Ozlem Uzuner
Progress Note Understanding -- Assessment and Plan Reasoning: Overview of the 2022 N2C2 Track 3 Shared Task
Mar 14, 2023



Abstract:Daily progress notes are common types in the electronic health record (EHR) where healthcare providers document the patient's daily progress and treatment plans. The EHR is designed to document all the care provided to patients, but it also enables note bloat with extraneous information that distracts from the diagnoses and treatment plans. Applications of natural language processing (NLP) in the EHR is a growing field with the majority of methods in information extraction. Few tasks use NLP methods for downstream diagnostic decision support. We introduced the 2022 National NLP Clinical Challenge (N2C2) Track 3: Progress Note Understanding - Assessment and Plan Reasoning as one step towards a new suite of tasks. The Assessment and Plan Reasoning task focuses on the most critical components of progress notes, Assessment and Plan subsections where health problems and diagnoses are contained. The goal of the task was to develop and evaluate NLP systems that automatically predict causal relations between the overall status of the patient contained in the Assessment section and its relation to each component of the Plan section which contains the diagnoses and treatment plans. The goal of the task was to identify and prioritize diagnoses as the first steps in diagnostic decision support to find the most relevant information in long documents like daily progress notes. We present the results of 2022 n2c2 Track 3 and provide a description of the data, evaluation, participation and system performance.
The Leaf Clinical Trials Corpus: a new resource for query generation from clinical trial eligibility criteria
Jul 27, 2022Abstract:Identifying cohorts of patients based on eligibility criteria such as medical conditions, procedures, and medication use is critical to recruitment for clinical trials. Such criteria are often most naturally described in free-text, using language familiar to clinicians and researchers. In order to identify potential participants at scale, these criteria must first be translated into queries on clinical databases, which can be labor-intensive and error-prone. Natural language processing (NLP) methods offer a potential means of such conversion into database queries automatically. However they must first be trained and evaluated using corpora which capture clinical trials criteria in sufficient detail. In this paper, we introduce the Leaf Clinical Trials (LCT) corpus, a human-annotated corpus of over 1,000 clinical trial eligibility criteria descriptions using highly granular structured labels capturing a range of biomedical phenomena. We provide details of our schema, annotation process, corpus quality, and statistics. Additionally, we present baseline information extraction results on this corpus as benchmarks for future work.
Online User Profiling to Detect Social Bots on Twitter
Mar 09, 2022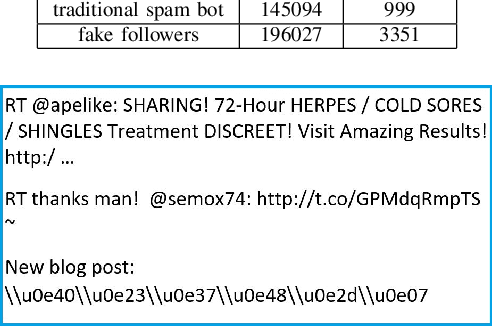
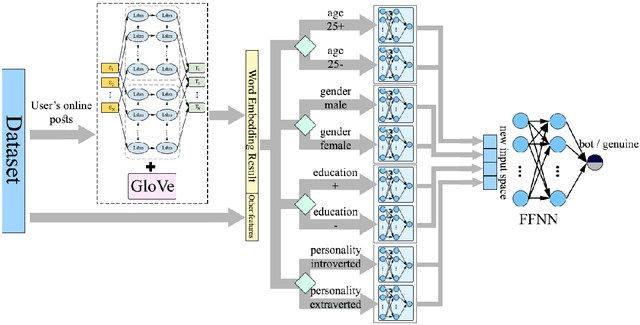
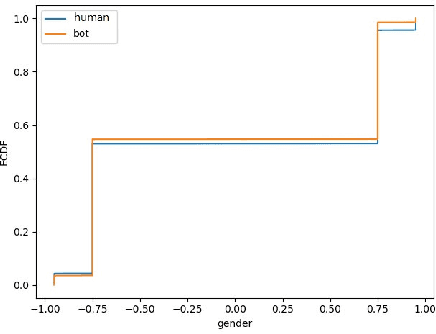
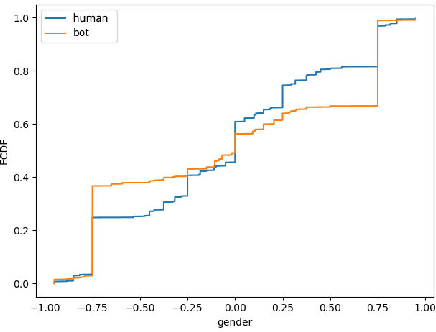
Abstract:Social media platforms can expose influential trends in many aspects of everyday life. However, the movements they represent can be contaminated by disinformation. Social bots are one of the significant sources of disinformation in social media. Social bots can pose serious cyber threats to society and public opinion. This research aims to develop machine learning models to detect bots based on the extracted user's profile from a Tweet's text. Online users' profile shows the user's personal information, such as age, gender, education, and personality. In this work, the user's profile is constructed based on the user's online posts. This work's main contribution is three-fold: First, we aim to improve bot detection through machine learning models based on the user's personal information generated by the user's online comments. When comparing two online posts, the similarity of personal information makes it difficult to differentiate a bot from a human user. However, this research turns personal information similarity among two online posts into an advantage for the new bot detection model. The new proposed model for bot detection creates user profiles based on personal information such as age, personality, gender, education from users' online posts and introduces a machine learning model to detect social bots with high prediction accuracy based on personal information. Second, create a new public data set that shows the user's profile for more than 6900 Twitter accounts in the Cresci 2017 data set.
A Scoping Review of Publicly Available Language Tasks in Clinical Natural Language Processing
Dec 07, 2021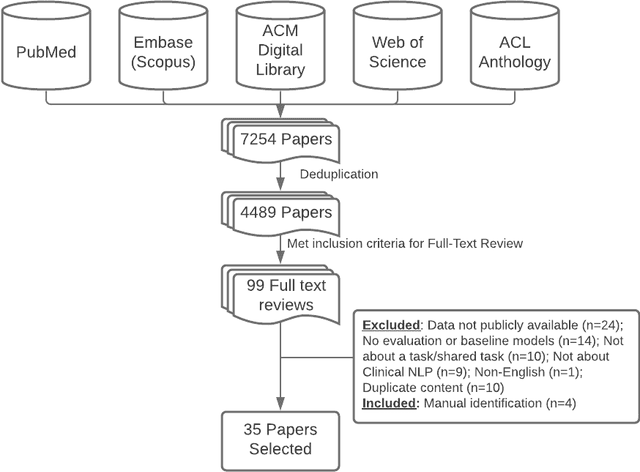
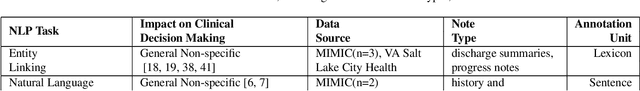
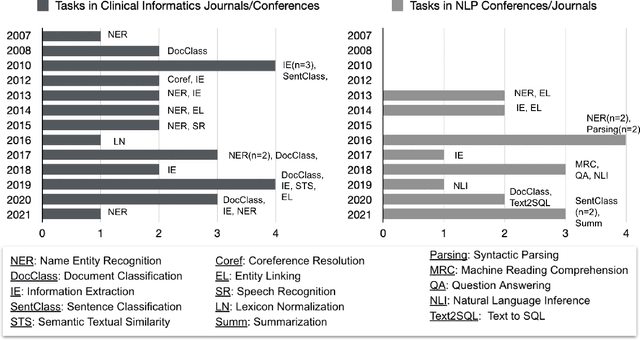

Abstract:Objective: to provide a scoping review of papers on clinical natural language processing (NLP) tasks that use publicly available electronic health record data from a cohort of patients. Materials and Methods: We searched six databases, including biomedical research and computer science literature database. A round of title/abstract screening and full-text screening were conducted by two reviewers. Our method followed the Preferred Reporting Items for Systematic Reviews and Meta-Analysis (PRISMA) guidelines. Results: A total of 35 papers with 47 clinical NLP tasks met inclusion criteria between 2007 and 2021. We categorized the tasks by the type of NLP problems, including name entity recognition, summarization, and other NLP tasks. Some tasks were introduced with a topic of clinical decision support applications, such as substance abuse, phenotyping, cohort selection for clinical trial. We summarized the tasks by publication and dataset information. Discussion: The breadth of clinical NLP tasks keeps growing as the field of NLP evolves with advancements in language systems. However, gaps exist in divergent interests between general domain NLP community and clinical informatics community, and in generalizability of the data sources. We also identified issues in data selection and preparation including the lack of time-sensitive data, and invalidity of problem size and evaluation. Conclusions: The existing clinical NLP tasks cover a wide range of topics and the field will continue to grow and attract more attention from both general domain NLP and clinical informatics community. We encourage future work to incorporate multi-disciplinary collaboration, reporting transparency, and standardization in data preparation.
Extracting Radiological Findings With Normalized Anatomical Information Using a Span-Based BERT Relation Extraction Model
Aug 20, 2021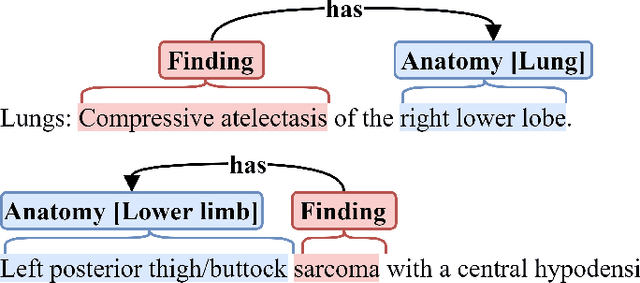
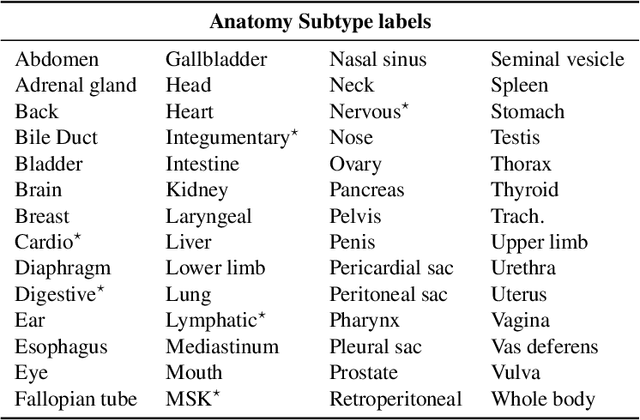
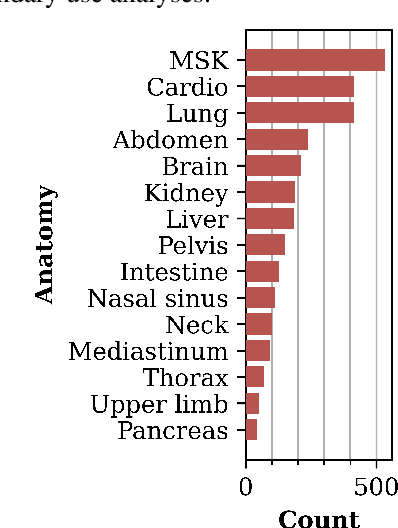
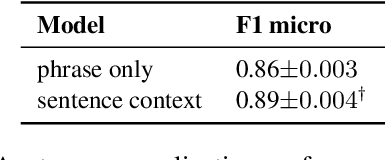
Abstract:Medical imaging is critical to the diagnosis and treatment of numerous medical problems, including many forms of cancer. Medical imaging reports distill the findings and observations of radiologists, creating an unstructured textual representation of unstructured medical images. Large-scale use of this text-encoded information requires converting the unstructured text to a structured, semantic representation. We explore the extraction and normalization of anatomical information in radiology reports that is associated with radiological findings. We investigate this extraction and normalization task using a span-based relation extraction model that jointly extracts entities and relations using BERT. This work examines the factors that influence extraction and normalization performance, including the body part/organ system, frequency of occurrence, span length, and span diversity. It discusses approaches for improving performance and creating high-quality semantic representations of radiological phenomena.
Jointly Learning Clinical Entities and Relations with Contextual Language Models and Explicit Context
Feb 17, 2021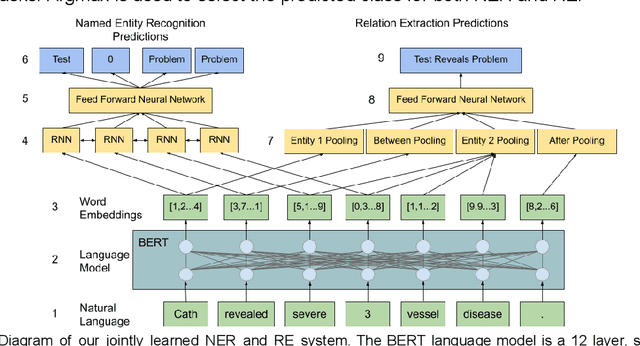
Abstract:We hypothesize that explicit integration of contextual information into an Multi-task Learning framework would emphasize the significance of context for boosting performance in jointly learning Named Entity Recognition (NER) and Relation Extraction (RE). Our work proves this hypothesis by segmenting entities from their surrounding context and by building contextual representations using each independent segment. This relation representation allows for a joint NER/RE system that achieves near state-of-the-art (SOTA) performance on both NER and RE tasks while beating the SOTA RE system at end-to-end NER & RE with a 49.07 F1.
Transfer Learning Approach for Arabic Offensive Language Detection System -- BERT-Based Model
Feb 09, 2021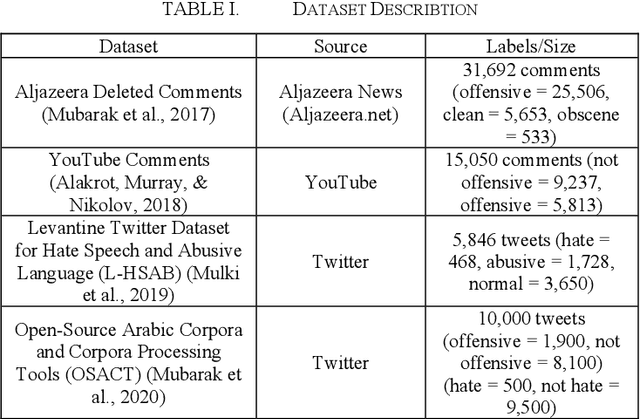
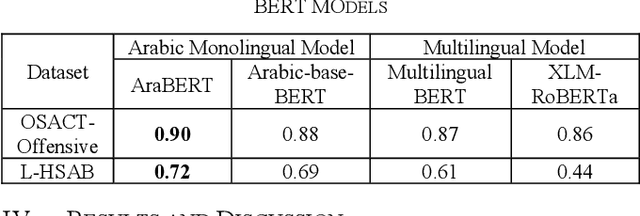
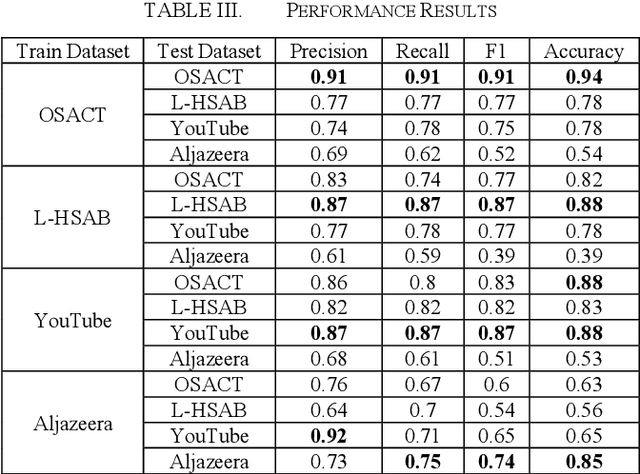
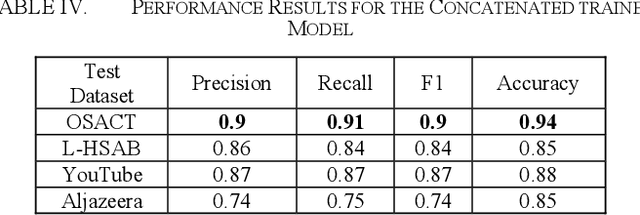
Abstract:Developing a system to detect online offensive language is very important to the health and the security of online users. Studies have shown that cyberhate, online harassment and other misuses of technology are on the rise, particularly during the global Coronavirus pandemic in 2020. According to the latest report by the Anti-Defamation League (ADL), 35% of online users reported online harassment related to their identity-based characteristics, which is a 3% increase over 2019. Applying advanced techniques from the Natural Language Processing (NLP) field to support the development of an online hate-free community is a critical task for social justice. Transfer learning enhances the performance of the classifier by allowing the transfer of knowledge from one domain or one dataset to others that have not been seen before, thus, supporting the classifier to be more generalizable. In our study, we apply the principles of transfer learning cross multiple Arabic offensive language datasets to compare the effects on system performance. This study aims at investigating the effects of fine-tuning and training Bidirectional Encoder Representations from Transformers (BERT) model on multiple Arabic offensive language datasets individually and testing it using other datasets individually. Our experiment starts with a comparison among multiple BERT models to guide the selection of the main model that is used for our study. The study also investigates the effects of concatenating all datasets to be used for fine-tuning and training BERT model. Our results demonstrate the limited effects of transfer learning on the performance of the classifiers, particularly for highly dialectic comments.
Exploratory Arabic Offensive Language Dataset Analysis
Jan 20, 2021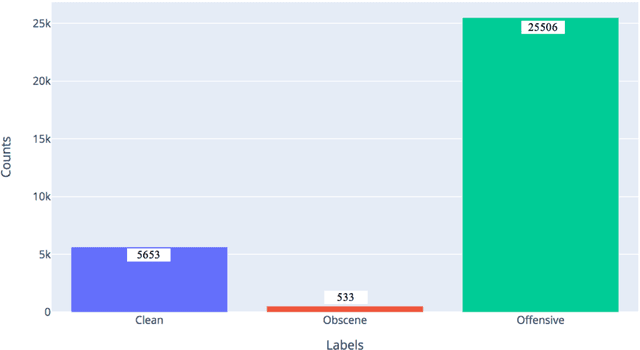
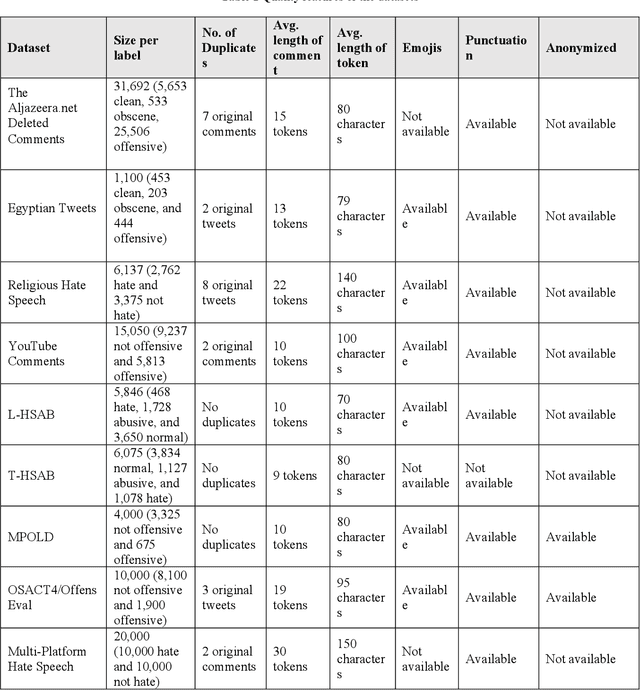

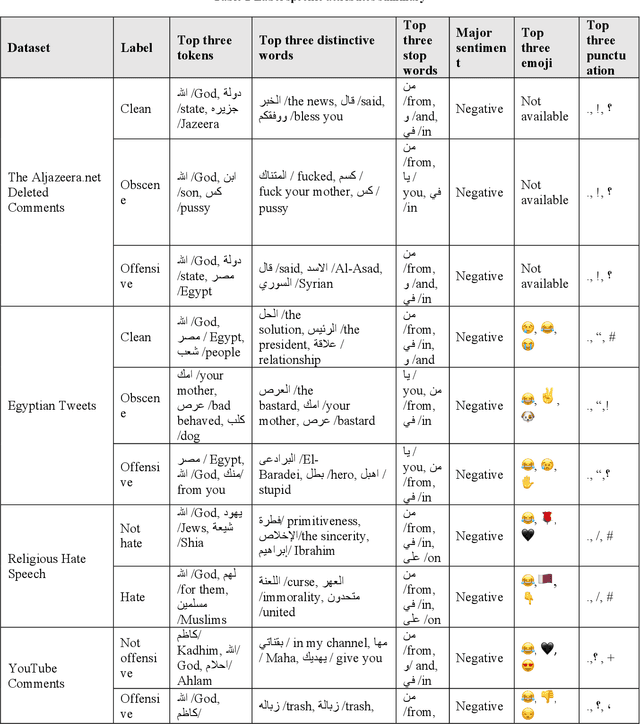
Abstract:This paper adding more insights towards resources and datasets used in Arabic offensive language research. The main goal of this paper is to guide researchers in Arabic offensive language in selecting appropriate datasets based on their content, and in creating new Arabic offensive language resources to support and complement the available ones.
SalamNET at SemEval-2020 Task12: Deep Learning Approach for Arabic Offensive Language Detection
Jul 28, 2020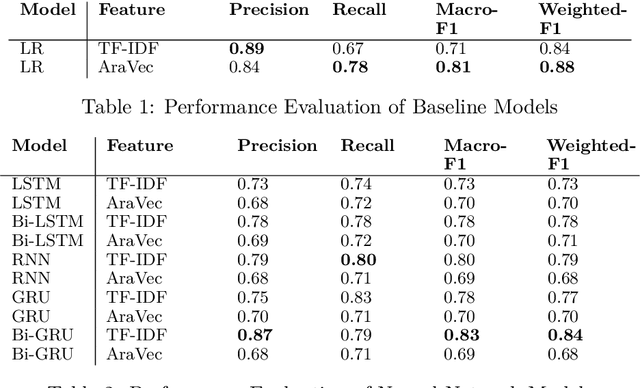
Abstract:This paper describes SalamNET, an Arabic offensive language detection system that has been submitted to SemEval 2020 shared task 12: Multilingual Offensive Language Identification in Social Media. Our approach focuses on applying multiple deep learning models and conducting in depth error analysis of results to provide system implications for future development considerations. To pursue our goal, a Recurrent Neural Network (RNN), a Gated Recurrent Unit (GRU), and Long-Short Term Memory (LSTM) models with different design architectures have been developed and evaluated. The SalamNET, a Bi-directional Gated Recurrent Unit (Bi-GRU) based model, reports a macro-F1 score of 0.83.
Extraction and Analysis of Clinically Important Follow-up Recommendations in a Large Radiology Dataset
May 14, 2019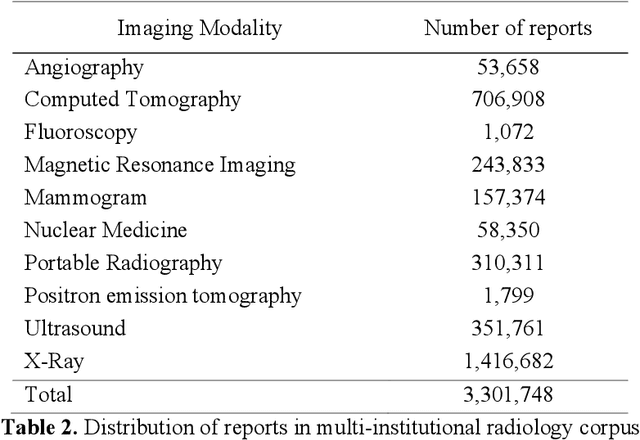
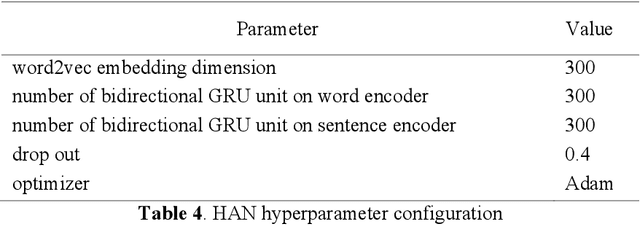
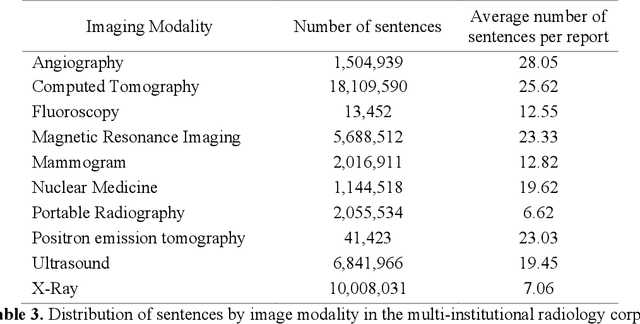

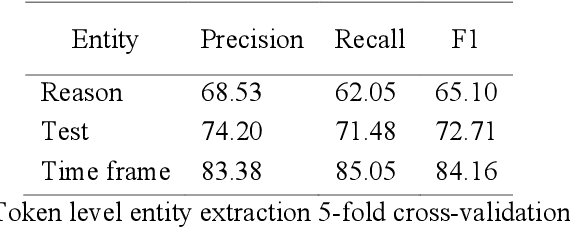
Abstract:Communication of follow-up recommendations when abnormalities are identified on imaging studies is prone to error. In this paper, we present a natural language processing approach based on deep learning to automatically identify clinically important recommendations in radiology reports. Our approach first identifies the recommendation sentences and then extracts reason, test, and time frame of the identified recommendations. To train our extraction models, we created a corpus of 567 radiology reports annotated for recommendation information. Our extraction models achieved 0.92 f-score for recommendation sentence, 0.65 f-score for reason, 0.73 f-score for test, and 0.84 f-score for time frame. We applied the extraction models to a set of over 3.3 million radiology reports and analyzed the adherence of follow-up recommendations.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge