Mihaela van der Schaar
Synthetic Model Combination: An Instance-wise Approach to Unsupervised Ensemble Learning
Oct 11, 2022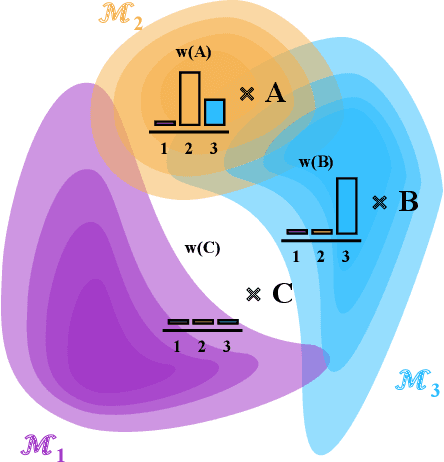
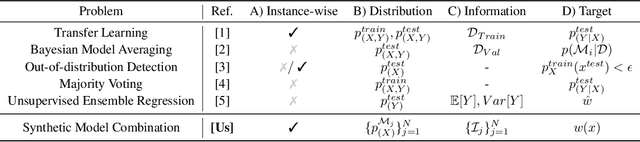

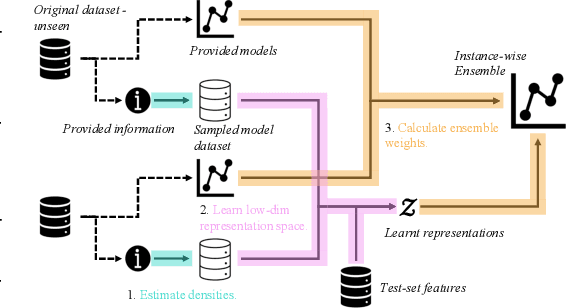
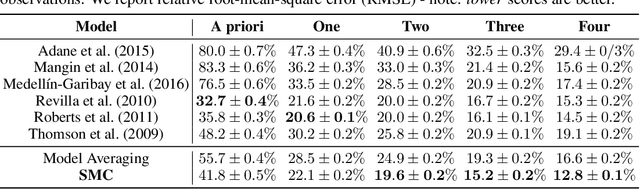
Abstract:Consider making a prediction over new test data without any opportunity to learn from a training set of labelled data - instead given access to a set of expert models and their predictions alongside some limited information about the dataset used to train them. In scenarios from finance to the medical sciences, and even consumer practice, stakeholders have developed models on private data they either cannot, or do not want to, share. Given the value and legislation surrounding personal information, it is not surprising that only the models, and not the data, will be released - the pertinent question becoming: how best to use these models? Previous work has focused on global model selection or ensembling, with the result of a single final model across the feature space. Machine learning models perform notoriously poorly on data outside their training domain however, and so we argue that when ensembling models the weightings for individual instances must reflect their respective domains - in other words models that are more likely to have seen information on that instance should have more attention paid to them. We introduce a method for such an instance-wise ensembling of models, including a novel representation learning step for handling sparse high-dimensional domains. Finally, we demonstrate the need and generalisability of our method on classical machine learning tasks as well as highlighting a real world use case in the pharmacological setting of vancomycin precision dosing.
Transfer Learning on Heterogeneous Feature Spaces for Treatment Effects Estimation
Oct 08, 2022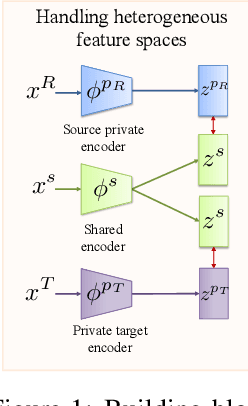
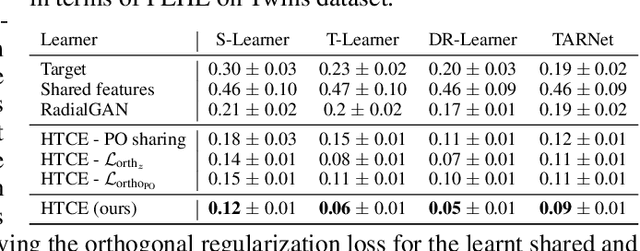
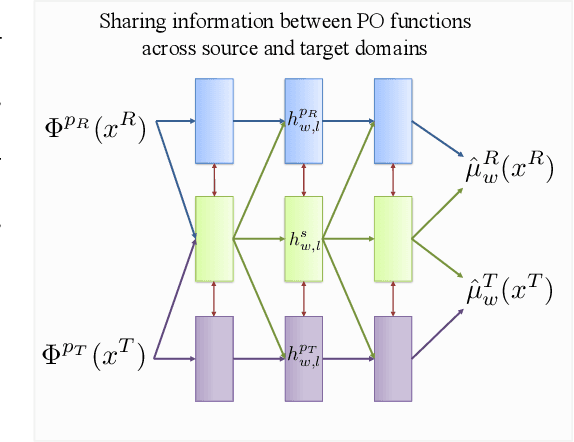
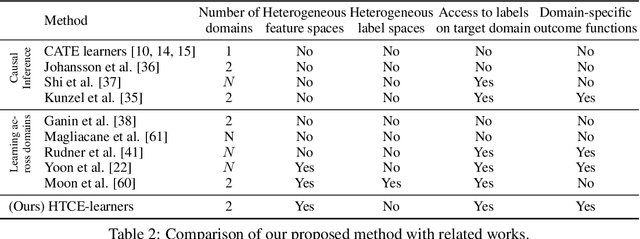
Abstract:Consider the problem of improving the estimation of conditional average treatment effects (CATE) for a target domain of interest by leveraging related information from a source domain with a different feature space. This heterogeneous transfer learning problem for CATE estimation is ubiquitous in areas such as healthcare where we may wish to evaluate the effectiveness of a treatment for a new patient population for which different clinical covariates and limited data are available. In this paper, we address this problem by introducing several building blocks that use representation learning to handle the heterogeneous feature spaces and a flexible multi-task architecture with shared and private layers to transfer information between potential outcome functions across domains. Then, we show how these building blocks can be used to recover transfer learning equivalents of the standard CATE learners. On a new semi-synthetic data simulation benchmark for heterogeneous transfer learning we not only demonstrate performance improvements of our heterogeneous transfer causal effect learners across datasets, but also provide insights into the differences between these learners from a transfer perspective.
Concept Activation Regions: A Generalized Framework For Concept-Based Explanations
Sep 29, 2022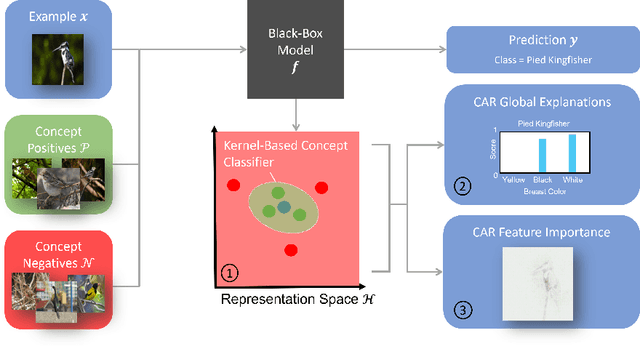
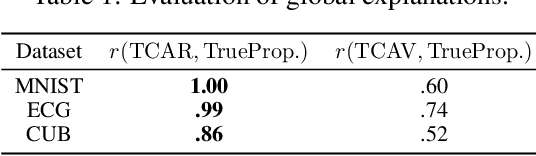
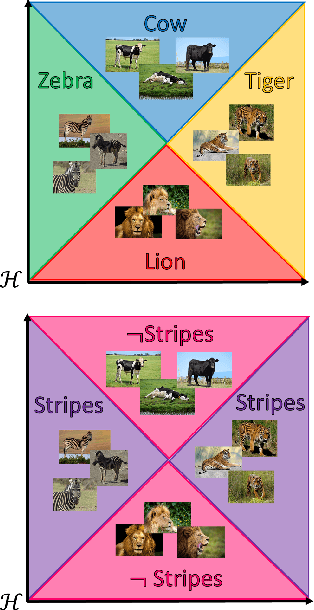
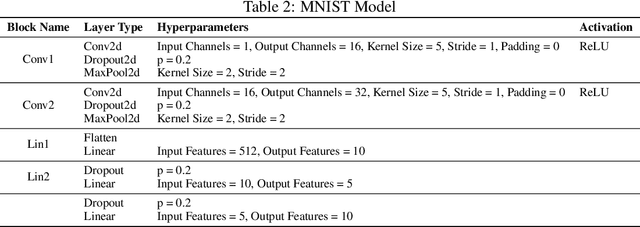
Abstract:Concept-based explanations permit to understand the predictions of a deep neural network (DNN) through the lens of concepts specified by users. Existing methods assume that the examples illustrating a concept are mapped in a fixed direction of the DNN's latent space. When this holds true, the concept can be represented by a concept activation vector (CAV) pointing in that direction. In this work, we propose to relax this assumption by allowing concept examples to be scattered across different clusters in the DNN's latent space. Each concept is then represented by a region of the DNN's latent space that includes these clusters and that we call concept activation region (CAR). To formalize this idea, we introduce an extension of the CAV formalism that is based on the kernel trick and support vector classifiers. This CAR formalism yields global concept-based explanations and local concept-based feature importance. We prove that CAR explanations built with radial kernels are invariant under latent space isometries. In this way, CAR assigns the same explanations to latent spaces that have the same geometry. We further demonstrate empirically that CARs offer (1) more accurate descriptions of how concepts are scattered in the DNN's latent space; (2) global explanations that are closer to human concept annotations and (3) concept-based feature importance that meaningfully relate concepts with each other. Finally, we use CARs to show that DNNs can autonomously rediscover known scientific concepts, such as the prostate cancer grading system.
Adaptively Identifying Patient Populations With Treatment Benefit in Clinical Trials
Aug 11, 2022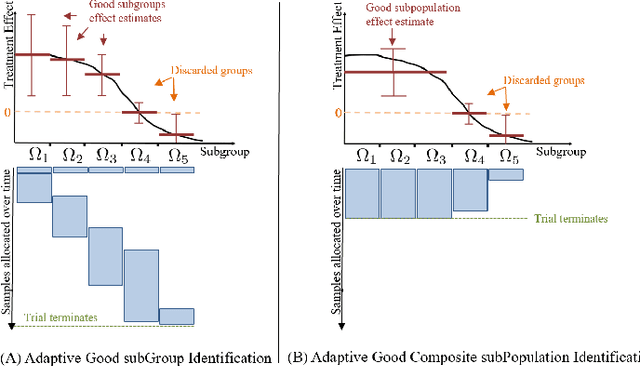
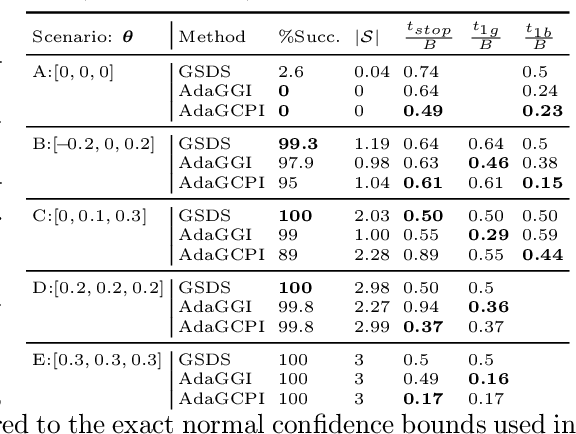
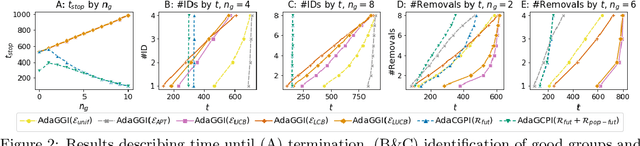
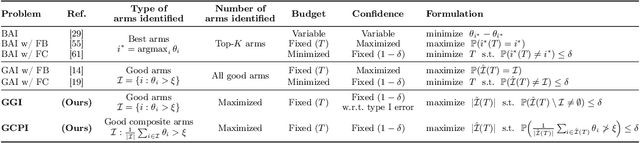
Abstract:We study the problem of adaptively identifying patient subpopulations that benefit from a given treatment during a confirmatory clinical trial. This type of adaptive clinical trial, often referred to as adaptive enrichment design, has been thoroughly studied in biostatistics with a focus on a limited number of subgroups (typically two) which make up (sub)populations, and a small number of interim analysis points. In this paper, we aim to relax classical restrictions on such designs and investigate how to incorporate ideas from the recent machine learning literature on adaptive and online experimentation to make trials more flexible and efficient. We find that the unique characteristics of the subpopulation selection problem -- most importantly that (i) one is usually interested in finding subpopulations with any treatment benefit (and not necessarily the single subgroup with largest effect) given a limited budget and that (ii) effectiveness only has to be demonstrated across the subpopulation on average -- give rise to interesting challenges and new desiderata when designing algorithmic solutions. Building on these findings, we propose AdaGGI and AdaGCPI, two meta-algorithms for subpopulation construction, which focus on identifying good subgroups and good composite subpopulations, respectively. We empirically investigate their performance across a range of simulation scenarios and derive insights into their (dis)advantages across different settings.
DAPDAG: Domain Adaptation via Perturbed DAG Reconstruction
Aug 02, 2022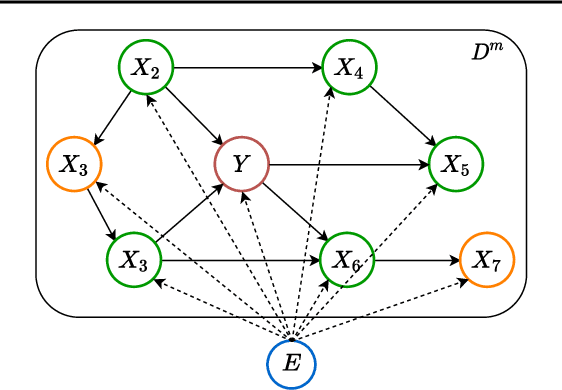
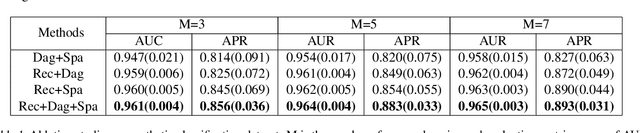
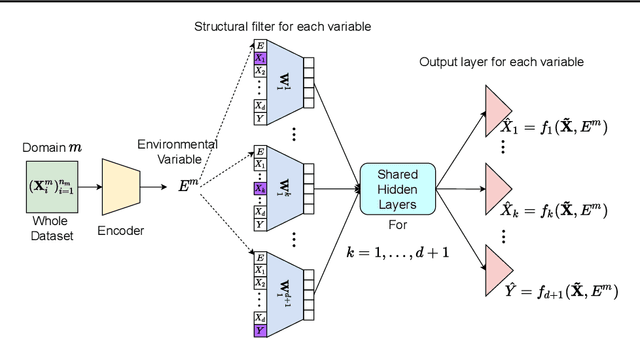
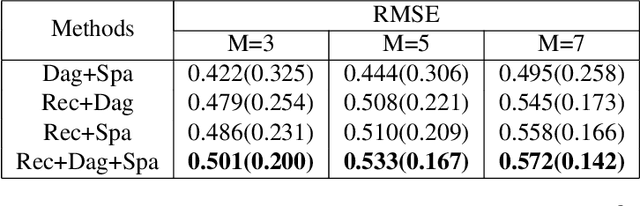
Abstract:Leveraging labelled data from multiple domains to enable prediction in another domain without labels is a significant, yet challenging problem. To address this problem, we introduce the framework DAPDAG (\textbf{D}omain \textbf{A}daptation via \textbf{P}erturbed \textbf{DAG} Reconstruction) and propose to learn an auto-encoder that undertakes inference on population statistics given features and reconstructing a directed acyclic graph (DAG) as an auxiliary task. The underlying DAG structure is assumed invariant among observed variables whose conditional distributions are allowed to vary across domains led by a latent environmental variable $E$. The encoder is designed to serve as an inference device on $E$ while the decoder reconstructs each observed variable conditioned on its graphical parents in the DAG and the inferred $E$. We train the encoder and decoder jointly in an end-to-end manner and conduct experiments on synthetic and real datasets with mixed variables. Empirical results demonstrate that reconstructing the DAG benefits the approximate inference. Furthermore, our approach can achieve competitive performance against other benchmarks in prediction tasks, with better adaptation ability, especially in the target domain significantly different from the source domains.
DAUX: a Density-based Approach for Uncertainty eXplanations
Jul 11, 2022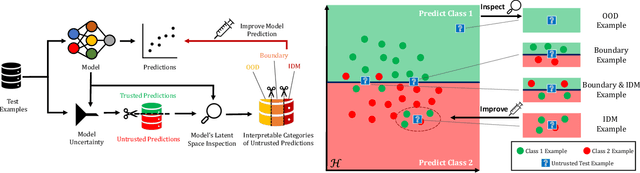
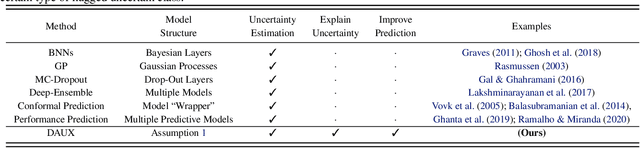
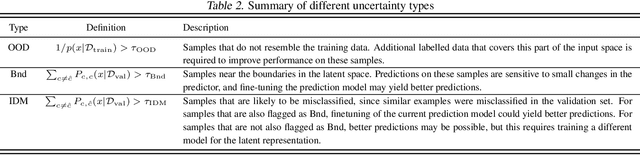
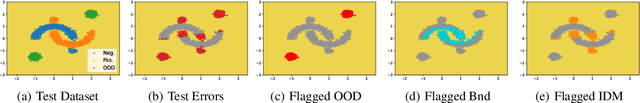
Abstract:Uncertainty quantification (UQ) is essential for creating trustworthy machine learning models. Recent years have seen a steep rise in UQ methods that can flag suspicious examples, however, it is often unclear what exactly these methods identify. In this work, we propose an assumption-light method for interpreting UQ models themselves. We introduce the confusion density matrix -- a kernel-based approximation of the misclassification density -- and use this to categorize suspicious examples identified by a given UQ method into three classes: out-of-distribution (OOD) examples, boundary (Bnd) examples, and examples in regions of high in-distribution misclassification (IDM). Through extensive experiments, we shed light on existing UQ methods and show that the cause of the uncertainty differs across models. Additionally, we show how the proposed framework can make use of the categorized examples to improve predictive performance.
D-CIPHER: Discovery of Closed-form PDEs
Jun 21, 2022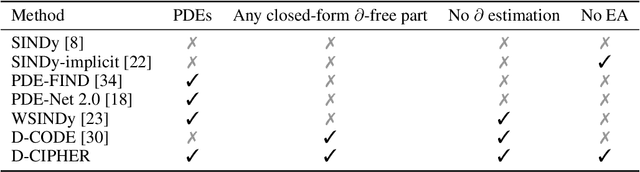
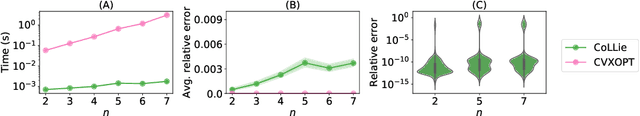
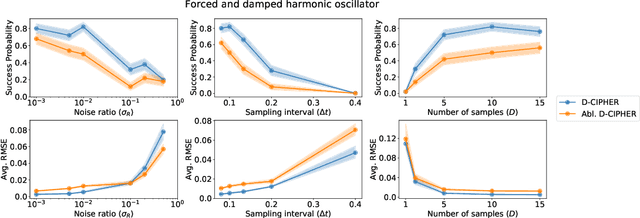
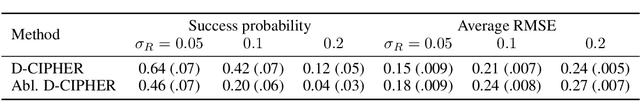
Abstract:Closed-form differential equations, including partial differential equations and higher-order ordinary differential equations, are one of the most important tools used by scientists to model and better understand natural phenomena. Discovering these equations directly from data is challenging because it requires modeling relationships between various derivatives that are not observed in the data (\textit{equation-data mismatch}) and it involves searching across a huge space of possible equations. Current approaches make strong assumptions about the form of the equation and thus fail to discover many well-known systems. Moreover, many of them resolve the equation-data mismatch by estimating the derivatives, which makes them inadequate for noisy and infrequently sampled systems. To this end, we propose D-CIPHER, which is robust to measurement artifacts and can uncover a new and very general class of differential equations. We further design a novel optimization procedure, CoLLie, to help D-CIPHER search through this class efficiently. Finally, we demonstrate empirically that it can discover many well-known equations that are beyond the capabilities of current methods.
Benchmarking Heterogeneous Treatment Effect Models through the Lens of Interpretability
Jun 16, 2022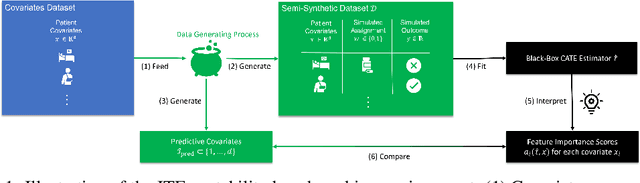
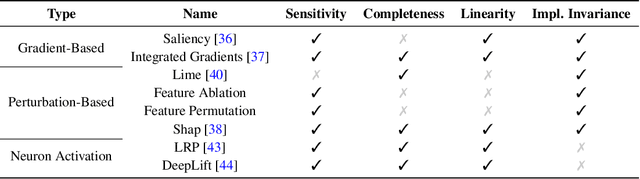

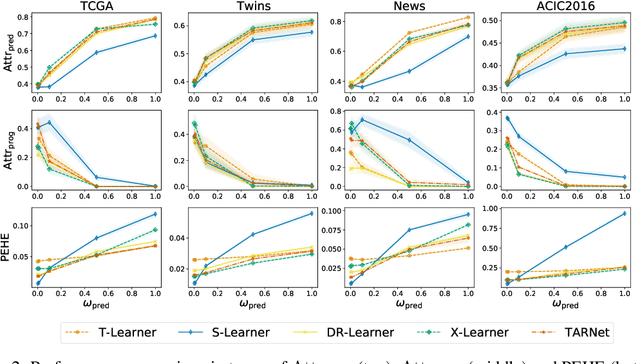
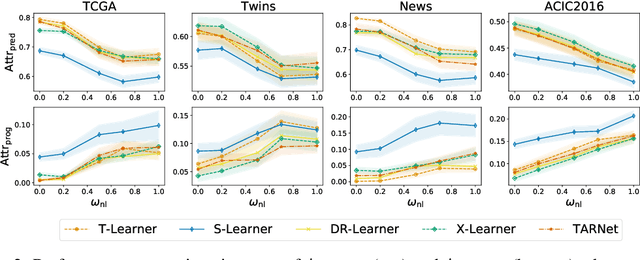
Abstract:Estimating personalized effects of treatments is a complex, yet pervasive problem. To tackle it, recent developments in the machine learning (ML) literature on heterogeneous treatment effect estimation gave rise to many sophisticated, but opaque, tools: due to their flexibility, modularity and ability to learn constrained representations, neural networks in particular have become central to this literature. Unfortunately, the assets of such black boxes come at a cost: models typically involve countless nontrivial operations, making it difficult to understand what they have learned. Yet, understanding these models can be crucial -- in a medical context, for example, discovered knowledge on treatment effect heterogeneity could inform treatment prescription in clinical practice. In this work, we therefore use post-hoc feature importance methods to identify features that influence the model's predictions. This allows us to evaluate treatment effect estimators along a new and important dimension that has been overlooked in previous work: We construct a benchmarking environment to empirically investigate the ability of personalized treatment effect models to identify predictive covariates -- covariates that determine differential responses to treatment. Our benchmarking environment then enables us to provide new insight into the strengths and weaknesses of different types of treatment effects models as we modulate different challenges specific to treatment effect estimation -- e.g. the ratio of prognostic to predictive information, the possible nonlinearity of potential outcomes and the presence and type of confounding.
Continuous-Time Modeling of Counterfactual Outcomes Using Neural Controlled Differential Equations
Jun 16, 2022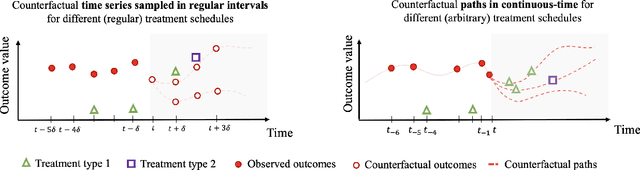
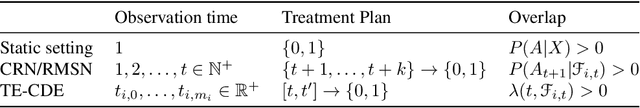

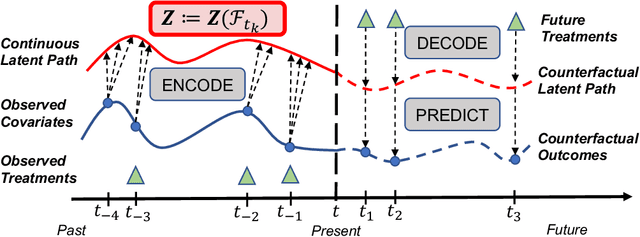
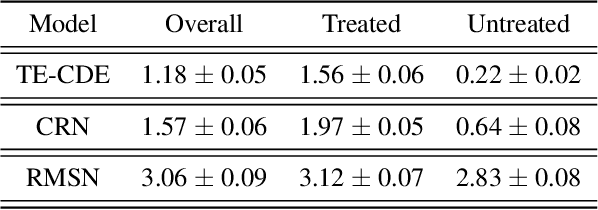
Abstract:Estimating counterfactual outcomes over time has the potential to unlock personalized healthcare by assisting decision-makers to answer ''what-iF'' questions. Existing causal inference approaches typically consider regular, discrete-time intervals between observations and treatment decisions and hence are unable to naturally model irregularly sampled data, which is the common setting in practice. To handle arbitrary observation patterns, we interpret the data as samples from an underlying continuous-time process and propose to model its latent trajectory explicitly using the mathematics of controlled differential equations. This leads to a new approach, the Treatment Effect Neural Controlled Differential Equation (TE-CDE), that allows the potential outcomes to be evaluated at any time point. In addition, adversarial training is used to adjust for time-dependent confounding which is critical in longitudinal settings and is an added challenge not encountered in conventional time-series. To assess solutions to this problem, we propose a controllable simulation environment based on a model of tumor growth for a range of scenarios with irregular sampling reflective of a variety of clinical scenarios. TE-CDE consistently outperforms existing approaches in all simulated scenarios with irregular sampling.
HyperImpute: Generalized Iterative Imputation with Automatic Model Selection
Jun 15, 2022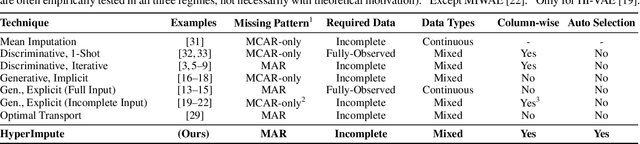
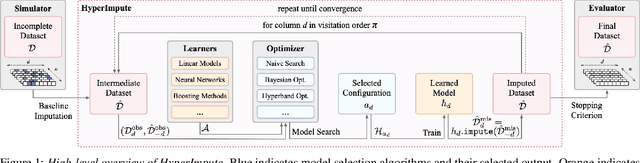
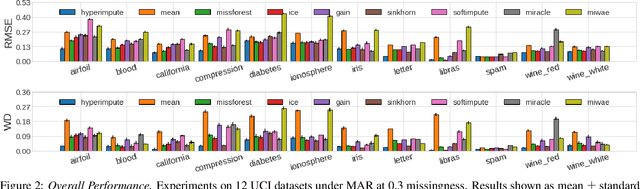
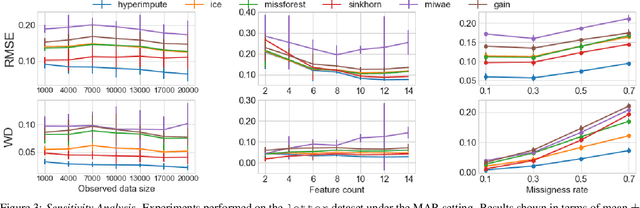
Abstract:Consider the problem of imputing missing values in a dataset. One the one hand, conventional approaches using iterative imputation benefit from the simplicity and customizability of learning conditional distributions directly, but suffer from the practical requirement for appropriate model specification of each and every variable. On the other hand, recent methods using deep generative modeling benefit from the capacity and efficiency of learning with neural network function approximators, but are often difficult to optimize and rely on stronger data assumptions. In this work, we study an approach that marries the advantages of both: We propose *HyperImpute*, a generalized iterative imputation framework for adaptively and automatically configuring column-wise models and their hyperparameters. Practically, we provide a concrete implementation with out-of-the-box learners, optimizers, simulators, and extensible interfaces. Empirically, we investigate this framework via comprehensive experiments and sensitivities on a variety of public datasets, and demonstrate its ability to generate accurate imputations relative to a strong suite of benchmarks. Contrary to recent work, we believe our findings constitute a strong defense of the iterative imputation paradigm.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge