Paul Swoboda
Max Planck Institute for Informatics, Saarbrücken
A Multidimensional Analysis of Social Biases in Vision Transformers
Aug 03, 2023Abstract:The embedding spaces of image models have been shown to encode a range of social biases such as racism and sexism. Here, we investigate specific factors that contribute to the emergence of these biases in Vision Transformers (ViT). Therefore, we measure the impact of training data, model architecture, and training objectives on social biases in the learned representations of ViTs. Our findings indicate that counterfactual augmentation training using diffusion-based image editing can mitigate biases, but does not eliminate them. Moreover, we find that larger models are less biased than smaller models, and that models trained using discriminative objectives are less biased than those trained using generative objectives. In addition, we observe inconsistencies in the learned social biases. To our surprise, ViTs can exhibit opposite biases when trained on the same data set using different self-supervised objectives. Our findings give insights into the factors that contribute to the emergence of social biases and suggests that we could achieve substantial fairness improvements based on model design choices.
LVM-Med: Learning Large-Scale Self-Supervised Vision Models for Medical Imaging via Second-order Graph Matching
Jul 09, 2023



Abstract:Obtaining large pre-trained models that can be fine-tuned to new tasks with limited annotated samples has remained an open challenge for medical imaging data. While pre-trained deep networks on ImageNet and vision-language foundation models trained on web-scale data are prevailing approaches, their effectiveness on medical tasks is limited due to the significant domain shift between natural and medical images. To bridge this gap, we introduce LVM-Med, the first family of deep networks trained on large-scale medical datasets. We have collected approximately 1.3 million medical images from 55 publicly available datasets, covering a large number of organs and modalities such as CT, MRI, X-ray, and Ultrasound. We benchmark several state-of-the-art self-supervised algorithms on this dataset and propose a novel self-supervised contrastive learning algorithm using a graph-matching formulation. The proposed approach makes three contributions: (i) it integrates prior pair-wise image similarity metrics based on local and global information; (ii) it captures the structural constraints of feature embeddings through a loss function constructed via a combinatorial graph-matching objective; and (iii) it can be trained efficiently end-to-end using modern gradient-estimation techniques for black-box solvers. We thoroughly evaluate the proposed LVM-Med on 15 downstream medical tasks ranging from segmentation and classification to object detection, and both for the in and out-of-distribution settings. LVM-Med empirically outperforms a number of state-of-the-art supervised, self-supervised, and foundation models. For challenging tasks such as Brain Tumor Classification or Diabetic Retinopathy Grading, LVM-Med improves previous vision-language models trained on 1 billion masks by 6-7% while using only a ResNet-50.
ClusterFuG: Clustering Fully connected Graphs by Multicut
Jan 28, 2023



Abstract:We propose a graph clustering formulation based on multicut (a.k.a. weighted correlation clustering) on the complete graph. Our formulation does not need specification of the graph topology as in the original sparse formulation of multicut, making our approach simpler and potentially better performing. In contrast to unweighted correlation clustering we allow for a more expressive weighted cost structure. In dense multicut, the clustering objective is given in a factorized form as inner products of node feature vectors. This allows for an efficient formulation and inference in contrast to multicut/weighted correlation clustering, which has at least quadratic representation and computation complexity when working on the complete graph. We show how to rewrite classical greedy algorithms for multicut in our dense setting and how to modify them for greater efficiency and solution quality. In particular, our algorithms scale to graphs with tens of thousands of nodes. Empirical evidence on instance segmentation on Cityscapes and clustering of ImageNet datasets shows the merits of our approach.
Joint Self-Supervised Image-Volume Representation Learning with Intra-Inter Contrastive Clustering
Dec 04, 2022



Abstract:Collecting large-scale medical datasets with fully annotated samples for training of deep networks is prohibitively expensive, especially for 3D volume data. Recent breakthroughs in self-supervised learning (SSL) offer the ability to overcome the lack of labeled training samples by learning feature representations from unlabeled data. However, most current SSL techniques in the medical field have been designed for either 2D images or 3D volumes. In practice, this restricts the capability to fully leverage unlabeled data from numerous sources, which may include both 2D and 3D data. Additionally, the use of these pre-trained networks is constrained to downstream tasks with compatible data dimensions. In this paper, we propose a novel framework for unsupervised joint learning on 2D and 3D data modalities. Given a set of 2D images or 2D slices extracted from 3D volumes, we construct an SSL task based on a 2D contrastive clustering problem for distinct classes. The 3D volumes are exploited by computing vectored embedding at each slice and then assembling a holistic feature through deformable self-attention mechanisms in Transformer, allowing incorporating long-range dependencies between slices inside 3D volumes. These holistic features are further utilized to define a novel 3D clustering agreement-based SSL task and masking embedding prediction inspired by pre-trained language models. Experiments on downstream tasks, such as 3D brain segmentation, lung nodule detection, 3D heart structures segmentation, and abnormal chest X-ray detection, demonstrate the effectiveness of our joint 2D and 3D SSL approach. We improve plain 2D Deep-ClusterV2 and SwAV by a significant margin and also surpass various modern 2D and 3D SSL approaches.
A Comparative Study of Graph Matching Algorithms in Computer Vision
Jul 01, 2022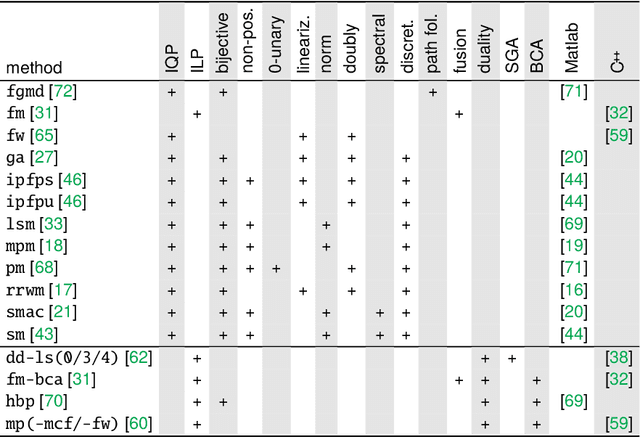
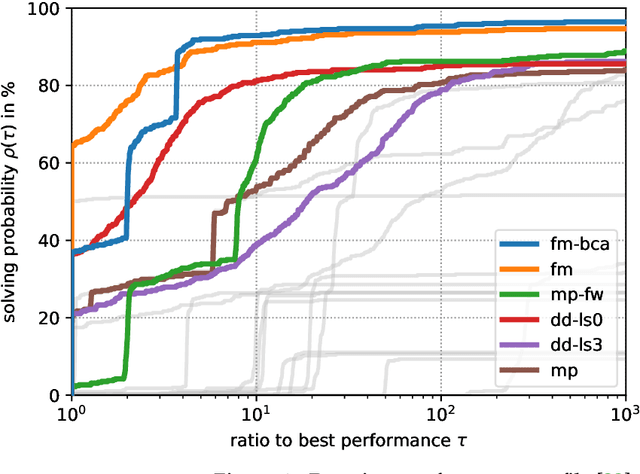

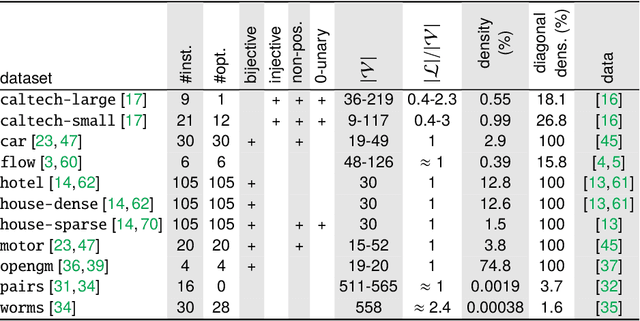
Abstract:The graph matching optimization problem is an essential component for many tasks in computer vision, such as bringing two deformable objects in correspondence. Naturally, a wide range of applicable algorithms have been proposed in the last decades. Since a common standard benchmark has not been developed, their performance claims are often hard to verify as evaluation on differing problem instances and criteria make the results incomparable. To address these shortcomings, we present a comparative study of graph matching algorithms. We create a uniform benchmark where we collect and categorize a large set of existing and publicly available computer vision graph matching problems in a common format. At the same time we collect and categorize the most popular open-source implementations of graph matching algorithms. Their performance is evaluated in a way that is in line with the best practices for comparing optimization algorithms. The study is designed to be reproducible and extensible to serve as a valuable resource in the future. Our study provides three notable insights: 1.) popular problem instances are exactly solvable in substantially less than 1 second and, therefore, are insufficient for future empirical evaluations; 2.) the most popular baseline methods are highly inferior to the best available methods; 3.) despite the NP-hardness of the problem, instances coming from vision applications are often solvable in a few seconds even for graphs with more than 500 vertices.
DOGE-Train: Discrete Optimization on GPU with End-to-end Training
May 23, 2022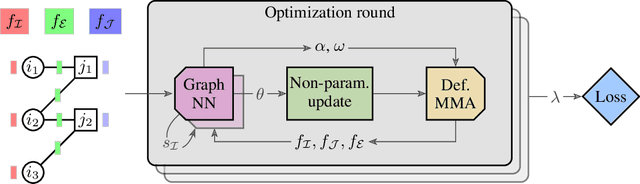
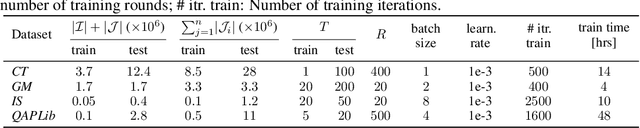
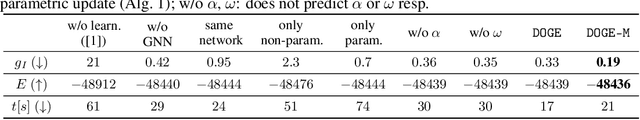
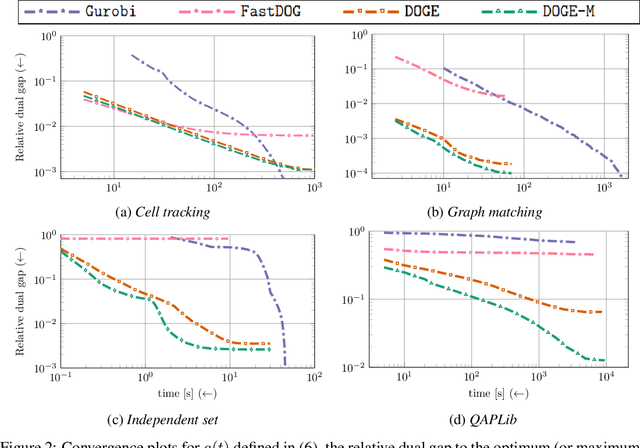
Abstract:We present a fast, scalable, data-driven approach for solving linear relaxations of 0-1 integer linear programs using a graph neural network. Our solver is based on the Lagrange decomposition based algorithm FastDOG (Abbas et al. (2022)). We make the algorithm differentiable and perform backpropagation through the dual update scheme for end-to-end training of its algorithmic parameters. This allows to preserve the algorithm's theoretical properties including feasibility and guaranteed non-decrease in the lower bound. Since FastDOG can get stuck in suboptimal fixed points, we provide additional freedom to our graph neural network to predict non-parametric update steps for escaping such points while maintaining dual feasibility. For training of the graph neural network we use an unsupervised loss and perform experiments on large-scale real world datasets. We train on smaller problems and test on larger ones showing strong generalization performance with a graph neural network comprising only around 10k parameters. Our solver achieves significantly faster performance and better dual objectives than its non-learned version. In comparison to commercial solvers our learned solver achieves close to optimal objective values of LP relaxations and is faster by up to an order of magnitude on very large problems from structured prediction and on selected combinatorial optimization problems.
A Scalable Combinatorial Solver for Elastic Geometrically Consistent 3D Shape Matching
Apr 27, 2022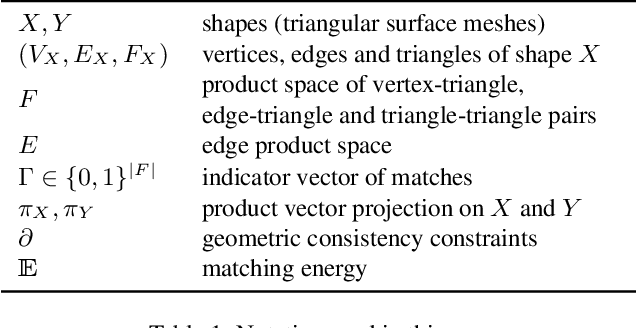
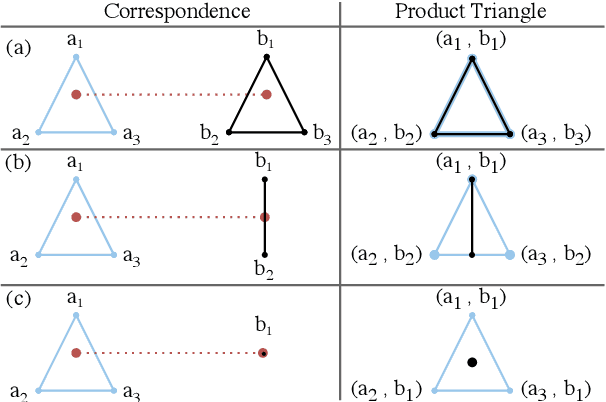
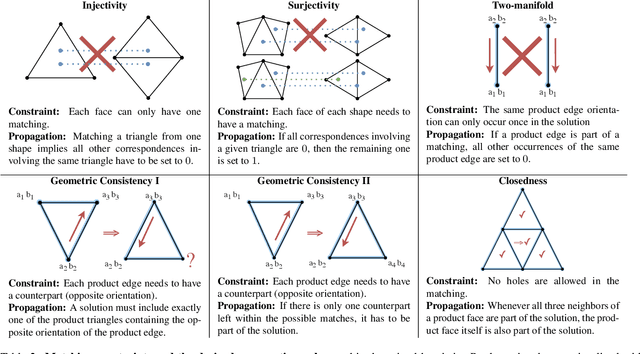
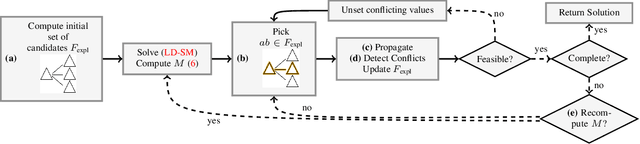
Abstract:We present a scalable combinatorial algorithm for globally optimizing over the space of geometrically consistent mappings between 3D shapes. We use the mathematically elegant formalism proposed by Windheuser et al. (ICCV 2011) where 3D shape matching was formulated as an integer linear program over the space of orientation-preserving diffeomorphisms. Until now, the resulting formulation had limited practical applicability due to its complicated constraint structure and its large size. We propose a novel primal heuristic coupled with a Lagrange dual problem that is several orders of magnitudes faster compared to previous solvers. This allows us to handle shapes with substantially more triangles than previously solvable. We demonstrate compelling results on diverse datasets, and, even showcase that we can address the challenging setting of matching two partial shapes without availability of complete shapes. Our code is publicly available at http://github.com/paul0noah/sm-comb .
Structured Prediction Problem Archive
Feb 04, 2022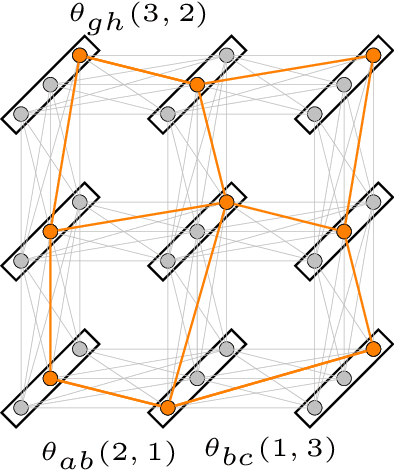
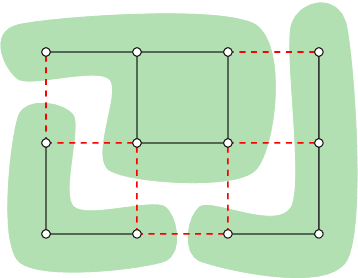
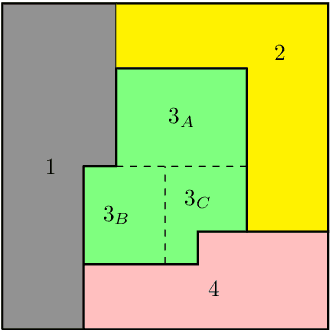
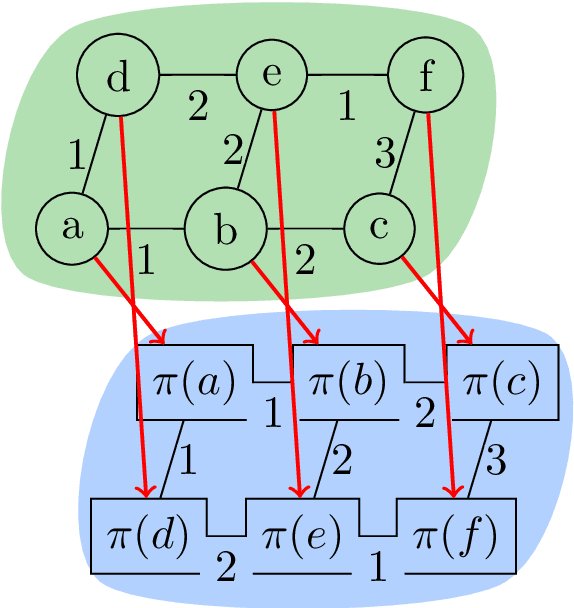
Abstract:Structured prediction problems are one of the fundamental tools in machine learning. In order to facilitate algorithm development for their numerical solution, we collect in one place a large number of datasets in easy to read formats for a diverse set of problem classes. We provide archival links to datasets, description of the considered problems and problem formats, and a short summary of problem characteristics including size, number of instances etc. For reference we also give a non-exhaustive selection of algorithms proposed in the literature for their solution. We hope that this central repository will make benchmarking and comparison to established works easier. We welcome submission of interesting new datasets and algorithms for inclusion in our archive.
LMGP: Lifted Multicut Meets Geometry Projections for Multi-Camera Multi-Object Tracking
Nov 23, 2021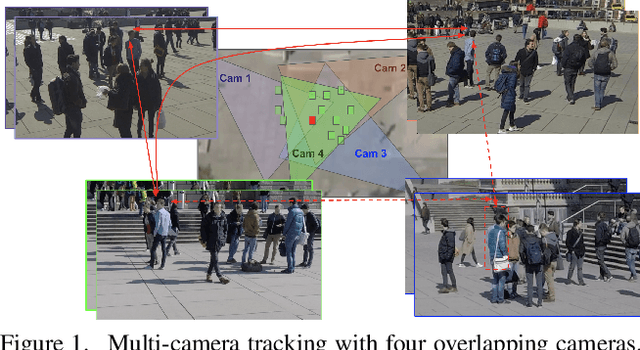
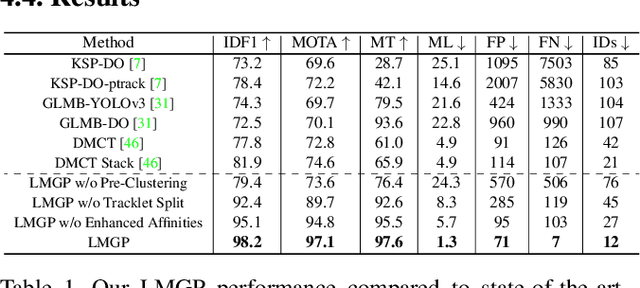
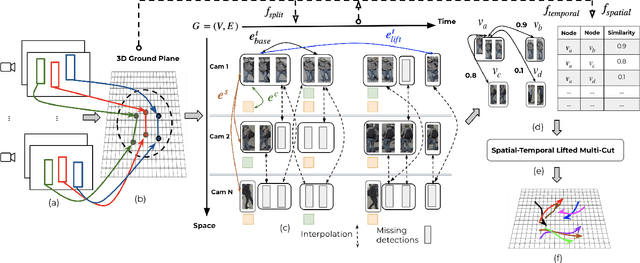
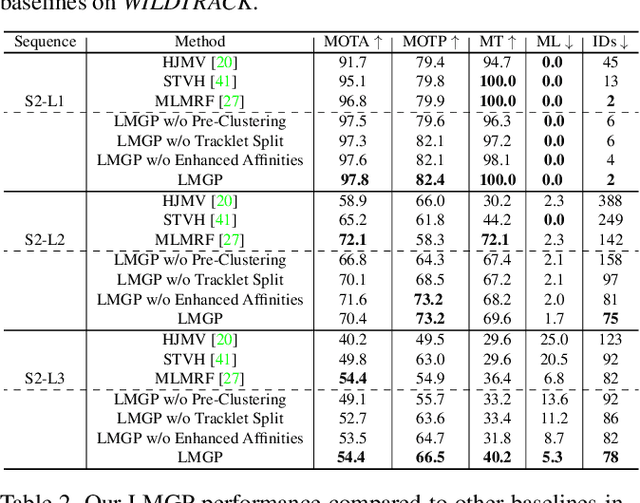
Abstract:Multi-Camera Multi-Object Tracking is currently drawing attention in the computer vision field due to its superior performance in real-world applications such as video surveillance with crowded scenes or in vast space. In this work, we propose a mathematically elegant multi-camera multiple object tracking approach based on a spatial-temporal lifted multicut formulation. Our model utilizes state-of-the-art tracklets produced by single-camera trackers as proposals. As these tracklets may contain ID-Switch errors, we refine them through a novel pre-clustering obtained from 3D geometry projections. As a result, we derive a better tracking graph without ID switches and more precise affinity costs for the data association phase. Tracklets are then matched to multi-camera trajectories by solving a global lifted multicut formulation that incorporates short and long-range temporal interactions on tracklets located in the same camera as well as inter-camera ones. Experimental results on the WildTrack dataset yield near-perfect result, outperforming state-of-the-art trackers on Campus while being on par on the PETS-09 dataset. We will make our implementations available upon acceptance of the paper.
FastDOG: Fast Discrete Optimization on GPU
Nov 19, 2021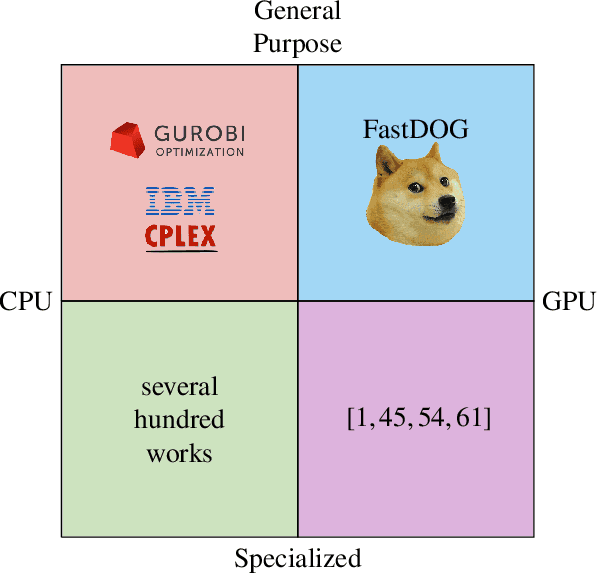
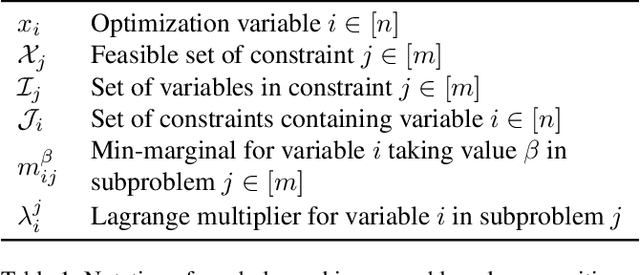

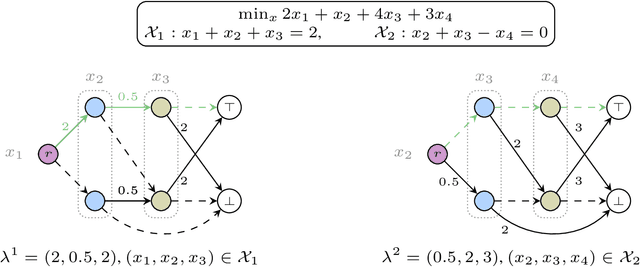
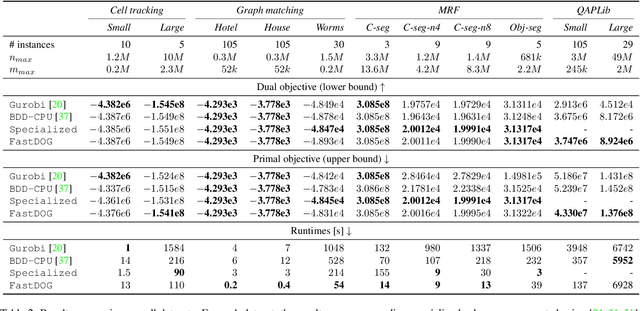
Abstract:We present a massively parallel Lagrange decomposition method for solving 0-1 integer linear programs occurring in structured prediction. We propose a new iterative update scheme for solving the Lagrangean dual and a perturbation technique for decoding primal solutions. For representing subproblems we follow Lange et al. (2021) and use binary decision diagrams (BDDs). Our primal and dual algorithms require little synchronization between subproblems and optimization over BDDs needs only elementary operations without complicated control flow. This allows us to exploit the parallelism offered by GPUs for all components of our method. We present experimental results on combinatorial problems from MAP inference for Markov Random Fields, quadratic assignment and cell tracking for developmental biology. Our highly parallel GPU implementation improves upon the running times of the algorithms from Lange et al. (2021) by up to an order of magnitude. In particular, we come close to or outperform some state-of-the-art specialized heuristics while being problem agnostic.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge