Nathan Nussbaum
Model-assisted cohort selection with bias analysis for generating large-scale cohorts from the EHR for oncology research
Jan 13, 2020
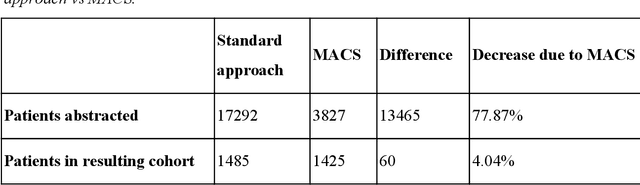
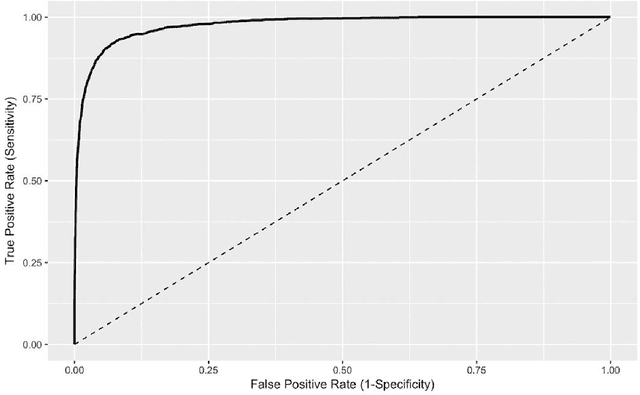
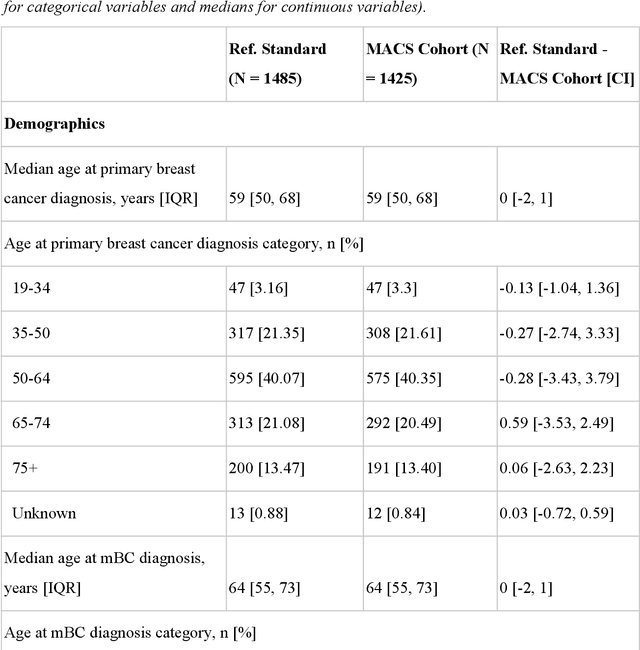
Abstract:Objective Electronic health records (EHRs) are a promising source of data for health outcomes research in oncology. A challenge in using EHR data is that selecting cohorts of patients often requires information in unstructured parts of the record. Machine learning has been used to address this, but even high-performing algorithms may select patients in a non-random manner and bias the resulting cohort. To improve the efficiency of cohort selection while measuring potential bias, we introduce a technique called Model-Assisted Cohort Selection (MACS) with Bias Analysis and apply it to the selection of metastatic breast cancer (mBC) patients. Materials and Methods We trained a model on 17,263 patients using term-frequency inverse-document-frequency (TF-IDF) and logistic regression. We used a test set of 17,292 patients to measure algorithm performance and perform Bias Analysis. We compared the cohort generated by MACS to the cohort that would have been generated without MACS as reference standard, first by comparing distributions of an extensive set of clinical and demographic variables and then by comparing the results of two analyses addressing existing example research questions. Results Our algorithm had an area under the curve (AUC) of 0.976, a sensitivity of 96.0%, and an abstraction efficiency gain of 77.9%. During Bias Analysis, we found no large differences in baseline characteristics and no differences in the example analyses. Conclusion MACS with bias analysis can significantly improve the efficiency of cohort selection on EHR data while instilling confidence that outcomes research performed on the resulting cohort will not be biased.
TIFTI: A Framework for Extracting Drug Intervals from Longitudinal Clinic Notes
Dec 03, 2018
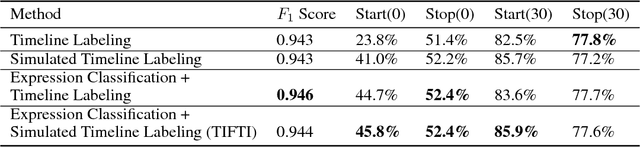

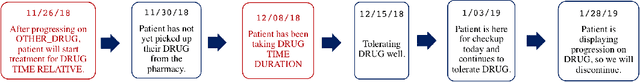
Abstract:Oral drugs are becoming increasingly common in oncology care. In contrast to intravenous chemotherapy, which is administered in the clinic and carefully tracked via structure electronic health records (EHRs), oral drug treatment is self-administered and therefore not tracked as well. Often, the details of oral cancer treatment occur only in unstructured clinic notes. Extracting this information is critical to understanding a patient's treatment history. Yet, this a challenging task because treatment intervals must be inferred longitudinally from both explicit mentions in the text as well as from document timestamps. In this work, we present TIFTI (Temporally Integrated Framework for Treatment Intervals), a robust framework for extracting oral drug treatment intervals from a patient's unstructured notes. TIFTI leverages distinct sources of temporal information by breaking the problem down into two separate subtasks: document-level sequence labeling and date extraction. On a labeled dataset of metastatic renal-cell carcinoma (RCC) patients, it exactly matched the labeled start date in 46% of the examples (86% of the examples within 30 days), and it exactly matched the labeled end date in 52% of the examples (78% of the examples within 30 days). Without retraining, the model achieved a similar level of performance on a labeled dataset of advanced non-small-cell lung cancer (NSCLC) patients.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge