Ole Winther
Implicit Transfer Operator Learning: Multiple Time-Resolution Surrogates for Molecular Dynamics
May 29, 2023Abstract:Computing properties of molecular systems rely on estimating expectations of the (unnormalized) Boltzmann distribution. Molecular dynamics (MD) is a broadly adopted technique to approximate such quantities. However, stable simulations rely on very small integration time-steps ($10^{-15}\,\mathrm{s}$), whereas convergence of some moments, e.g. binding free energy or rates, might rely on sampling processes on time-scales as long as $10^{-1}\, \mathrm{s}$, and these simulations must be repeated for every molecular system independently. Here, we present Implict Transfer Operator (ITO) Learning, a framework to learn surrogates of the simulation process with multiple time-resolutions. We implement ITO with denoising diffusion probabilistic models with a new SE(3) equivariant architecture and show the resulting models can generate self-consistent stochastic dynamics across multiple time-scales, even when the system is only partially observed. Finally, we present a coarse-grained CG-SE3-ITO model which can quantitatively model all-atom molecular dynamics using only coarse molecular representations. As such, ITO provides an important step towards multiple time- and space-resolution acceleration of MD.
Graph Neural Network Interatomic Potential Ensembles with Calibrated Aleatoric and Epistemic Uncertainty on Energy and Forces
May 10, 2023Abstract:Inexpensive machine learning potentials are increasingly being used to speed up structural optimization and molecular dynamics simulations of materials by iteratively predicting and applying interatomic forces. In these settings, it is crucial to detect when predictions are unreliable to avoid wrong or misleading results. Here, we present a complete framework for training and recalibrating graph neural network ensemble models to produce accurate predictions of energy and forces with calibrated uncertainty estimates. The proposed method considers both epistemic and aleatoric uncertainty and the total uncertainties are recalibrated post hoc using a nonlinear scaling function to achieve good calibration on previously unseen data, without loss of predictive accuracy. The method is demonstrated and evaluated on two challenging, publicly available datasets, ANI-1x (Smith et al.) and Transition1x (Schreiner et al.), both containing diverse conformations far from equilibrium. A detailed analysis of the predictive performance and uncertainty calibration is provided. In all experiments, the proposed method achieved low prediction error and good uncertainty calibration, with predicted uncertainty correlating with expected error, on energy and forces. To the best of our knowledge, the method presented in this paper is the first to consider a complete framework for obtaining calibrated epistemic and aleatoric uncertainty predictions on both energy and forces in ML potentials.
Dermatological Diagnosis Explainability Benchmark for Convolutional Neural Networks
Feb 23, 2023Abstract:In recent years, large strides have been taken in developing machine learning methods for dermatological applications, supported in part by the success of deep learning (DL). To date, diagnosing diseases from images is one of the most explored applications of DL within dermatology. Convolutional neural networks (ConvNets) are the most common (DL) method in medical imaging due to their training efficiency and accuracy, although they are often described as black boxes because of their limited explainability. One popular way to obtain insight into a ConvNet's decision mechanism is gradient class activation maps (Grad-CAM). A quantitative evaluation of the Grad-CAM explainability has been recently made possible by the release of DermXDB, a skin disease diagnosis explainability dataset which enables explainability benchmarking of ConvNet architectures. In this paper, we perform a literature review to identify the most common ConvNet architectures used for this task, and compare their Grad-CAM explanations with the explanation maps provided by DermXDB. We identified 11 architectures: DenseNet121, EfficientNet-B0, InceptionV3, InceptionResNetV2, MobileNet, MobileNetV2, NASNetMobile, ResNet50, ResNet50V2, VGG16, and Xception. We pre-trained all architectures on an clinical skin disease dataset, and fine-tuned them on a DermXDB subset. Validation results on the DermXDB holdout subset show an explainability F1 score of between 0.35-0.46, with Xception displaying the highest explainability performance. NASNetMobile reports the highest characteristic-level explainability sensitivity, despite it's mediocre diagnosis performance. These results highlight the importance of choosing the right architecture for the desired application and target market, underline need for additional explainability datasets, and further confirm the need for explainability benchmarking that relies on quantitative analyses.
Unifying Molecular and Textual Representations via Multi-task Language Modelling
Jan 29, 2023Abstract:The recent advances in neural language models have also been successfully applied to the field of chemistry, offering generative solutions for classical problems in molecular design and synthesis planning. These new methods have the potential to optimize laboratory operations and fuel a new era of data-driven automation in scientific discovery. However, specialized models are still typically required for each task, leading to the need for problem-specific fine-tuning and neglecting task interrelations. The main obstacle in this field is the lack of a unified representation between natural language and chemical representations, complicating and limiting human-machine interaction. Here, we propose a multi-domain, multi-task language model to solve a wide range of tasks in both the chemical and natural language domains. By leveraging multi-task learning, our model can handle chemical and natural language concurrently, without requiring expensive pre-training on single domains or task-specific models. Interestingly, sharing weights across domains remarkably improves our model when benchmarked against state-of-the-art baselines on single-domain and cross-domain tasks. In particular, sharing information across domains and tasks gives rise to large improvements in cross-domain tasks, the magnitude of which increase with scale, as measured by more than a dozen of relevant metrics. Our work suggests that such models can robustly and efficiently accelerate discovery in physical sciences by superseding problem-specific fine-tuning and enhancing human-model interactions.
ThoughtSource: A central hub for large language model reasoning data
Jan 27, 2023Abstract:Large language models (LLMs) such as GPT-3 and ChatGPT have recently demonstrated impressive results across a wide range of tasks. LLMs are still limited, however, in that they frequently fail at complex reasoning, their reasoning processes are opaque, they are prone to 'hallucinate' facts, and there are concerns about their underlying biases. Letting models verbalize reasoning steps as natural language, a technique known as chain-of-thought prompting, has recently been proposed as a way to address some of these issues. Here we present the first release of ThoughtSource, a meta-dataset and software library for chain-of-thought (CoT) reasoning. The goal of ThoughtSource is to improve future artificial intelligence systems by facilitating qualitative understanding of CoTs, enabling empirical evaluations, and providing training data. This first release of ThoughtSource integrates six scientific/medical, three general-domain and five math word question answering datasets.
Variational Open-Domain Question Answering
Sep 23, 2022
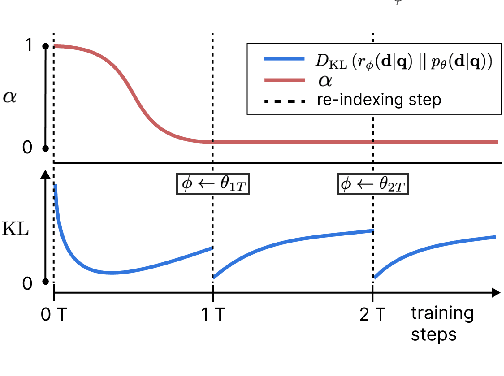
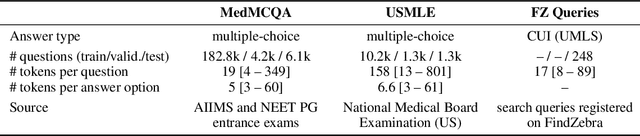
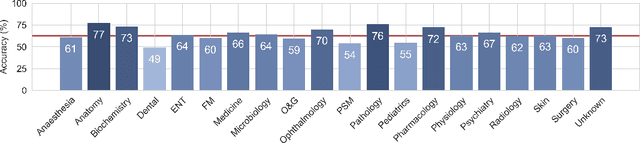
Abstract:We introduce the Variational Open-Domain (VOD) framework for end-to-end training and evaluation of retrieval-augmented models (open-domain question answering and language modelling). We show that the R\'enyi variational bound, a lower bound to the task marginal likelihood, can be exploited to aid optimization and use importance sampling to estimate the task log-likelihood lower bound and its gradients using samples drawn from an auxiliary retriever (approximate posterior). The framework can be used to train modern retrieval-augmented systems end-to-end using tractable and consistent estimates of the R\'enyi variational bound and its gradients. We demonstrate the framework's versatility by training reader-retriever BERT-based models on multiple-choice medical exam questions (MedMCQA and USMLE). We registered a new state-of-the-art for both datasets (MedMCQA: $62.9$\%, USMLE: $55.0$\%). Last, we show that the retriever part of the learned reader-retriever model trained on the medical board exam questions can be used in search engines for a medical knowledge base.
Explainable Image Quality Assessments in Teledermatological Photography
Sep 10, 2022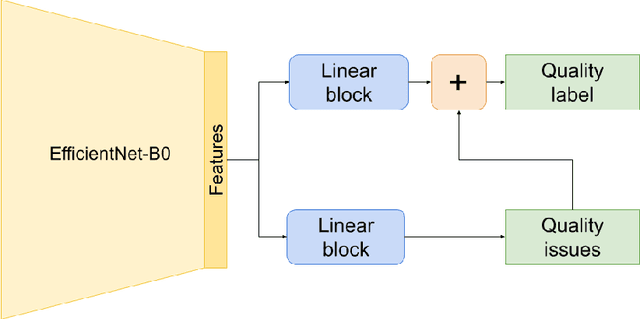
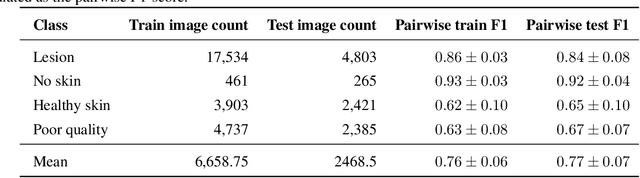
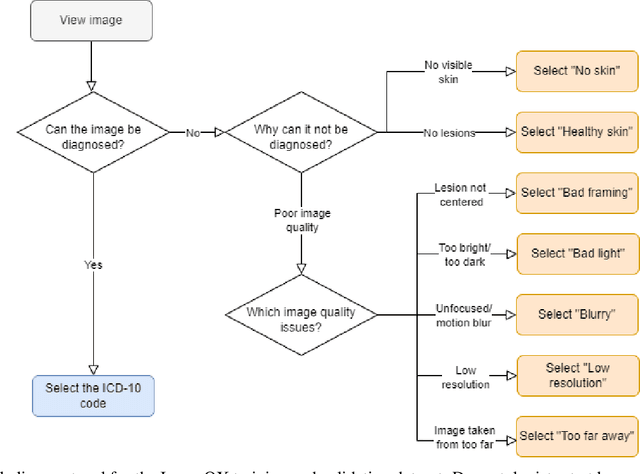
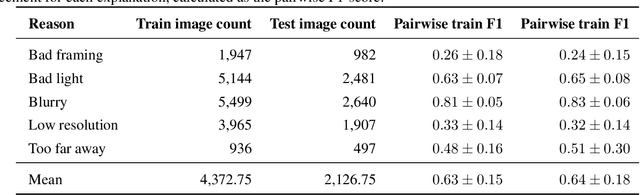
Abstract:Image quality is a crucial factor in the success of teledermatological consultations. However, up to 50% of images sent by patients have quality issues, thus increasing the time to diagnosis and treatment. An automated, easily deployable, explainable method for assessing image quality is necessary to improve the current teledermatological consultation flow. We introduce ImageQX, a convolutional neural network trained for image quality assessment with a learning mechanism for identifying the most common poor image quality explanations: bad framing, bad lighting, blur, low resolution, and distance issues. ImageQX was trained on 26635 photographs and validated on 9874 photographs, each annotated with image quality labels and poor image quality explanations by up to 12 board-certified dermatologists. The photographic images were taken between 2017-2019 using a mobile skin disease tracking application accessible worldwide. Our method achieves expert-level performance for both image quality assessment and poor image quality explanation. For image quality assessment, ImageQX obtains a macro F1-score of 0.73 which places it within standard deviation of the pairwise inter-rater F1-score of 0.77. For poor image quality explanations, our method obtains F1-scores of between 0.37 and 0.70, similar to the inter-rater pairwise F1-score of between 0.24 and 0.83. Moreover, with a size of only 15 MB, ImageQX is easily deployable on mobile devices. With an image quality detection performance similar to that of dermatologists, incorporating ImageQX into the teledermatology flow can reduce the image evaluation burden on dermatologists, while at the same time reducing the time to diagnosis and treatment for patients. We introduce ImageQX, a first of its kind explainable image quality assessor which leverages domain expertise to improve the quality and efficiency of dermatological care in a virtual setting.
Transition1x -- a Dataset for Building Generalizable Reactive Machine Learning Potentials
Jul 25, 2022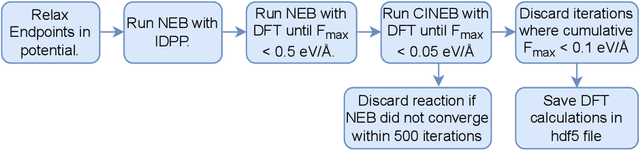
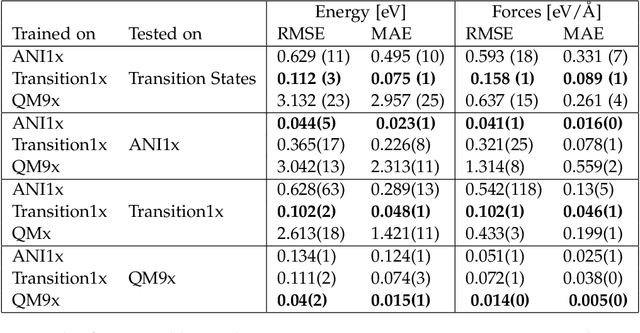
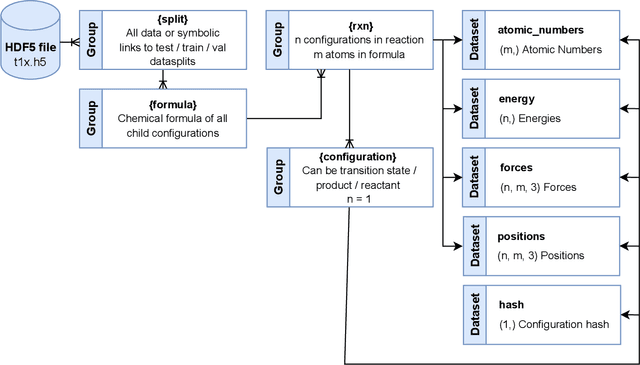
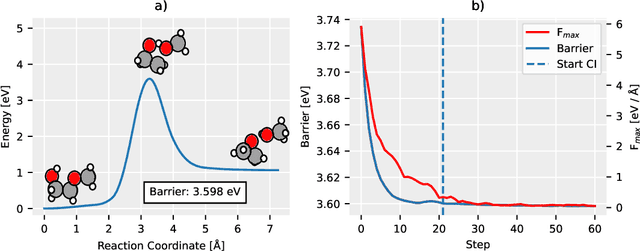
Abstract:Machine Learning (ML) models have, in contrast to their usefulness in molecular dynamics studies, had limited success as surrogate potentials for reaction barrier search. It is due to the scarcity of training data in relevant transition state regions of chemical space. Currently, available datasets for training ML models on small molecular systems almost exclusively contain configurations at or near equilibrium. In this work, we present the dataset Transition1x containing 9.6 million Density Functional Theory (DFT) calculations of forces and energies of molecular configurations on and around reaction pathways at the wB97x/6-31G(d) level of theory. The data was generated by running Nudged Elastic Band (NEB) calculations with DFT on 10k reactions while saving intermediate calculations. We train state-of-the-art equivariant graph message-passing neural network models on Transition1x and cross-validate on the popular ANI1x and QM9 datasets. We show that ML models cannot learn features in transition-state regions solely by training on hitherto popular benchmark datasets. Transition1x is a new challenging benchmark that will provide an important step towards developing next-generation ML force fields that also work far away from equilibrium configurations and reactive systems.
NeuralNEB -- Neural Networks can find Reaction Paths Fast
Jul 21, 2022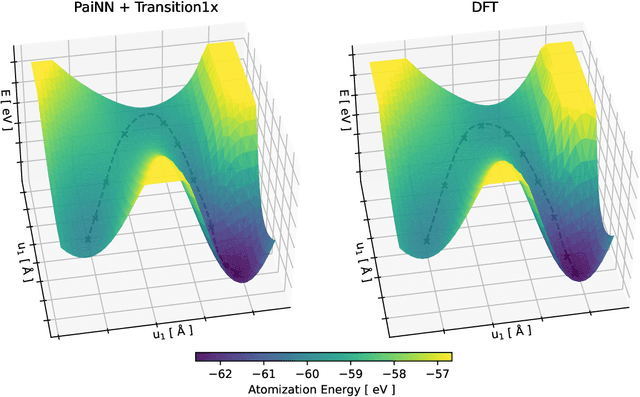
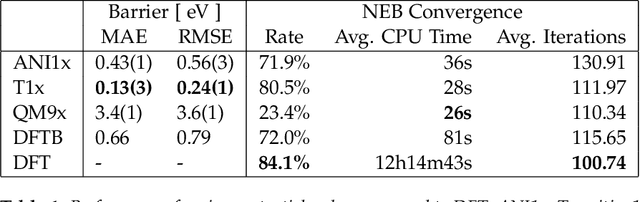
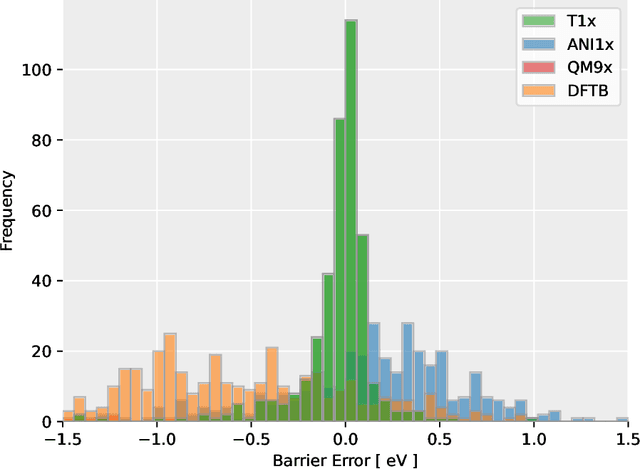
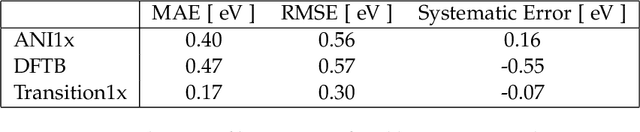
Abstract:Quantum mechanical methods like Density Functional Theory (DFT) are used with great success alongside efficient search algorithms for studying kinetics of reactive systems. However, DFT is prohibitively expensive for large scale exploration. Machine Learning (ML) models have turned out to be excellent emulators of small molecule DFT calculations and could possibly replace DFT in such tasks. For kinetics, success relies primarily on the models capability to accurately predict the Potential Energy Surface (PES) around transition-states and Minimal Energy Paths (MEPs). Previously this has not been possible due to scarcity of relevant data in the literature. In this paper we train state of the art equivariant Graph Neural Network (GNN)-based models on around 10.000 elementary reactions from the Transition1x dataset. We apply the models as potentials for the Nudged Elastic Band (NEB) algorithm and achieve a Mean Average Error (MAE) of 0.13+/-0.03 eV on barrier energies on unseen reactions. We compare the results against equivalent models trained on QM9 and ANI1x. We also compare with and outperform Density Functional based Tight Binding (DFTB) on both accuracy and computational resource. The implication is that ML models, given relevant data, are now at a level where they can be applied for downstream tasks in quantum chemistry transcending prediction of simple molecular features.
Can large language models reason about medical questions?
Jul 17, 2022
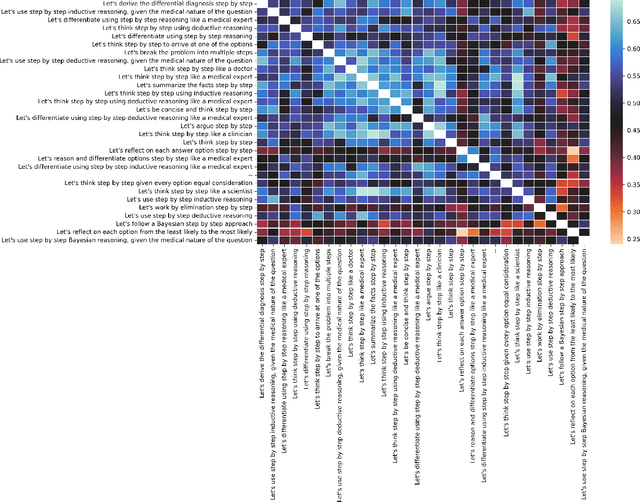

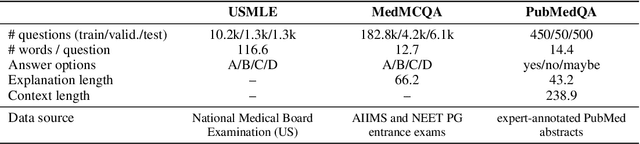
Abstract:Although large language models (LLMs) often produce impressive outputs, they also fail to reason and be factual. We set out to investigate how these limitations affect the LLM's ability to answer and reason about difficult real-world based questions. We applied the human-aligned GPT-3 (InstructGPT) to answer multiple-choice medical exam questions (USMLE and MedMCQA) and medical research questions (PubMedQA). We investigated Chain-of-thought (think step by step) prompts, grounding (augmenting the prompt with search results) and few-shot (prepending the question with question-answer exemplars). For a subset of the USMLE questions, a medical domain expert reviewed and annotated the model's reasoning. Overall, GPT-3 achieved a substantial improvement in state-of-the-art machine learning performance. We observed that GPT-3 is often knowledgeable and can reason about medical questions. GPT-3, when confronted with a question it cannot answer, will still attempt to answer, often resulting in a biased predictive distribution. LLMs are not on par with human performance but our results suggest the emergence of reasoning patterns that are compatible with medical problem-solving. We speculate that scaling model and data, enhancing prompt alignment and allowing for better contextualization of the completions will be sufficient for LLMs to reach human-level performance on this type of task.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge