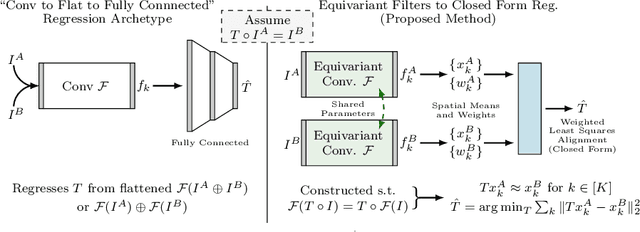

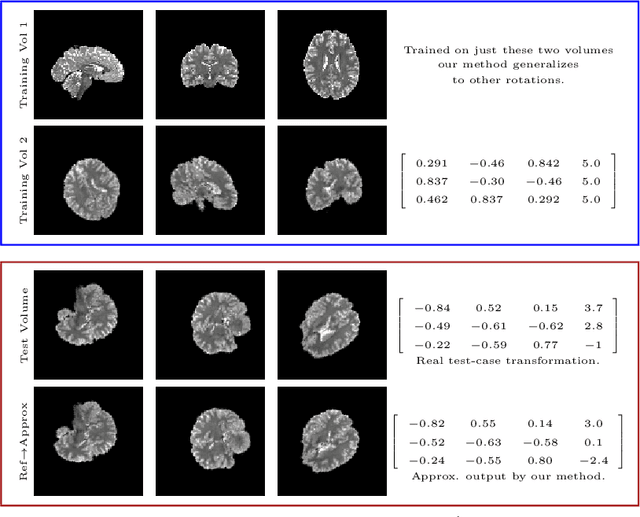
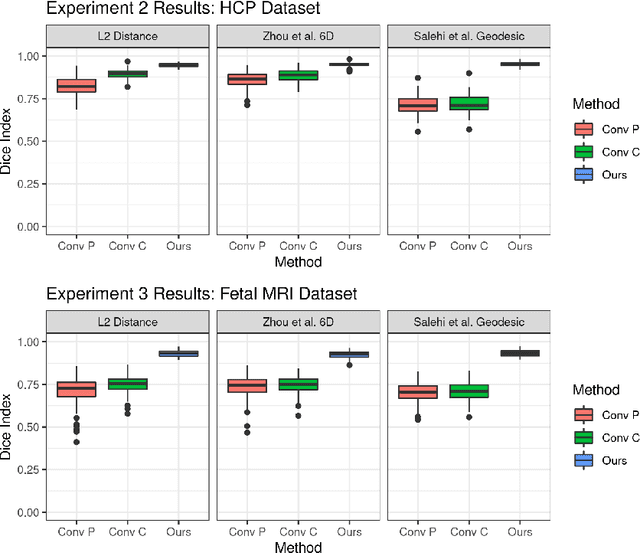
Abstract:We demonstrate an object tracking method for {3D} images with fixed computational cost and state-of-the-art performance. Previous methods predicted transformation parameters from convolutional layers. We instead propose an architecture that does not include either flattening of convolutional features or fully connected layers, but instead relies on equivariant filters to preserve transformations between inputs and outputs (e.g. rot./trans. of inputs rotate/translate outputs). The transformation is then derived in closed form from the outputs of the filters. This method is useful for applications requiring low latency, such as real-time tracking. We demonstrate our model on synthetically augmented adult brain MRI, as well as fetal brain MRI, which is the intended use-case.
Multimodal Representation Learning via Maximization of Local Mutual Information
Mar 08, 2021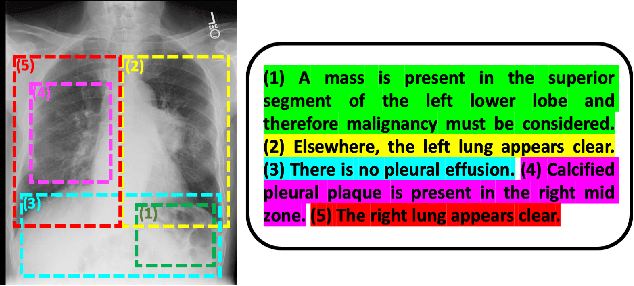
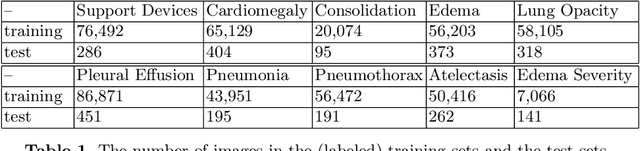
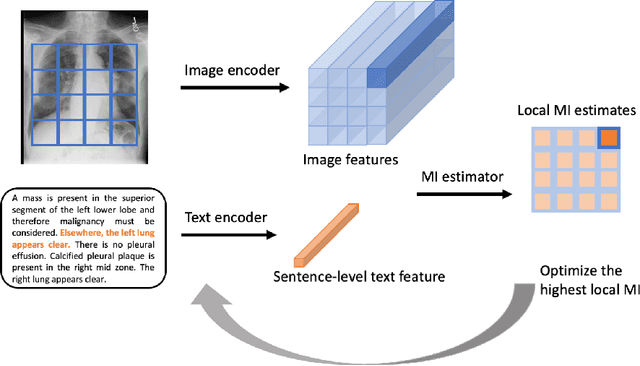
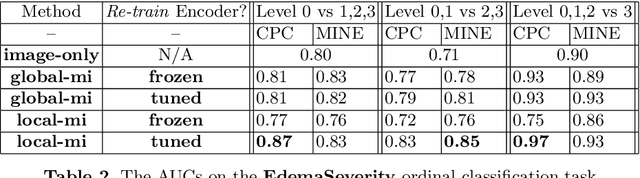
Abstract:We propose and demonstrate a representation learning approach by maximizing the mutual information between local features of images and text. The goal of this approach is to learn useful image representations by taking advantage of the rich information contained in the free text that describes the findings in the image. Our method learns image and text encoders by encouraging the resulting representations to exhibit high local mutual information. We make use of recent advances in mutual information estimation with neural network discriminators. We argue that, typically, the sum of local mutual information is a lower bound on the global mutual information. Our experimental results in the downstream image classification tasks demonstrate the advantages of using local features for image-text representation learning.
Harmonization and the Worst Scanner Syndrome
Jan 15, 2021Abstract:We show that for a wide class of harmonization/domain-invariance schemes several undesirable properties are unavoidable. If a predictive machine is made invariant to a set of domains, the accuracy of the output predictions (as measured by mutual information) is limited by the domain with the least amount of information to begin with. If a real label value is highly informative about the source domain, it cannot be accurately predicted by an invariant predictor. These results are simple and intuitive, but we believe that it is beneficial to state them for medical imaging harmonization.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge