Paris Perdikaris
Scalable Bayesian optimization with high-dimensional outputs using randomized prior networks
Feb 14, 2023



Abstract:Several fundamental problems in science and engineering consist of global optimization tasks involving unknown high-dimensional (black-box) functions that map a set of controllable variables to the outcomes of an expensive experiment. Bayesian Optimization (BO) techniques are known to be effective in tackling global optimization problems using a relatively small number objective function evaluations, but their performance suffers when dealing with high-dimensional outputs. To overcome the major challenge of dimensionality, here we propose a deep learning framework for BO and sequential decision making based on bootstrapped ensembles of neural architectures with randomized priors. Using appropriate architecture choices, we show that the proposed framework can approximate functional relationships between design variables and quantities of interest, even in cases where the latter take values in high-dimensional vector spaces or even infinite-dimensional function spaces. In the context of BO, we augmented the proposed probabilistic surrogates with re-parameterized Monte Carlo approximations of multiple-point (parallel) acquisition functions, as well as methodological extensions for accommodating black-box constraints and multi-fidelity information sources. We test the proposed framework against state-of-the-art methods for BO and demonstrate superior performance across several challenging tasks with high-dimensional outputs, including a constrained optimization task involving shape optimization of rotor blades in turbo-machinery.
Random Weight Factorization Improves the Training of Continuous Neural Representations
Oct 05, 2022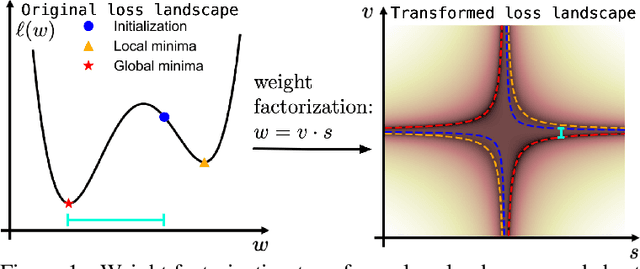
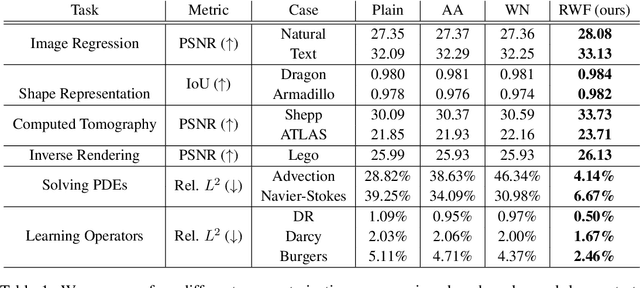
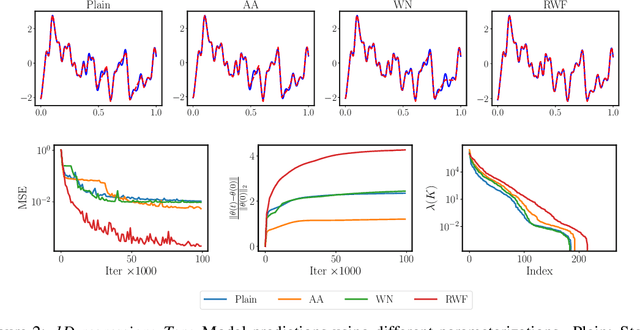
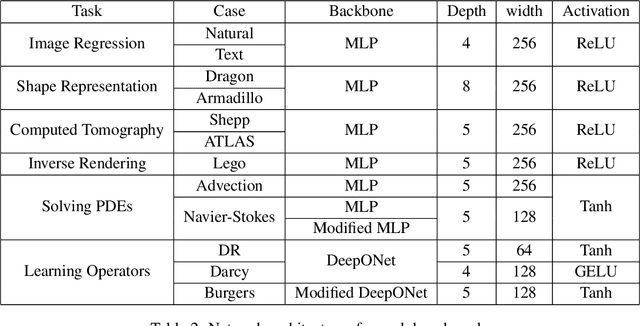
Abstract:Continuous neural representations have recently emerged as a powerful and flexible alternative to classical discretized representations of signals. However, training them to capture fine details in multi-scale signals is difficult and computationally expensive. Here we propose random weight factorization as a simple drop-in replacement for parameterizing and initializing conventional linear layers in coordinate-based multi-layer perceptrons (MLPs) that significantly accelerates and improves their training. We show how this factorization alters the underlying loss landscape and effectively enables each neuron in the network to learn using its own self-adaptive learning rate. This not only helps with mitigating spectral bias, but also allows networks to quickly recover from poor initializations and reach better local minima. We demonstrate how random weight factorization can be leveraged to improve the training of neural representations on a variety of tasks, including image regression, shape representation, computed tomography, inverse rendering, solving partial differential equations, and learning operators between function spaces.
$Δ$-PINNs: physics-informed neural networks on complex geometries
Sep 08, 2022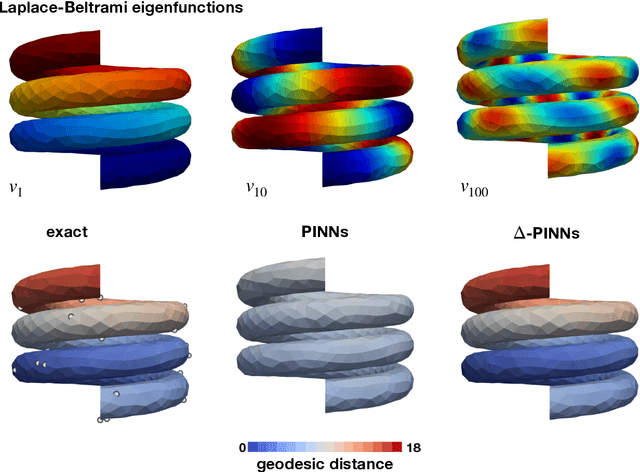
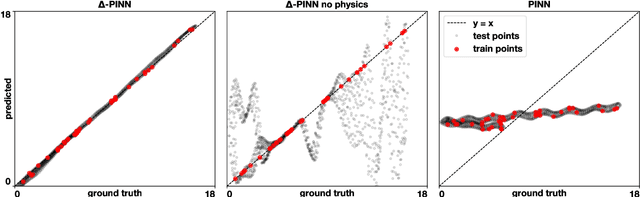
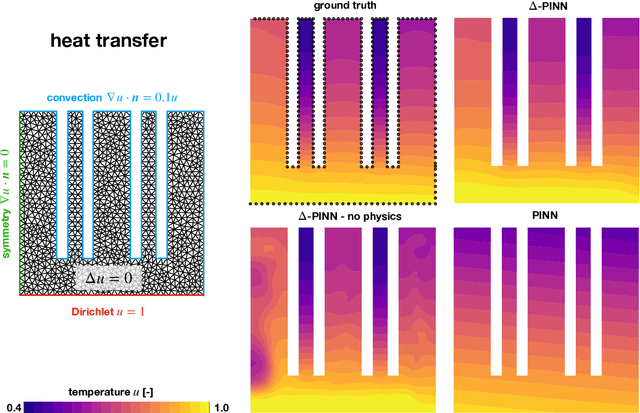
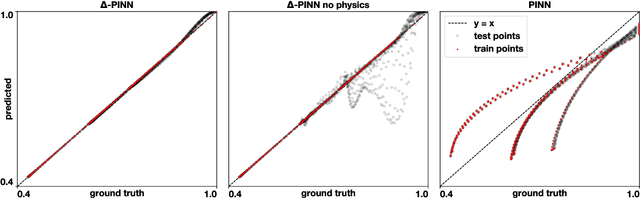
Abstract:Physics-informed neural networks (PINNs) have demonstrated promise in solving forward and inverse problems involving partial differential equations. Despite recent progress on expanding the class of problems that can be tackled by PINNs, most of existing use-cases involve simple geometric domains. To date, there is no clear way to inform PINNs about the topology of the domain where the problem is being solved. In this work, we propose a novel positional encoding mechanism for PINNs based on the eigenfunctions of the Laplace-Beltrami operator. This technique allows to create an input space for the neural network that represents the geometry of a given object. We approximate the eigenfunctions as well as the operators involved in the partial differential equations with finite elements. We extensively test and compare the proposed methodology against traditional PINNs in complex shapes, such as a coil, a heat sink and a bunny, with different physics, such as the Eikonal equation and heat transfer. We also study the sensitivity of our method to the number of eigenfunctions used, as well as the discretization used for the eigenfunctions and the underlying operators. Our results show excellent agreement with the ground truth data in cases where traditional PINNs fail to produce a meaningful solution. We envision this new technique will expand the effectiveness of PINNs to more realistic applications.
Semi-supervised Invertible DeepONets for Bayesian Inverse Problems
Sep 08, 2022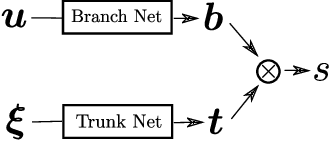
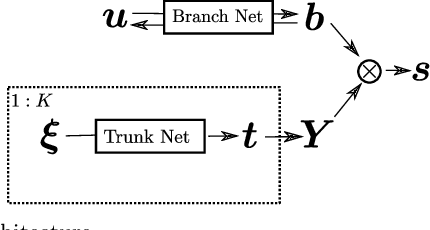
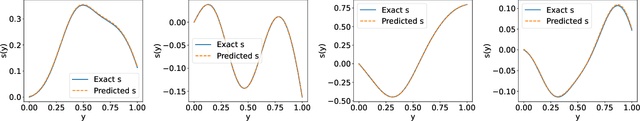
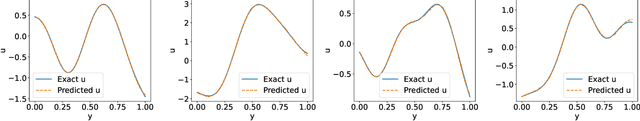
Abstract:Deep Operator Networks (DeepONets) offer a powerful, data-driven tool for solving parametric PDEs by learning operators, i.e. maps between infinite-dimensional function spaces. In this work, we employ physics-informed DeepONets in the context of high-dimensional, Bayesian inverse problems. Traditional solution strategies necessitate an enormous, and frequently infeasible, number of forward model solves, as well as the computation of parametric derivatives. In order to enable efficient solutions, we extend DeepONets by employing a realNVP architecture which yields an invertible and differentiable map between the parametric input and the branch net output. This allows us to construct accurate approximations of the full posterior which can be readily adapted irrespective of the number of observations and the magnitude of the observation noise. As a result, no additional forward solves are required, nor is there any need for costly sampling procedures. We demonstrate the efficacy and accuracy of the proposed methodology in the context of inverse problems based on a anti-derivative, a reaction-diffusion and a Darcy-flow equation.
Rethinking the Importance of Sampling in Physics-informed Neural Networks
Jul 05, 2022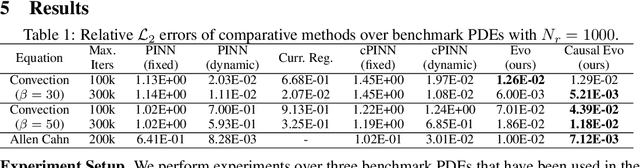
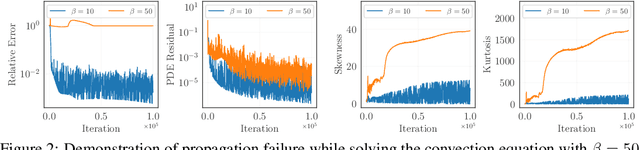
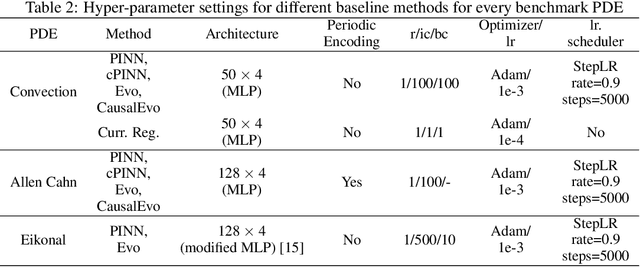
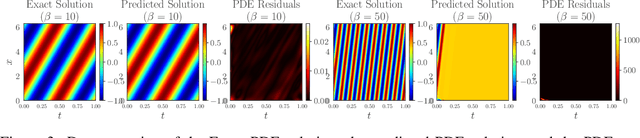
Abstract:Physics-informed neural networks (PINNs) have emerged as a powerful tool for solving partial differential equations (PDEs) in a variety of domains. While previous research in PINNs has mainly focused on constructing and balancing loss functions during training to avoid poor minima, the effect of sampling collocation points on the performance of PINNs has largely been overlooked. In this work, we find that the performance of PINNs can vary significantly with different sampling strategies, and using a fixed set of collocation points can be quite detrimental to the convergence of PINNs to the correct solution. In particular, (1) we hypothesize that training of PINNs rely on successful "propagation" of solution from initial and/or boundary condition points to interior points, and PINNs with poor sampling strategies can get stuck at trivial solutions if there are \textit{propagation failures}. (2) We demonstrate that propagation failures are characterized by highly imbalanced PDE residual fields where very high residuals are observed over very narrow regions. (3) To mitigate propagation failure, we propose a novel \textit{evolutionary sampling} (Evo) method that can incrementally accumulate collocation points in regions of high PDE residuals. We further provide an extension of Evo to respect the principle of causality while solving time-dependent PDEs. We empirically demonstrate the efficacy and efficiency of our proposed methods in a variety of PDE problems.
NOMAD: Nonlinear Manifold Decoders for Operator Learning
Jun 07, 2022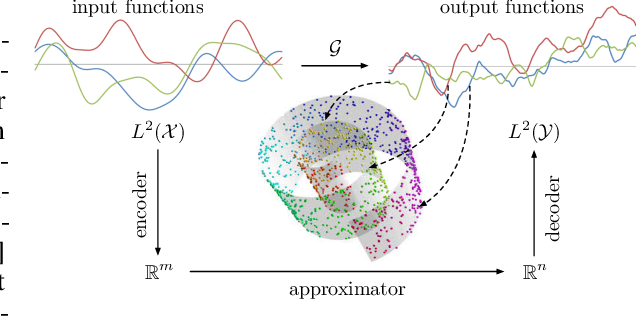
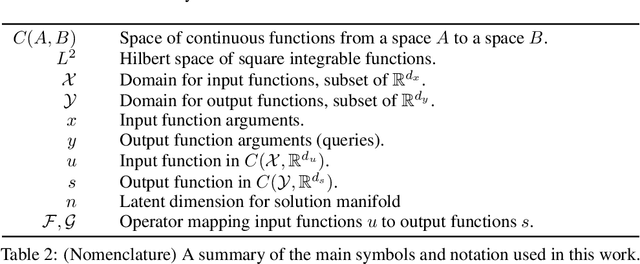
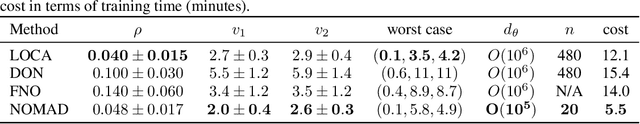
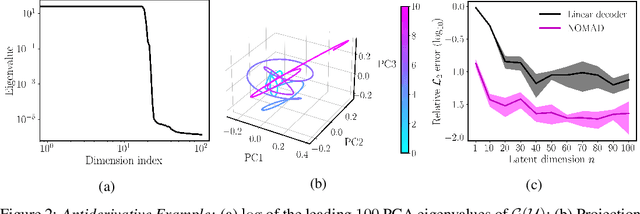

Abstract:Supervised learning in function spaces is an emerging area of machine learning research with applications to the prediction of complex physical systems such as fluid flows, solid mechanics, and climate modeling. By directly learning maps (operators) between infinite dimensional function spaces, these models are able to learn discretization invariant representations of target functions. A common approach is to represent such target functions as linear combinations of basis elements learned from data. However, there are simple scenarios where, even though the target functions form a low dimensional submanifold, a very large number of basis elements is needed for an accurate linear representation. Here we present NOMAD, a novel operator learning framework with a nonlinear decoder map capable of learning finite dimensional representations of nonlinear submanifolds in function spaces. We show this method is able to accurately learn low dimensional representations of solution manifolds to partial differential equations while outperforming linear models of larger size. Additionally, we compare to state-of-the-art operator learning methods on a complex fluid dynamics benchmark and achieve competitive performance with a significantly smaller model size and training cost.
Respecting causality is all you need for training physics-informed neural networks
Mar 14, 2022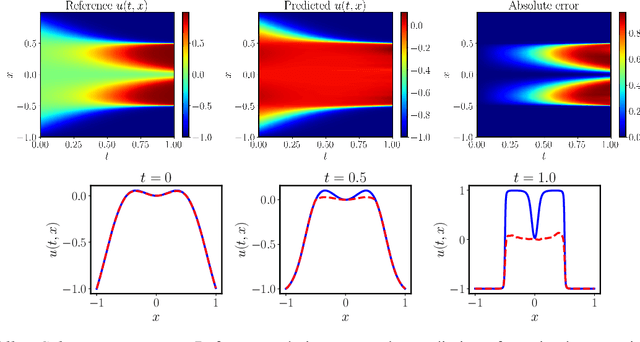
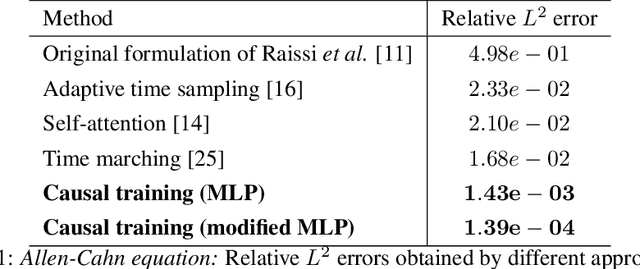
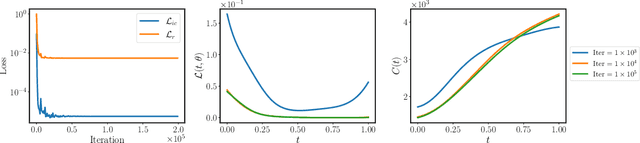
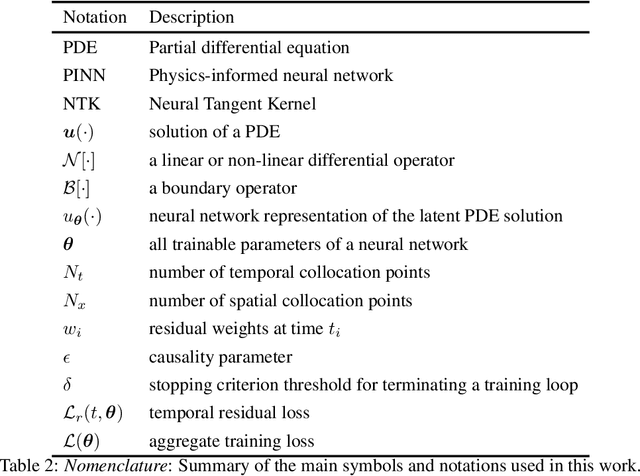
Abstract:While the popularity of physics-informed neural networks (PINNs) is steadily rising, to this date PINNs have not been successful in simulating dynamical systems whose solution exhibits multi-scale, chaotic or turbulent behavior. In this work we attribute this shortcoming to the inability of existing PINNs formulations to respect the spatio-temporal causal structure that is inherent to the evolution of physical systems. We argue that this is a fundamental limitation and a key source of error that can ultimately steer PINN models to converge towards erroneous solutions. We address this pathology by proposing a simple re-formulation of PINNs loss functions that can explicitly account for physical causality during model training. We demonstrate that this simple modification alone is enough to introduce significant accuracy improvements, as well as a practical quantitative mechanism for assessing the convergence of a PINNs model. We provide state-of-the-art numerical results across a series of benchmarks for which existing PINNs formulations fail, including the chaotic Lorenz system, the Kuramoto-Sivashinsky equation in the chaotic regime, and the Navier-Stokes equations in the turbulent regime. To the best of our knowledge, this is the first time that PINNs have been successful in simulating such systems, introducing new opportunities for their applicability to problems of industrial complexity.
Learning cardiac activation maps from 12-lead ECG with multi-fidelity Bayesian optimization on manifolds
Mar 11, 2022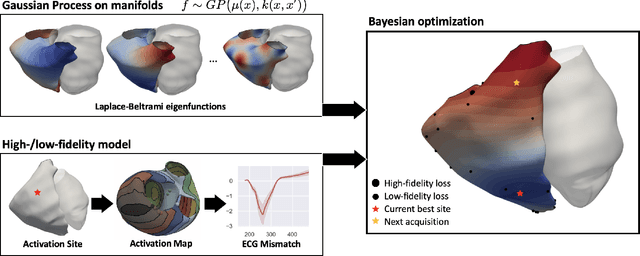
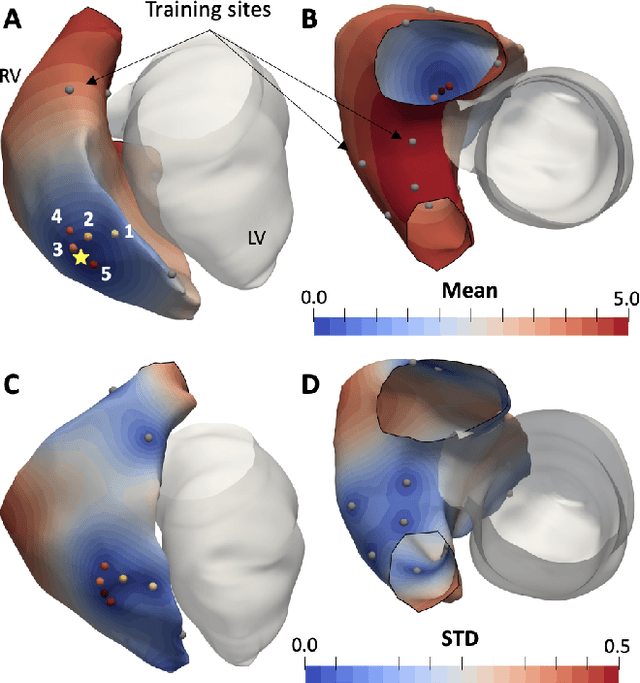
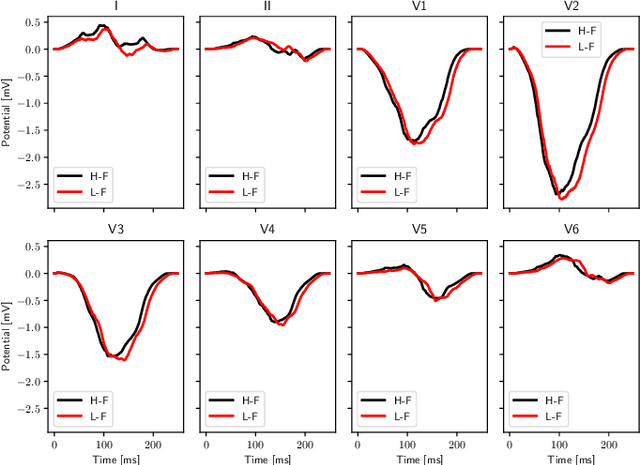
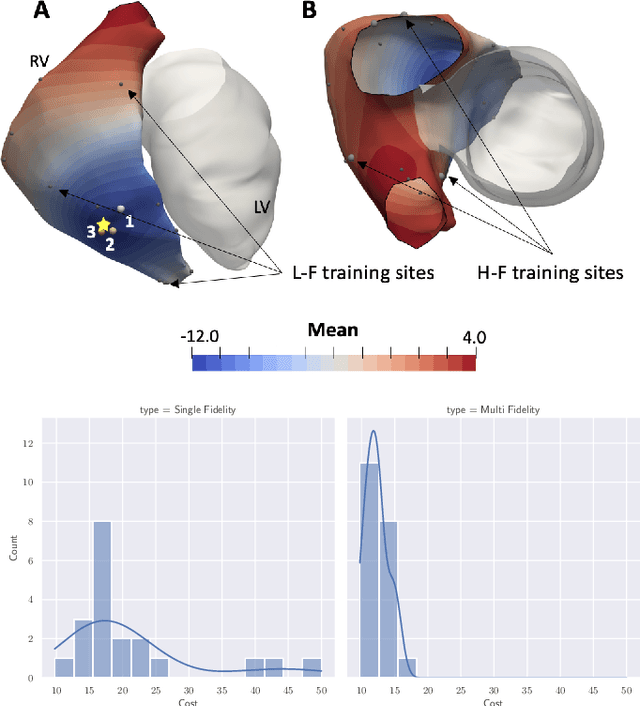
Abstract:We propose a method for identifying an ectopic activation in the heart non-invasively. Ectopic activity in the heart can trigger deadly arrhythmias. The localization of the ectopic foci or earliest activation sites (EASs) is therefore a critical information for cardiologists in deciding the optimal treatment. In this work, we formulate the identification problem as a global optimization problem, by minimizing the mismatch between the ECG predicted by a cardiac model, when paced at a given EAS, and the observed ECG during the ectopic activity. Our cardiac model amounts at solving an anisotropic eikonal equation for cardiac activation and the forward bidomain model in the torso with the lead field approach for computing the ECG. We build a Gaussian process surrogate model of the loss function on the heart surface to perform Bayesian optimization. In this procedure, we iteratively evaluate the loss function following the lower confidence bound criterion, which combines exploring the surface with exploitation of the minimum region. We also extend this framework to incorporate multiple levels of fidelity of the model. We show that our procedure converges to the minimum only after $11.7\pm10.4$ iterations (20 independent runs) for the single-fidelity case and $3.5\pm1.7$ iterations for the multi-fidelity case. We envision that this tool could be applied in real time in a clinical setting to identify potentially dangerous EASs.
Scalable Uncertainty Quantification for Deep Operator Networks using Randomized Priors
Mar 06, 2022
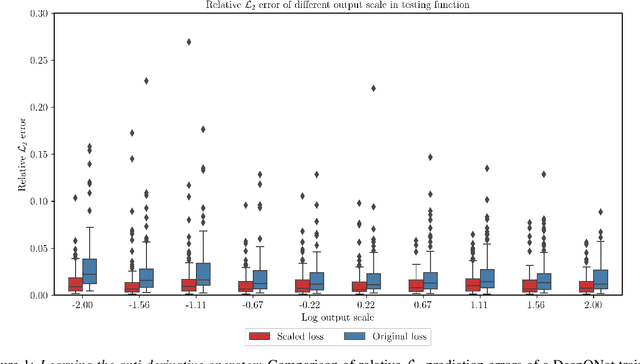
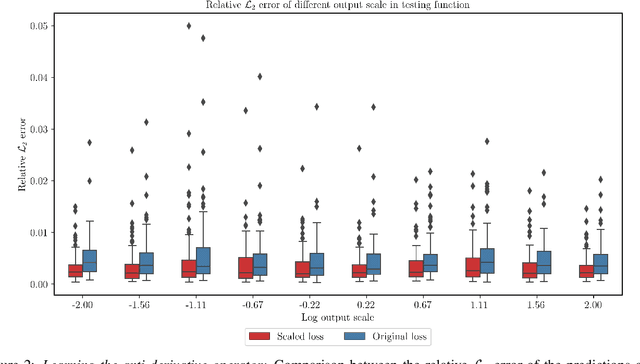
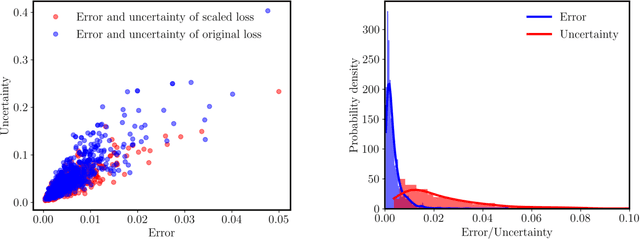
Abstract:We present a simple and effective approach for posterior uncertainty quantification in deep operator networks (DeepONets); an emerging paradigm for supervised learning in function spaces. We adopt a frequentist approach based on randomized prior ensembles, and put forth an efficient vectorized implementation for fast parallel inference on accelerated hardware. Through a collection of representative examples in computational mechanics and climate modeling, we show that the merits of the proposed approach are fourfold. (1) It can provide more robust and accurate predictions when compared against deterministic DeepONets. (2) It shows great capability in providing reliable uncertainty estimates on scarce data-sets with multi-scale function pairs. (3) It can effectively detect out-of-distribution and adversarial examples. (4) It can seamlessly quantify uncertainty due to model bias, as well as noise corruption in the data. Finally, we provide an optimized JAX library called {\em UQDeepONet} that can accommodate large model architectures, large ensemble sizes, as well as large data-sets with excellent parallel performance on accelerated hardware, thereby enabling uncertainty quantification for DeepONets in realistic large-scale applications.
Physics-informed neural networks to learn cardiac fiber orientation from multiple electroanatomical maps
Feb 01, 2022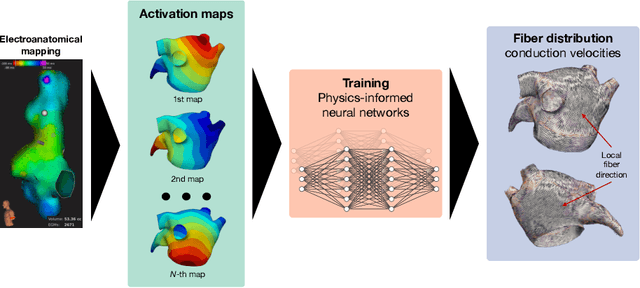

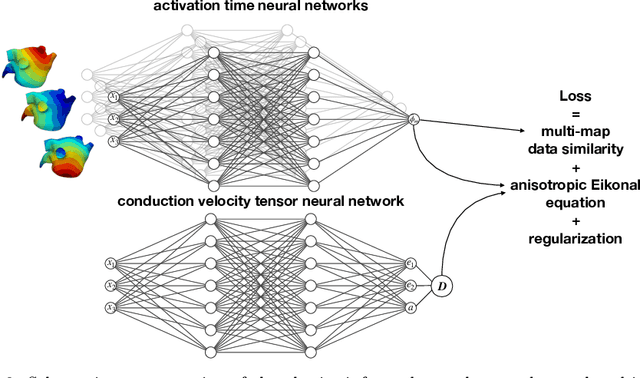
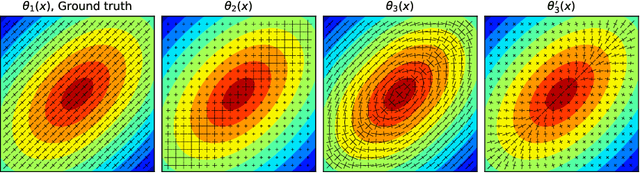
Abstract:We propose FiberNet, a method to estimate in-vivo the cardiac fiber architecture of the human atria from multiple catheter recordings of the electrical activation. Cardiac fibers play a central rolein the electro-mechanical function of the heart, yet they aredifficult to determine in-vivo, and hence rarely truly patient-specificin existing cardiac models.FiberNet learns the fibers arrangement by solvingan inverse problem with physics-informed neural networks. The inverse problem amounts to identifyingthe conduction velocity tensor of a cardiac propagation modelfrom a set of sparse activation maps. The use of multiple mapsenables the simultaneous identification of all the componentsof the conduction velocity tensor, including the local fiber angle.We extensively test FiberNet on synthetic 2-D and 3-D examples, diffusion tensor fibers, and a patient-specific case. We show that 3 maps are sufficient to accurately capture the fibers, also in thepresence of noise. With fewer maps, the role of regularization becomesprominent. Moreover, we show that the fitted model can robustlyreproduce unseen activation maps. We envision that FiberNet will help the creation of patient-specific models for personalized medicine.The full code is available at http://github.com/fsahli/FiberNet.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge