Omer Antverg
A Content-Based Framework for Cybersecurity Refusal Decisions in Large Language Models
Feb 17, 2026Abstract:Large language models and LLM-based agents are increasingly used for cybersecurity tasks that are inherently dual-use. Existing approaches to refusal, spanning academic policy frameworks and commercially deployed systems, often rely on broad topic-based bans or offensive-focused taxonomies. As a result, they can yield inconsistent decisions, over-restrict legitimate defenders, and behave brittlely under obfuscation or request segmentation. We argue that effective refusal requires explicitly modeling the trade-off between offensive risk and defensive benefit, rather than relying solely on intent or offensive classification. In this paper, we introduce a content-based framework for designing and auditing cyber refusal policies that makes offense-defense tradeoffs explicit. The framework characterizes requests along five dimensions: Offensive Action Contribution, Offensive Risk, Technical Complexity, Defensive Benefit, and Expected Frequency for Legitimate Users, grounded in the technical substance of the request rather than stated intent. We demonstrate that this content-grounded approach resolves inconsistencies in current frontier model behavior and allows organizations to construct tunable, risk-aware refusal policies.
Jamba-1.5: Hybrid Transformer-Mamba Models at Scale
Aug 22, 2024



Abstract:We present Jamba-1.5, new instruction-tuned large language models based on our Jamba architecture. Jamba is a hybrid Transformer-Mamba mixture of experts architecture, providing high throughput and low memory usage across context lengths, while retaining the same or better quality as Transformer models. We release two model sizes: Jamba-1.5-Large, with 94B active parameters, and Jamba-1.5-Mini, with 12B active parameters. Both models are fine-tuned for a variety of conversational and instruction-following capabilties, and have an effective context length of 256K tokens, the largest amongst open-weight models. To support cost-effective inference, we introduce ExpertsInt8, a novel quantization technique that allows fitting Jamba-1.5-Large on a machine with 8 80GB GPUs when processing 256K-token contexts without loss of quality. When evaluated on a battery of academic and chatbot benchmarks, Jamba-1.5 models achieve excellent results while providing high throughput and outperforming other open-weight models on long-context benchmarks. The model weights for both sizes are publicly available under the Jamba Open Model License and we release ExpertsInt8 as open source.
BLOOM: A 176B-Parameter Open-Access Multilingual Language Model
Nov 09, 2022Abstract:Large language models (LLMs) have been shown to be able to perform new tasks based on a few demonstrations or natural language instructions. While these capabilities have led to widespread adoption, most LLMs are developed by resource-rich organizations and are frequently kept from the public. As a step towards democratizing this powerful technology, we present BLOOM, a 176B-parameter open-access language model designed and built thanks to a collaboration of hundreds of researchers. BLOOM is a decoder-only Transformer language model that was trained on the ROOTS corpus, a dataset comprising hundreds of sources in 46 natural and 13 programming languages (59 in total). We find that BLOOM achieves competitive performance on a wide variety of benchmarks, with stronger results after undergoing multitask prompted finetuning. To facilitate future research and applications using LLMs, we publicly release our models and code under the Responsible AI License.
IDANI: Inference-time Domain Adaptation via Neuron-level Interventions
Jun 01, 2022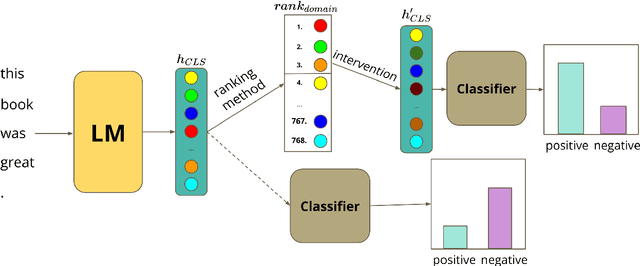
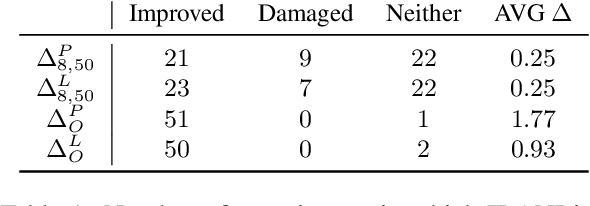

Abstract:Large pre-trained models are usually fine-tuned on downstream task data, and tested on unseen data. When the train and test data come from different domains, the model is likely to struggle, as it is not adapted to the test domain. We propose a new approach for domain adaptation (DA), using neuron-level interventions: We modify the representation of each test example in specific neurons, resulting in a counterfactual example from the source domain, which the model is more familiar with. The modified example is then fed back into the model. While most other DA methods are applied during training time, ours is applied during inference only, making it more efficient and applicable. Our experiments show that our method improves performance on unseen domains.
On the Pitfalls of Analyzing Individual Neurons in Language Models
Oct 14, 2021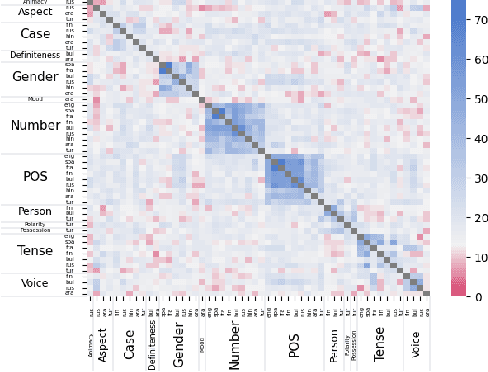
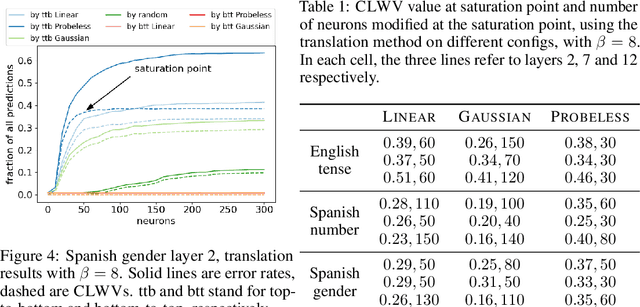
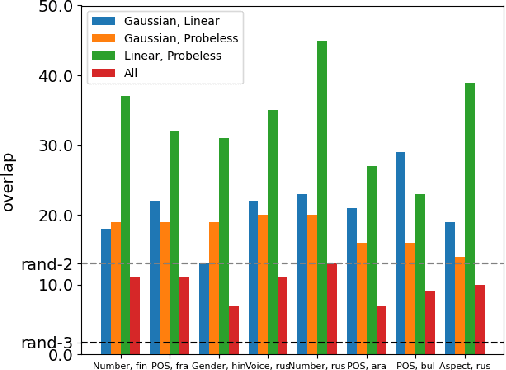
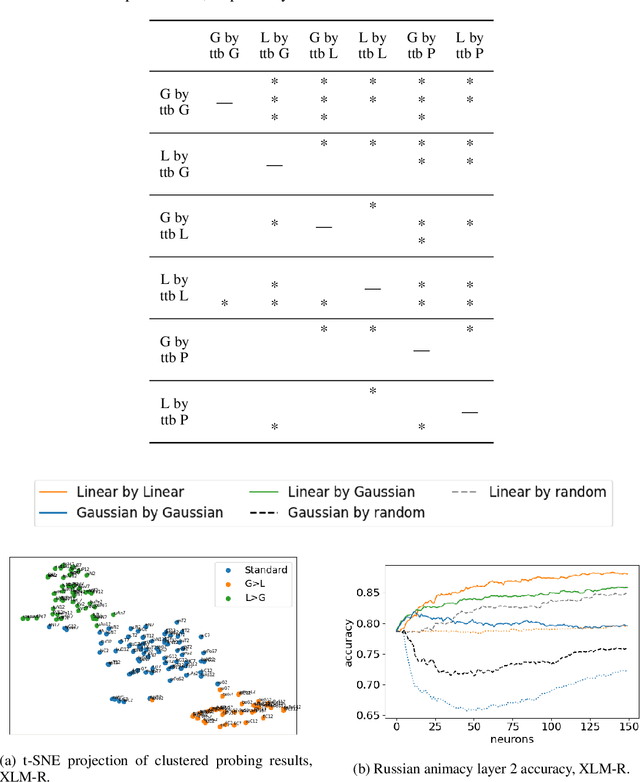
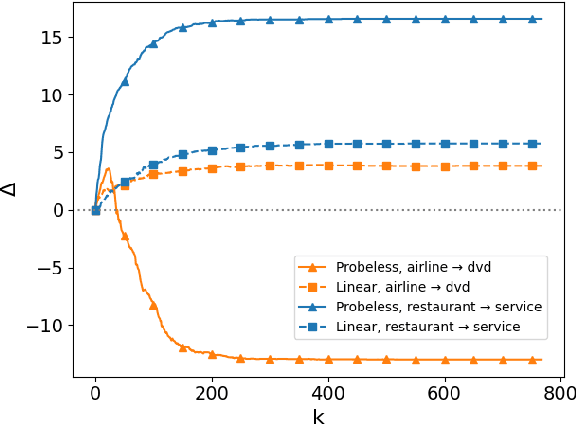
Abstract:While many studies have shown that linguistic information is encoded in hidden word representations, few have studied individual neurons, to show how and in which neurons it is encoded. Among these, the common approach is to use an external probe to rank neurons according to their relevance to some linguistic attribute, and to evaluate the obtained ranking using the same probe that produced it. We show two pitfalls in this methodology: 1. It confounds distinct factors: probe quality and ranking quality. We separate them and draw conclusions on each. 2. It focuses on encoded information, rather than information that is used by the model. We show that these are not the same. We compare two recent ranking methods and a simple one we introduce, and evaluate them with regard to both of these aspects.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge