Martin Bretzner
ISLA: A U-Net for MRI-based acute ischemic stroke lesion segmentation with deep supervision, attention, domain adaptation, and ensemble learning
Jan 13, 2026Abstract:Accurate delineation of acute ischemic stroke lesions in MRI is a key component of stroke diagnosis and management. In recent years, deep learning models have been successfully applied to the automatic segmentation of such lesions. While most proposed architectures are based on the U-Net framework, they primarily differ in their choice of loss functions and in the use of deep supervision, residual connections, and attention mechanisms. Moreover, many implementations are not publicly available, and the optimal configuration for acute ischemic stroke (AIS) lesion segmentation remains unclear. In this work, we introduce ISLA (Ischemic Stroke Lesion Analyzer), a new deep learning model for AIS lesion segmentation from diffusion MRI, trained on three multicenter databases totaling more than 1500 AIS participants. Through systematic optimization of the loss function, convolutional architecture, deep supervision, and attention mechanisms, we developed a robust segmentation framework. We further investigated unsupervised domain adaptation to improve generalization to an external clinical dataset. ISLA outperformed two state-of-the-art approaches for AIS lesion segmentation on an external test set. Codes and trained models will be made publicly available to facilitate reuse and reproducibility.
Federated Learning for MRI-based BrainAGE: a multicenter study on post-stroke functional outcome prediction
Jun 18, 2025



Abstract:$\textbf{Objective:}$ Brain-predicted age difference (BrainAGE) is a neuroimaging biomarker reflecting brain health. However, training robust BrainAGE models requires large datasets, often restricted by privacy concerns. This study evaluates the performance of federated learning (FL) for BrainAGE estimation in ischemic stroke patients treated with mechanical thrombectomy, and investigates its association with clinical phenotypes and functional outcomes. $\textbf{Methods:}$ We used FLAIR brain images from 1674 stroke patients across 16 hospital centers. We implemented standard machine learning and deep learning models for BrainAGE estimates under three data management strategies: centralized learning (pooled data), FL (local training at each site), and single-site learning. We reported prediction errors and examined associations between BrainAGE and vascular risk factors (e.g., diabetes mellitus, hypertension, smoking), as well as functional outcomes at three months post-stroke. Logistic regression evaluated BrainAGE's predictive value for these outcomes, adjusting for age, sex, vascular risk factors, stroke severity, time between MRI and arterial puncture, prior intravenous thrombolysis, and recanalisation outcome. $\textbf{Results:}$ While centralized learning yielded the most accurate predictions, FL consistently outperformed single-site models. BrainAGE was significantly higher in patients with diabetes mellitus across all models. Comparisons between patients with good and poor functional outcomes, and multivariate predictions of these outcomes showed the significance of the association between BrainAGE and post-stroke recovery. $\textbf{Conclusion:}$ FL enables accurate age predictions without data centralization. The strong association between BrainAGE, vascular risk factors, and post-stroke recovery highlights its potential for prognostic modeling in stroke care.
Hypernet-Ensemble Learning of Segmentation Probability for Medical Image Segmentation with Ambiguous Labels
Dec 13, 2021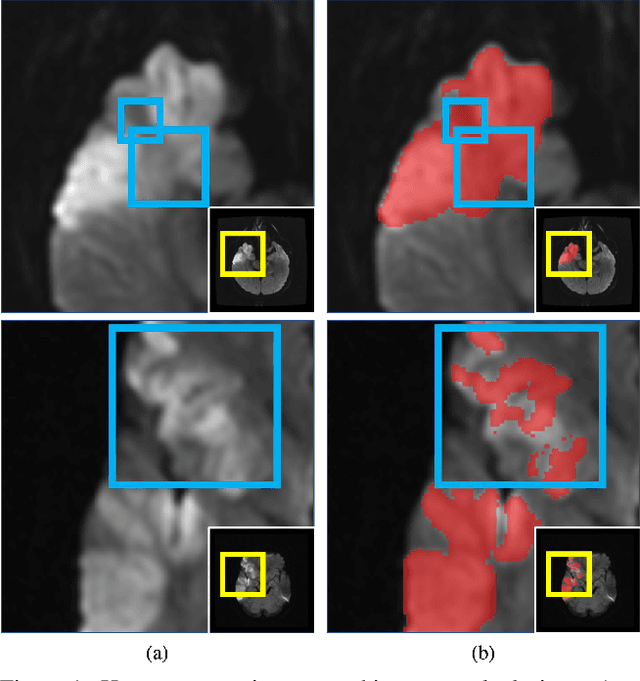
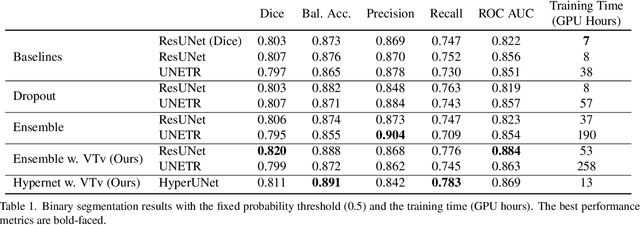
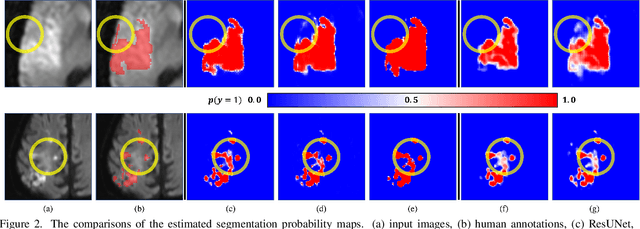
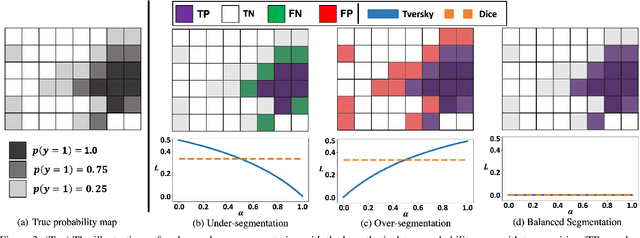
Abstract:Despite the superior performance of Deep Learning (DL) on numerous segmentation tasks, the DL-based approaches are notoriously overconfident about their prediction with highly polarized label probability. This is often not desirable for many applications with the inherent label ambiguity even in human annotations. This challenge has been addressed by leveraging multiple annotations per image and the segmentation uncertainty. However, multiple per-image annotations are often not available in a real-world application and the uncertainty does not provide full control on segmentation results to users. In this paper, we propose novel methods to improve the segmentation probability estimation without sacrificing performance in a real-world scenario that we have only one ambiguous annotation per image. We marginalize the estimated segmentation probability maps of networks that are encouraged to under-/over-segment with the varying Tversky loss without penalizing balanced segmentation. Moreover, we propose a unified hypernetwork ensemble method to alleviate the computational burden of training multiple networks. Our approaches successfully estimated the segmentation probability maps that reflected the underlying structures and provided the intuitive control on segmentation for the challenging 3D medical image segmentation. Although the main focus of our proposed methods is not to improve the binary segmentation performance, our approaches marginally outperformed the state-of-the-arts. The codes are available at \url{https://github.com/sh4174/HypernetEnsemble}.
3D-StyleGAN: A Style-Based Generative Adversarial Network for Generative Modeling of Three-Dimensional Medical Images
Jul 20, 2021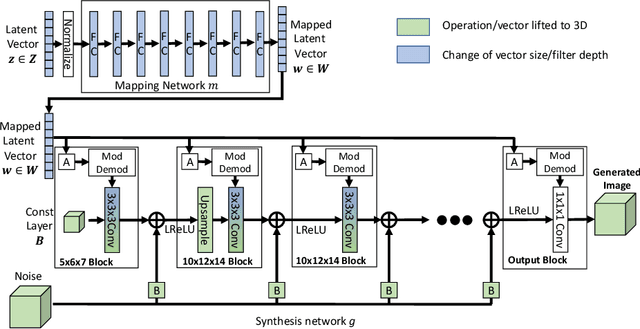
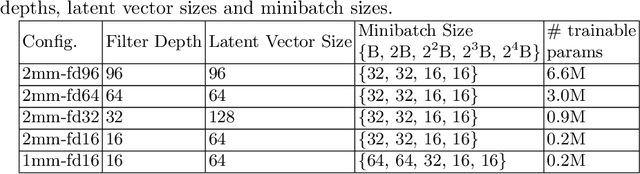
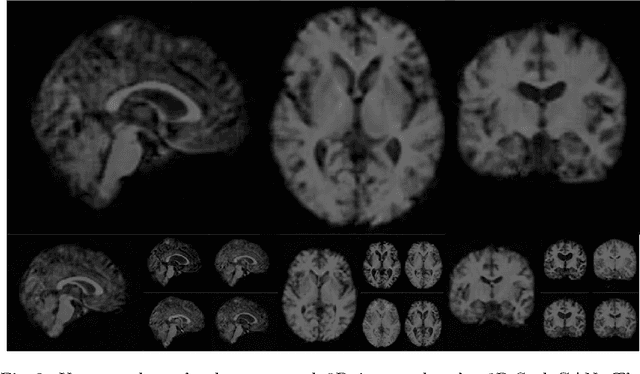
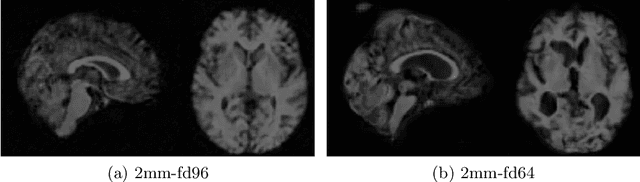
Abstract:Image synthesis via Generative Adversarial Networks (GANs) of three-dimensional (3D) medical images has great potential that can be extended to many medical applications, such as, image enhancement and disease progression modeling. However, current GAN technologies for 3D medical image synthesis need to be significantly improved to be readily adapted to real-world medical problems. In this paper, we extend the state-of-the-art StyleGAN2 model, which natively works with two-dimensional images, to enable 3D image synthesis. In addition to the image synthesis, we investigate the controllability and interpretability of the 3D-StyleGAN via style vectors inherited form the original StyleGAN2 that are highly suitable for medical applications: (i) the latent space projection and reconstruction of unseen real images, and (ii) style mixing. We demonstrate the 3D-StyleGAN's performance and feasibility with ~12,000 three-dimensional full brain MR T1 images, although it can be applied to any 3D volumetric images. Furthermore, we explore different configurations of hyperparameters to investigate potential improvement of the image synthesis with larger networks. The codes and pre-trained networks are available online: https://github.com/sh4174/3DStyleGAN.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge