uTRAND: Unsupervised Anomaly Detection in Traffic Trajectories
Apr 19, 2024Giacomo D'Amicantonio, Egor Bondarau, Peter H. N. de With
Deep learning-based approaches have achieved significant improvements on public video anomaly datasets, but often do not perform well in real-world applications. This paper addresses two issues: the lack of labeled data and the difficulty of explaining the predictions of a neural network. To this end, we present a framework called uTRAND, that shifts the problem of anomalous trajectory prediction from the pixel space to a semantic-topological domain. The framework detects and tracks all types of traffic agents in bird's-eye-view videos of traffic cameras mounted at an intersection. By conceptualizing the intersection as a patch-based graph, it is shown that the framework learns and models the normal behaviour of traffic agents without costly manual labeling. Furthermore, uTRAND allows to formulate simple rules to classify anomalous trajectories in a way suited for human interpretation. We show that uTRAND outperforms other state-of-the-art approaches on a dataset of anomalous trajectories collected in a real-world setting, while producing explainable detection results.
IndustReal: A Dataset for Procedure Step Recognition Handling Execution Errors in Egocentric Videos in an Industrial-Like Setting
Oct 26, 2023Tim J. Schoonbeek, Tim Houben, Hans Onvlee, Peter H. N. de With, Fons van der Sommen
Although action recognition for procedural tasks has received notable attention, it has a fundamental flaw in that no measure of success for actions is provided. This limits the applicability of such systems especially within the industrial domain, since the outcome of procedural actions is often significantly more important than the mere execution. To address this limitation, we define the novel task of procedure step recognition (PSR), focusing on recognizing the correct completion and order of procedural steps. Alongside the new task, we also present the multi-modal IndustReal dataset. Unlike currently available datasets, IndustReal contains procedural errors (such as omissions) as well as execution errors. A significant part of these errors are exclusively present in the validation and test sets, making IndustReal suitable to evaluate robustness of algorithms to new, unseen mistakes. Additionally, to encourage reproducibility and allow for scalable approaches trained on synthetic data, the 3D models of all parts are publicly available. Annotations and benchmark performance are provided for action recognition and assembly state detection, as well as the new PSR task. IndustReal, along with the code and model weights, is available at: https://github.com/TimSchoonbeek/IndustReal .
Investigating and Improving Latent Density Segmentation Models for Aleatoric Uncertainty Quantification in Medical Imaging
Aug 15, 2023M. M. Amaan Valiuddin, Christiaan G. A. Viviers, Ruud J. G. van Sloun, Peter H. N. de With, Fons van der Sommen
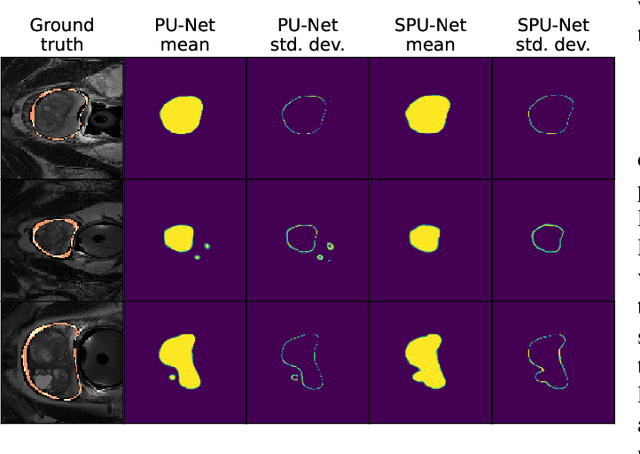
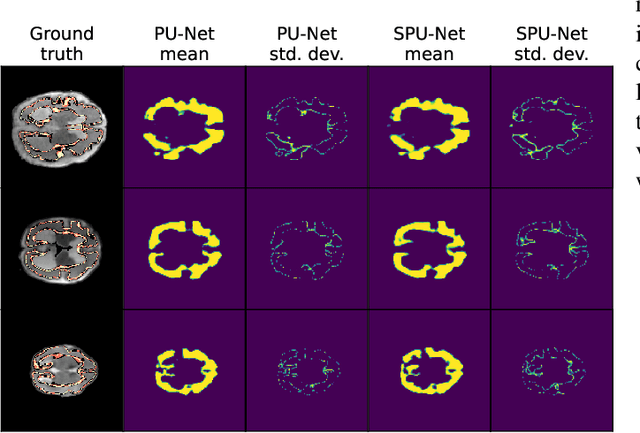



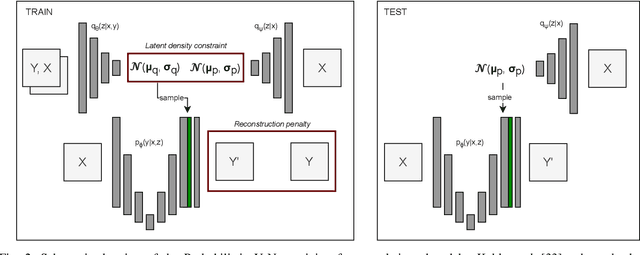
Data uncertainties, such as sensor noise or occlusions, can introduce irreducible ambiguities in images, which result in varying, yet plausible, semantic hypotheses. In Machine Learning, this ambiguity is commonly referred to as aleatoric uncertainty. Latent density models can be utilized to address this problem in image segmentation. The most popular approach is the Probabilistic U-Net (PU-Net), which uses latent Normal densities to optimize the conditional data log-likelihood Evidence Lower Bound. In this work, we demonstrate that the PU- Net latent space is severely inhomogenous. As a result, the effectiveness of gradient descent is inhibited and the model becomes extremely sensitive to the localization of the latent space samples, resulting in defective predictions. To address this, we present the Sinkhorn PU-Net (SPU-Net), which uses the Sinkhorn Divergence to promote homogeneity across all latent dimensions, effectively improving gradient-descent updates and model robustness. Our results show that by applying this on public datasets of various clinical segmentation problems, the SPU-Net receives up to 11% performance gains compared against preceding latent variable models for probabilistic segmentation on the Hungarian-Matched metric. The results indicate that by encouraging a homogeneous latent space, one can significantly improve latent density modeling for medical image segmentation.
A signal processing interpretation of noise-reduction convolutional neural networks
Jul 25, 2023Luis A. Zavala-Mondragón, Peter H. N. de With, Fons van der Sommen
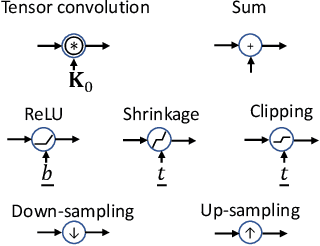
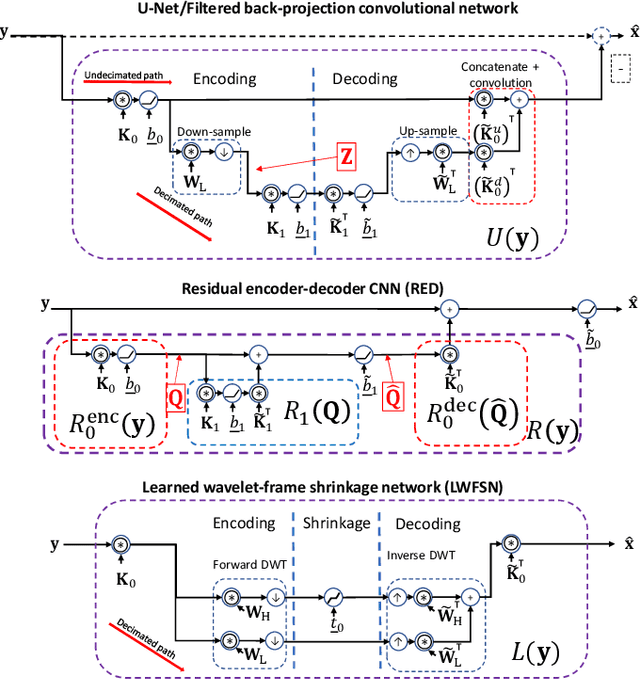
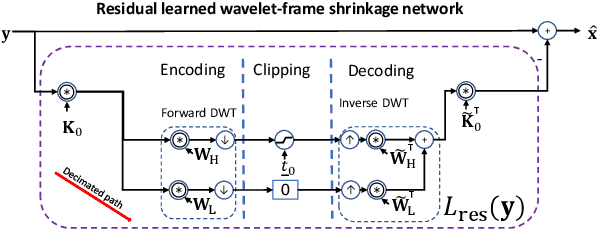
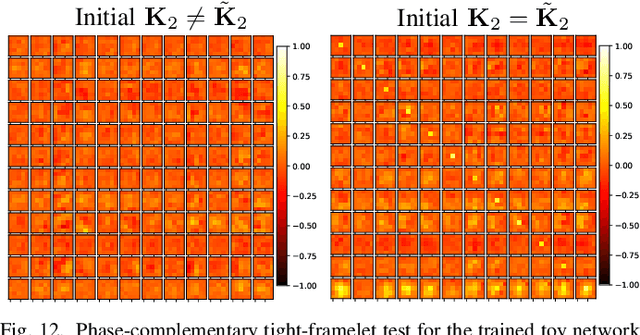
Encoding-decoding CNNs play a central role in data-driven noise reduction and can be found within numerous deep-learning algorithms. However, the development of these CNN architectures is often done in ad-hoc fashion and theoretical underpinnings for important design choices is generally lacking. Up to this moment there are different existing relevant works that strive to explain the internal operation of these CNNs. Still, these ideas are either scattered and/or may require significant expertise to be accessible for a bigger audience. In order to open up this exciting field, this article builds intuition on the theory of deep convolutional framelets and explains diverse ED CNN architectures in a unified theoretical framework. By connecting basic principles from signal processing to the field of deep learning, this self-contained material offers significant guidance for designing robust and efficient novel CNN architectures.
Probabilistic 3D segmentation for aleatoric uncertainty quantification in full 3D medical data
May 01, 2023Christiaan G. A. Viviers, Amaan M. M. Valiuddin, Peter H. N. de With, Fons van der Sommen
Uncertainty quantification in medical images has become an essential addition to segmentation models for practical application in the real world. Although there are valuable developments in accurate uncertainty quantification methods using 2D images and slices of 3D volumes, in clinical practice, the complete 3D volumes (such as CT and MRI scans) are used to evaluate and plan the medical procedure. As a result, the existing 2D methods miss the rich 3D spatial information when resolving the uncertainty. A popular approach for quantifying the ambiguity in the data is to learn a distribution over the possible hypotheses. In recent work, this ambiguity has been modeled to be strictly Gaussian. Normalizing Flows (NFs) are capable of modelling more complex distributions and thus, better fit the embedding space of the data. To this end, we have developed a 3D probabilistic segmentation framework augmented with NFs, to enable capturing the distributions of various complexity. To test the proposed approach, we evaluate the model on the LIDC-IDRI dataset for lung nodule segmentation and quantify the aleatoric uncertainty introduced by the multi-annotator setting and inherent ambiguity in the CT data. Following this approach, we are the first to present a 3D Squared Generalized Energy Distance (GED) of 0.401 and a high 0.468 Hungarian-matched 3D IoU. The obtained results reveal the value in capturing the 3D uncertainty, using a flexible posterior distribution augmented with a Normalizing Flow. Finally, we present the aleatoric uncertainty in a visual manner with the aim to provide clinicians with additional insight into data ambiguity and facilitating more informed decision-making.
Towards real-time 6D pose estimation of objects in single-view cone-beam X-ray
Nov 06, 2022Christiaan G. A. Viviers, Joel de Bruijn, Lena Filatova, Peter H. N. de With, Fons van der Sommen
Deep learning-based pose estimation algorithms can successfully estimate the pose of objects in an image, especially in the field of color images. 6D Object pose estimation based on deep learning models for X-ray images often use custom architectures that employ extensive CAD models and simulated data for training purposes. Recent RGB-based methods opt to solve pose estimation problems using small datasets, making them more attractive for the X-ray domain where medical data is scarcely available. We refine an existing RGB-based model (SingleShotPose) to estimate the 6D pose of a marked cube from grayscale X-ray images by creating a generic solution trained on only real X-ray data and adjusted for X-ray acquisition geometry. The model regresses 2D control points and calculates the pose through 2D/3D correspondences using Perspective-n-Point(PnP), allowing a single trained model to be used across all supporting cone-beam-based X-ray geometries. Since modern X-ray systems continuously adjust acquisition parameters during a procedure, it is essential for such a pose estimation network to consider these parameters in order to be deployed successfully and find a real use case. With a 5-cm/5-degree accuracy of 93% and an average 3D rotation error of 2.2 degrees, the results of the proposed approach are comparable with state-of-the-art alternatives, while requiring significantly less real training examples and being applicable in real-time applications.
Efficient Out-of-Distribution Detection of Melanoma with Wavelet-based Normalizing Flows
Aug 10, 2022M. M. Amaan Valiuddin, Christiaan G. A. Viviers, Ruud J. G. van Sloun, Peter H. N. de With, Fons van der Sommen
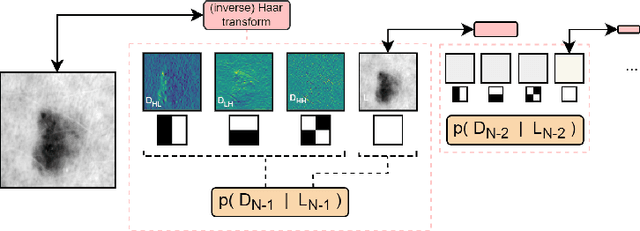

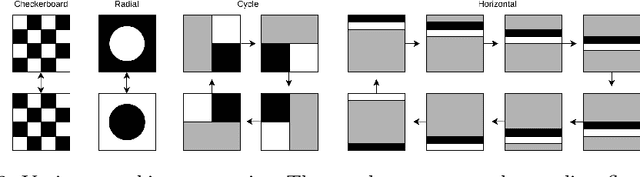
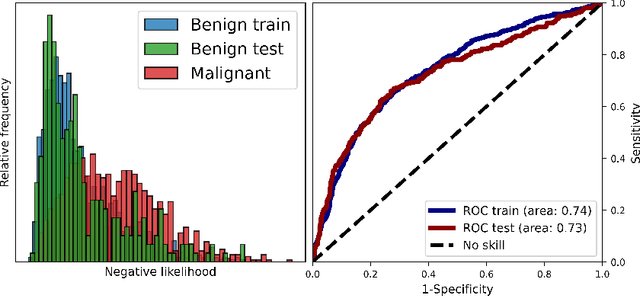
Melanoma is a serious form of skin cancer with high mortality rate at later stages. Fortunately, when detected early, the prognosis of melanoma is promising and malignant melanoma incidence rates are relatively low. As a result, datasets are heavily imbalanced which complicates training current state-of-the-art supervised classification AI models. We propose to use generative models to learn the benign data distribution and detect Out-of-Distribution (OOD) malignant images through density estimation. Normalizing Flows (NFs) are ideal candidates for OOD detection due to their ability to compute exact likelihoods. Nevertheless, their inductive biases towards apparent graphical features rather than semantic context hamper accurate OOD detection. In this work, we aim at using these biases with domain-level knowledge of melanoma, to improve likelihood-based OOD detection of malignant images. Our encouraging results demonstrate potential for OOD detection of melanoma using NFs. We achieve a 9% increase in Area Under Curve of the Receiver Operating Characteristics by using wavelet-based NFs. This model requires significantly less parameters for inference making it more applicable on edge devices. The proposed methodology can aid medical experts with diagnosis of skin-cancer patients and continuously increase survival rates. Furthermore, this research paves the way for other areas in oncology with similar data imbalance issues.
Improved Pancreatic Tumor Detection by Utilizing Clinically-Relevant Secondary Features
Aug 06, 2022Christiaan G. A. Viviers, Mark Ramaekers, Peter H. N. de With, Dimitrios Mavroeidis, Joost Nederend, Misha Luyer, Fons van der Sommen
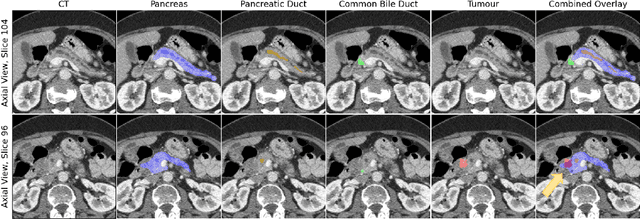
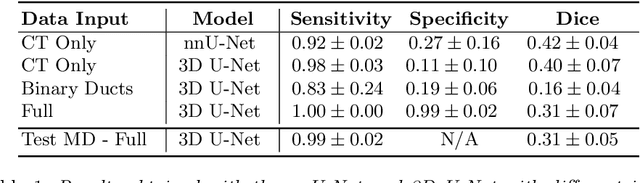
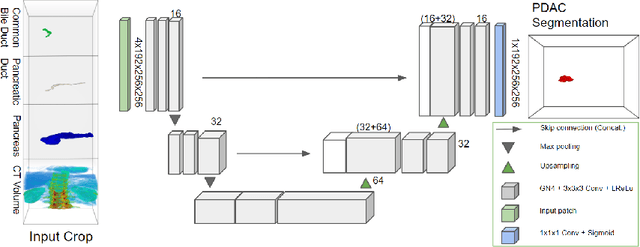

Pancreatic cancer is one of the global leading causes of cancer-related deaths. Despite the success of Deep Learning in computer-aided diagnosis and detection (CAD) methods, little attention has been paid to the detection of Pancreatic Cancer. We propose a method for detecting pancreatic tumor that utilizes clinically-relevant features in the surrounding anatomical structures, thereby better aiming to exploit the radiologist's knowledge compared to other, conventional deep learning approaches. To this end, we collect a new dataset consisting of 99 cases with pancreatic ductal adenocarcinoma (PDAC) and 97 control cases without any pancreatic tumor. Due to the growth pattern of pancreatic cancer, the tumor may not be always visible as a hypodense lesion, therefore experts refer to the visibility of secondary external features that may indicate the presence of the tumor. We propose a method based on a U-Net-like Deep CNN that exploits the following external secondary features: the pancreatic duct, common bile duct and the pancreas, along with a processed CT scan. Using these features, the model segments the pancreatic tumor if it is present. This segmentation for classification and localization approach achieves a performance of 99% sensitivity (one case missed) and 99% specificity, which realizes a 5% increase in sensitivity over the previous state-of-the-art method. The model additionally provides location information with reasonable accuracy and a shorter inference time compared to previous PDAC detection methods. These results offer a significant performance improvement and highlight the importance of incorporating the knowledge of the clinical expert when developing novel CAD methods.
Medical Instrument Segmentation in 3D US by Hybrid Constrained Semi-Supervised Learning
Jul 30, 2021Hongxu Yang, Caifeng Shan, R. Arthur Bouwman, Lukas R. C. Dekker, Alexander F. Kolen, Peter H. N. de With
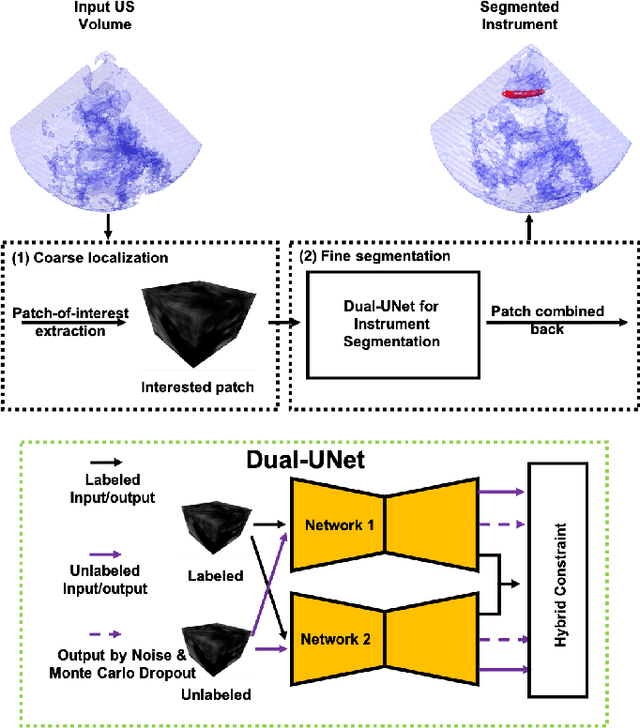
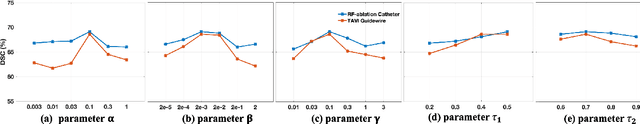
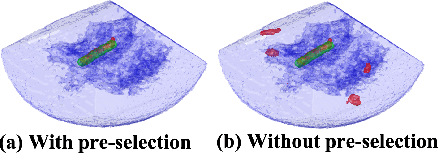
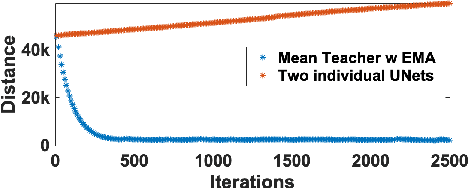
Medical instrument segmentation in 3D ultrasound is essential for image-guided intervention. However, to train a successful deep neural network for instrument segmentation, a large number of labeled images are required, which is expensive and time-consuming to obtain. In this article, we propose a semi-supervised learning (SSL) framework for instrument segmentation in 3D US, which requires much less annotation effort than the existing methods. To achieve the SSL learning, a Dual-UNet is proposed to segment the instrument. The Dual-UNet leverages unlabeled data using a novel hybrid loss function, consisting of uncertainty and contextual constraints. Specifically, the uncertainty constraints leverage the uncertainty estimation of the predictions of the UNet, and therefore improve the unlabeled information for SSL training. In addition, contextual constraints exploit the contextual information of the training images, which are used as the complementary information for voxel-wise uncertainty estimation. Extensive experiments on multiple ex-vivo and in-vivo datasets show that our proposed method achieves Dice score of about 68.6%-69.1% and the inference time of about 1 sec. per volume. These results are better than the state-of-the-art SSL methods and the inference time is comparable to the supervised approaches.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge