Max Welling
UC Irvine
Bayesian Dark Knowledge
Nov 06, 2015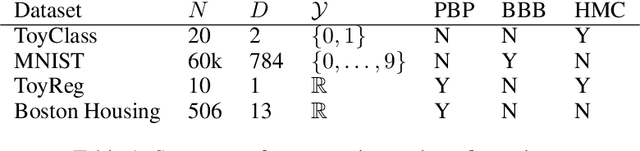
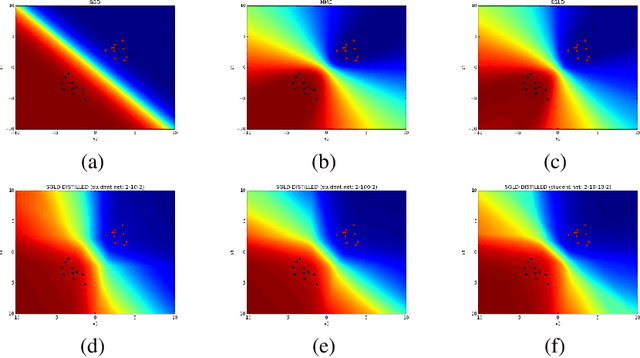
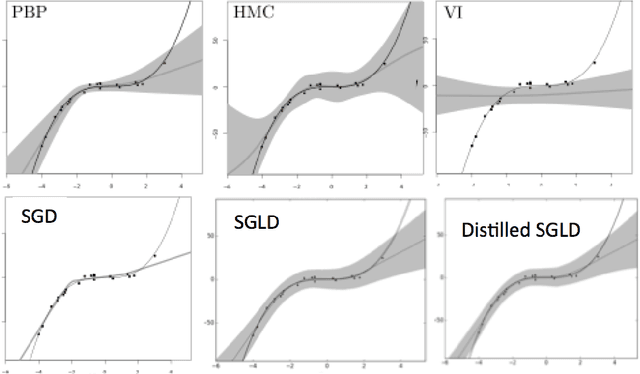
Abstract:We consider the problem of Bayesian parameter estimation for deep neural networks, which is important in problem settings where we may have little data, and/ or where we need accurate posterior predictive densities, e.g., for applications involving bandits or active learning. One simple approach to this is to use online Monte Carlo methods, such as SGLD (stochastic gradient Langevin dynamics). Unfortunately, such a method needs to store many copies of the parameters (which wastes memory), and needs to make predictions using many versions of the model (which wastes time). We describe a method for "distilling" a Monte Carlo approximation to the posterior predictive density into a more compact form, namely a single deep neural network. We compare to two very recent approaches to Bayesian neural networks, namely an approach based on expectation propagation [Hernandez-Lobato and Adams, 2015] and an approach based on variational Bayes [Blundell et al., 2015]. Our method performs better than both of these, is much simpler to implement, and uses less computation at test time.
Scalable MCMC for Mixed Membership Stochastic Blockmodels
Oct 22, 2015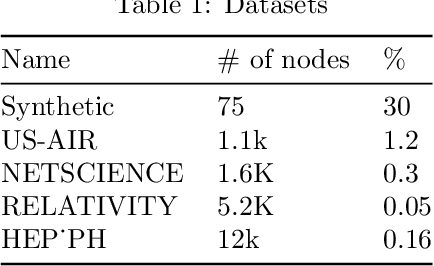
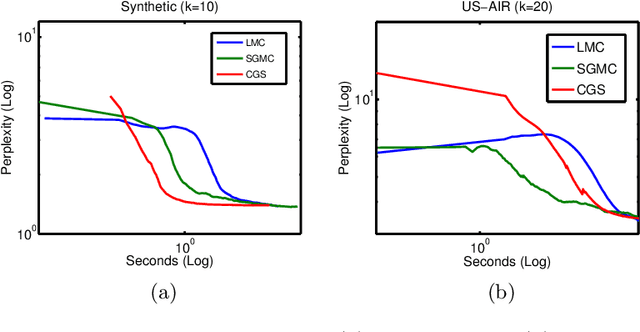
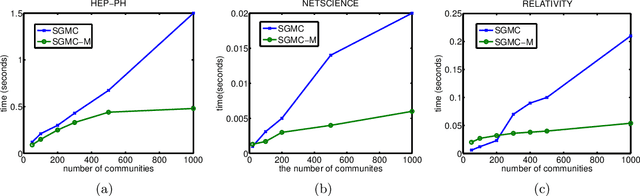
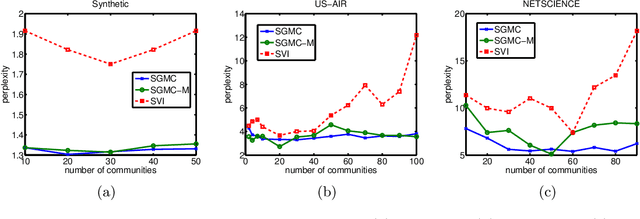
Abstract:We propose a stochastic gradient Markov chain Monte Carlo (SG-MCMC) algorithm for scalable inference in mixed-membership stochastic blockmodels (MMSB). Our algorithm is based on the stochastic gradient Riemannian Langevin sampler and achieves both faster speed and higher accuracy at every iteration than the current state-of-the-art algorithm based on stochastic variational inference. In addition we develop an approximation that can handle models that entertain a very large number of communities. The experimental results show that SG-MCMC strictly dominates competing algorithms in all cases.
MLitB: Machine Learning in the Browser
Jun 17, 2015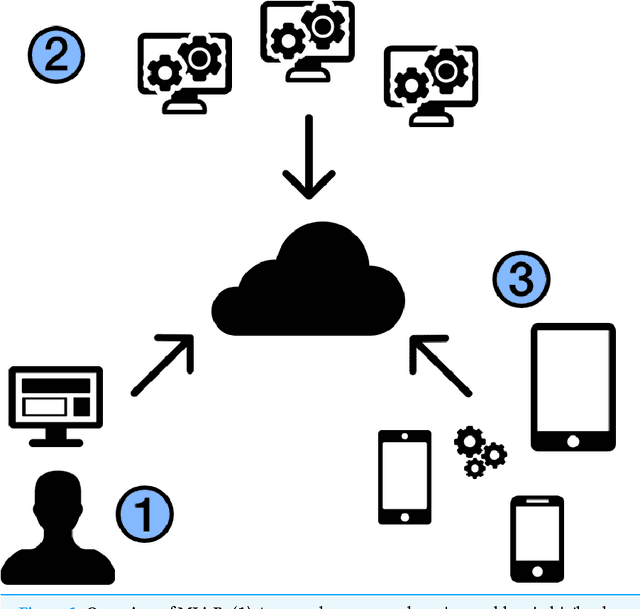
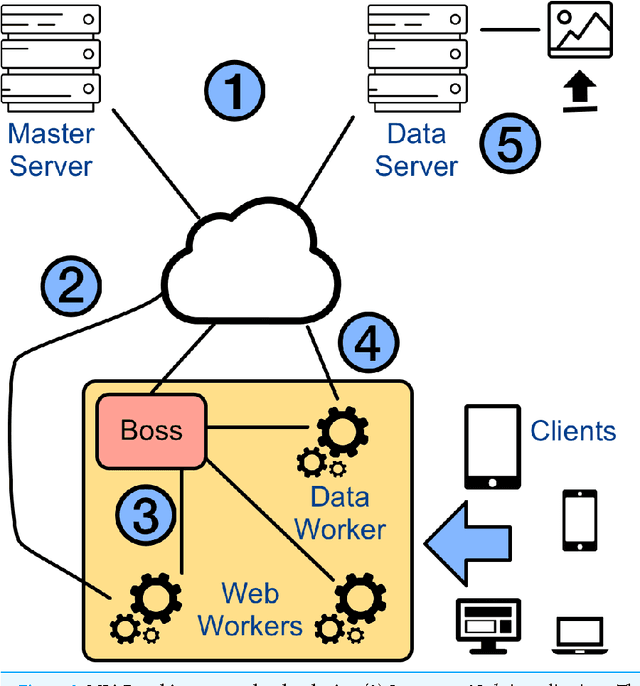
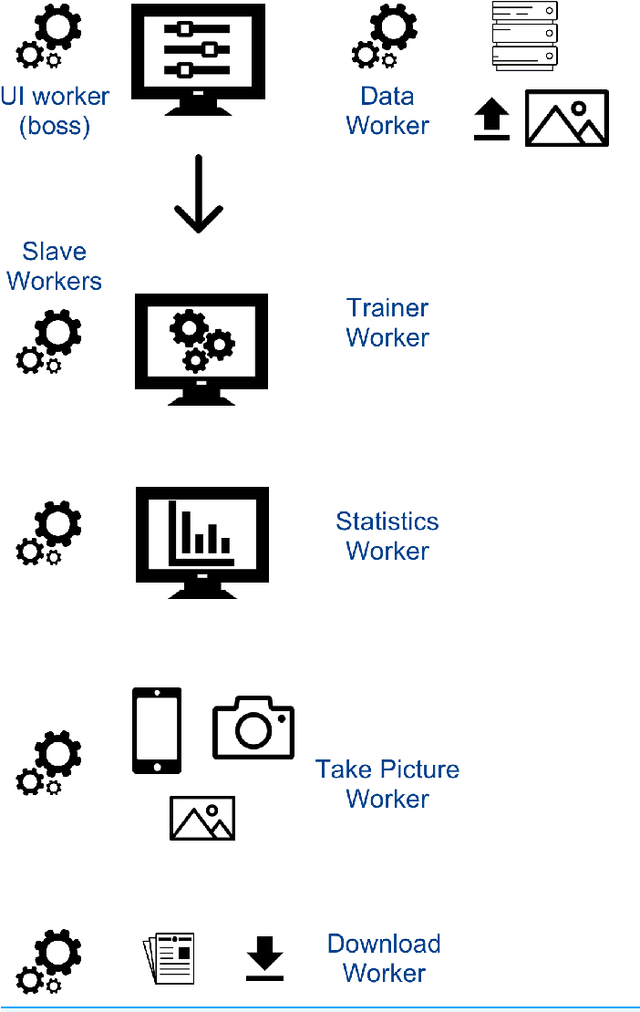
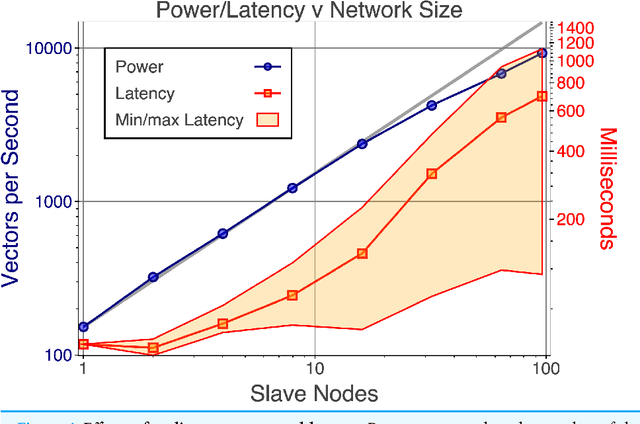
Abstract:With few exceptions, the field of Machine Learning (ML) research has largely ignored the browser as a computational engine. Beyond an educational resource for ML, the browser has vast potential to not only improve the state-of-the-art in ML research, but also, inexpensively and on a massive scale, to bring sophisticated ML learning and prediction to the public at large. This paper introduces MLitB, a prototype ML framework written entirely in JavaScript, capable of performing large-scale distributed computing with heterogeneous classes of devices. The development of MLitB has been driven by several underlying objectives whose aim is to make ML learning and usage ubiquitous (by using ubiquitous compute devices), cheap and effortlessly distributed, and collaborative. This is achieved by allowing every internet capable device to run training algorithms and predictive models with no software installation and by saving models in universally readable formats. Our prototype library is capable of training deep neural networks with synchronized, distributed stochastic gradient descent. MLitB offers several important opportunities for novel ML research, including: development of distributed learning algorithms, advancement of web GPU algorithms, novel field and mobile applications, privacy preserving computing, and green grid-computing. MLitB is available as open source software.
Harmonic Exponential Families on Manifolds
May 20, 2015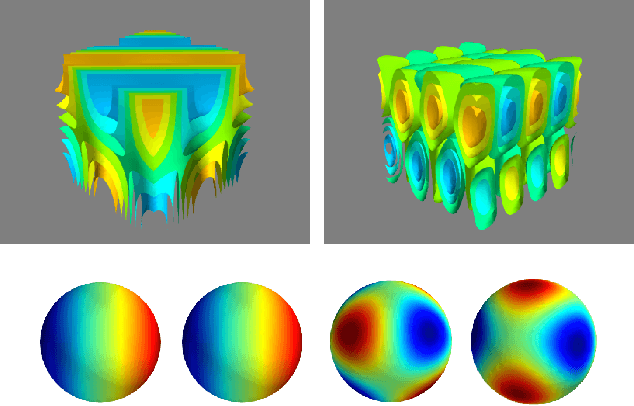
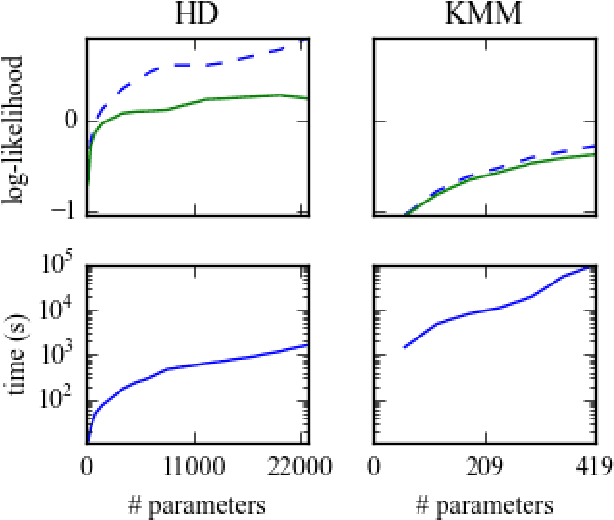
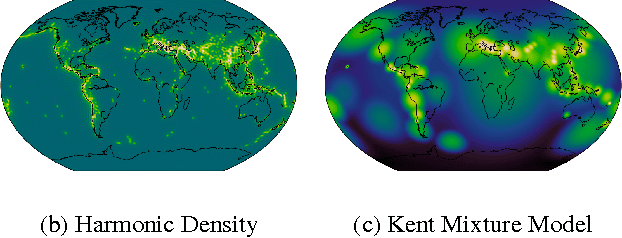
Abstract:In a range of fields including the geosciences, molecular biology, robotics and computer vision, one encounters problems that involve random variables on manifolds. Currently, there is a lack of flexible probabilistic models on manifolds that are fast and easy to train. We define an extremely flexible class of exponential family distributions on manifolds such as the torus, sphere, and rotation groups, and show that for these distributions the gradient of the log-likelihood can be computed efficiently using a non-commutative generalization of the Fast Fourier Transform (FFT). We discuss applications to Bayesian camera motion estimation (where harmonic exponential families serve as conjugate priors), and modelling of the spatial distribution of earthquakes on the surface of the earth. Our experimental results show that harmonic densities yield a significantly higher likelihood than the best competing method, while being orders of magnitude faster to train.
Markov Chain Monte Carlo and Variational Inference: Bridging the Gap
May 19, 2015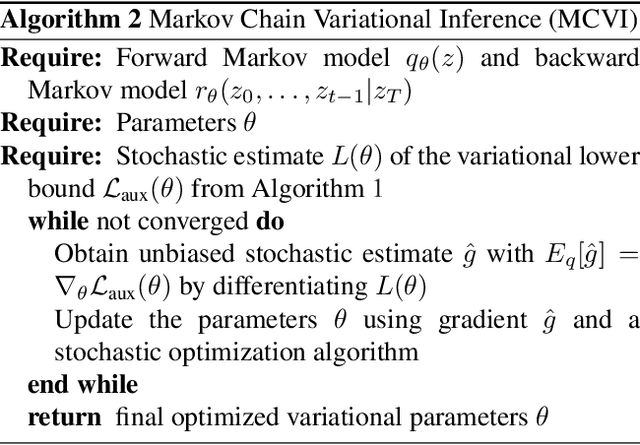
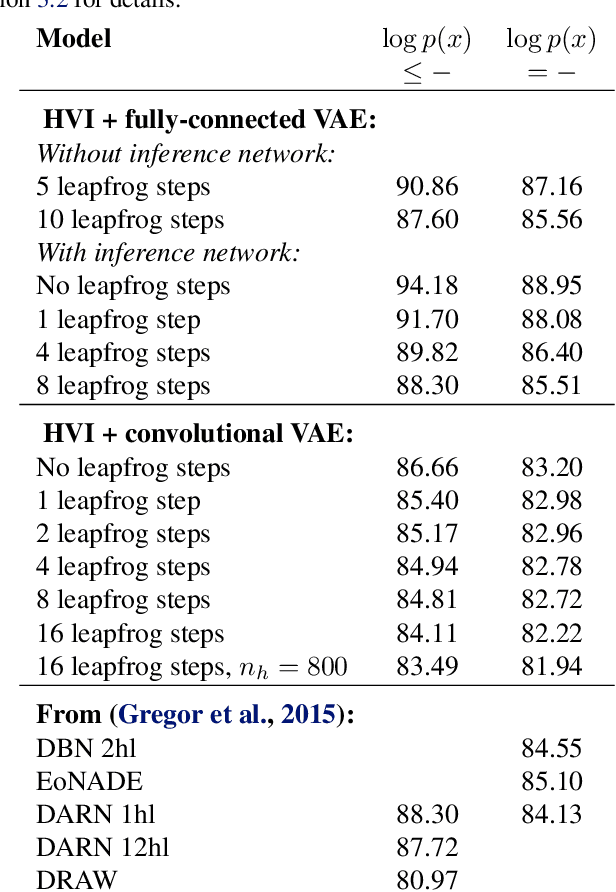

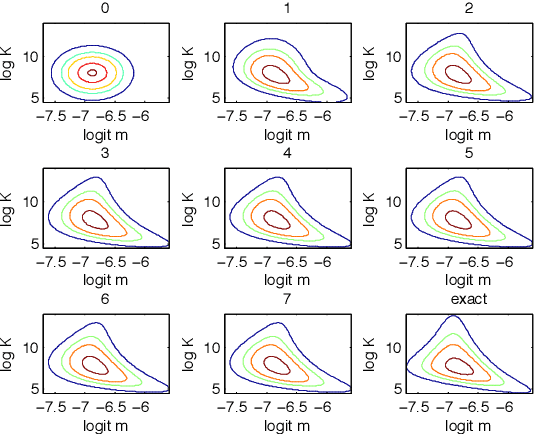
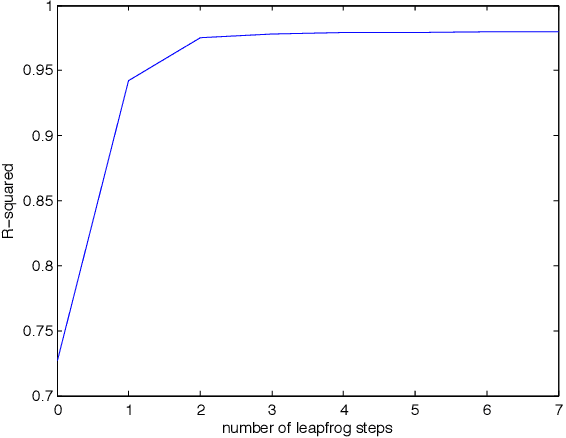
Abstract:Recent advances in stochastic gradient variational inference have made it possible to perform variational Bayesian inference with posterior approximations containing auxiliary random variables. This enables us to explore a new synthesis of variational inference and Monte Carlo methods where we incorporate one or more steps of MCMC into our variational approximation. By doing so we obtain a rich class of inference algorithms bridging the gap between variational methods and MCMC, and offering the best of both worlds: fast posterior approximation through the maximization of an explicit objective, with the option of trading off additional computation for additional accuracy. We describe the theoretical foundations that make this possible and show some promising first results.
Transformation Properties of Learned Visual Representations
Apr 07, 2015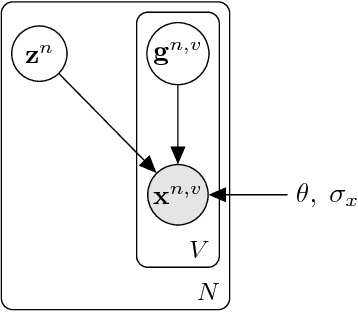
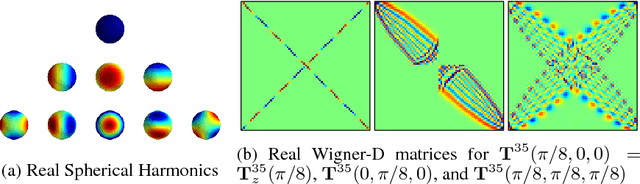

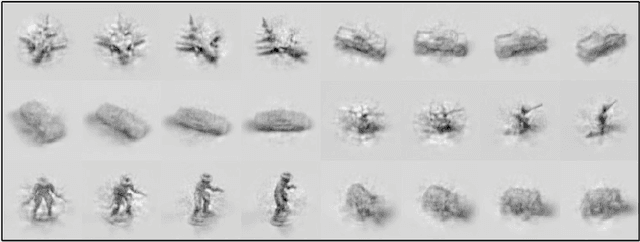
Abstract:When a three-dimensional object moves relative to an observer, a change occurs on the observer's image plane and in the visual representation computed by a learned model. Starting with the idea that a good visual representation is one that transforms linearly under scene motions, we show, using the theory of group representations, that any such representation is equivalent to a combination of the elementary irreducible representations. We derive a striking relationship between irreducibility and the statistical dependency structure of the representation, by showing that under restricted conditions, irreducible representations are decorrelated. Under partial observability, as induced by the perspective projection of a scene onto the image plane, the motion group does not have a linear action on the space of images, so that it becomes necessary to perform inference over a latent representation that does transform linearly. This idea is demonstrated in a model of rotating NORB objects that employs a latent representation of the non-commutative 3D rotation group SO(3).
Large-Scale Distributed Bayesian Matrix Factorization using Stochastic Gradient MCMC
Mar 10, 2015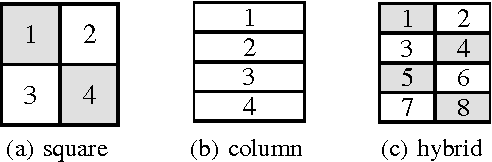
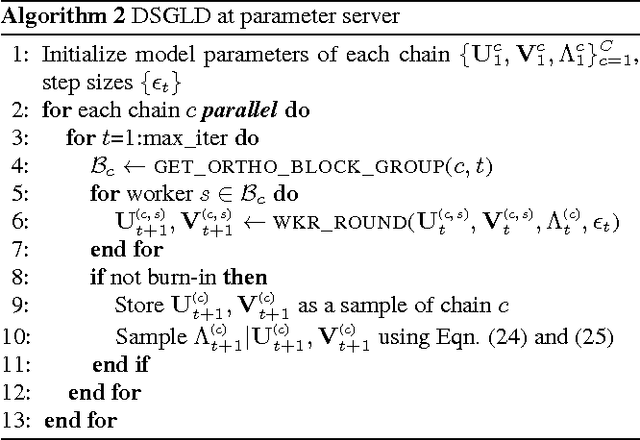

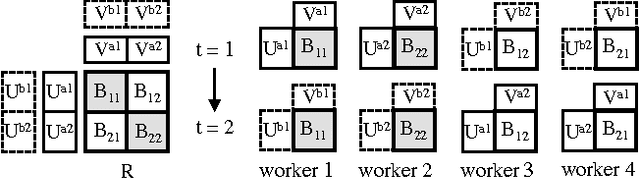
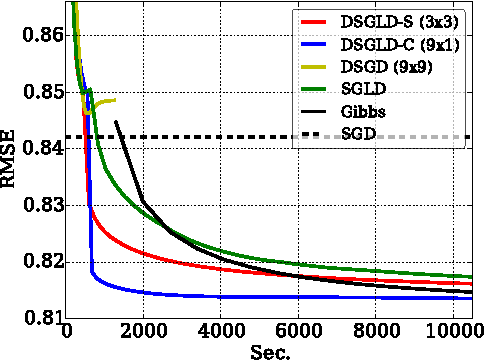
Abstract:Despite having various attractive qualities such as high prediction accuracy and the ability to quantify uncertainty and avoid over-fitting, Bayesian Matrix Factorization has not been widely adopted because of the prohibitive cost of inference. In this paper, we propose a scalable distributed Bayesian matrix factorization algorithm using stochastic gradient MCMC. Our algorithm, based on Distributed Stochastic Gradient Langevin Dynamics, can not only match the prediction accuracy of standard MCMC methods like Gibbs sampling, but at the same time is as fast and simple as stochastic gradient descent. In our experiments, we show that our algorithm can achieve the same level of prediction accuracy as Gibbs sampling an order of magnitude faster. We also show that our method reduces the prediction error as fast as distributed stochastic gradient descent, achieving a 4.1% improvement in RMSE for the Netflix dataset and an 1.8% for the Yahoo music dataset.
Hamiltonian ABC
Mar 06, 2015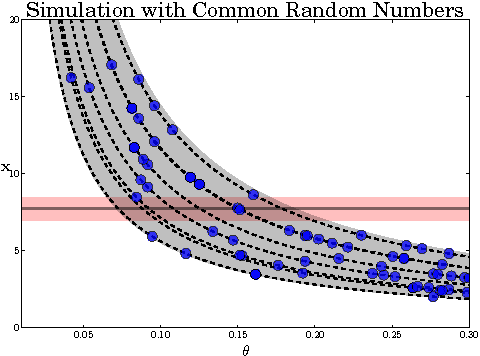
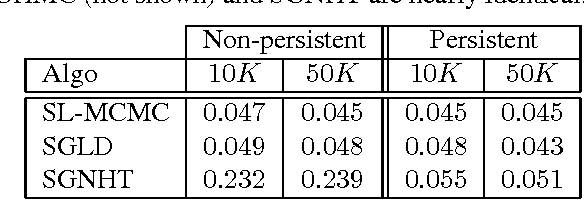
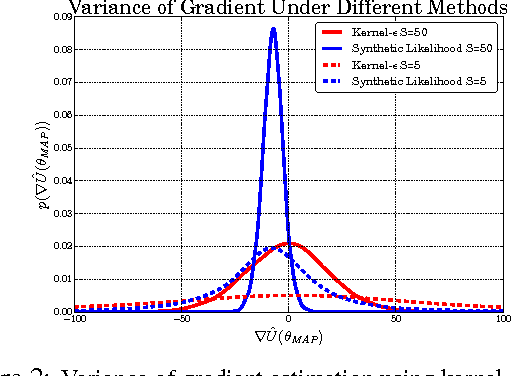

Abstract:Approximate Bayesian computation (ABC) is a powerful and elegant framework for performing inference in simulation-based models. However, due to the difficulty in scaling likelihood estimates, ABC remains useful for relatively low-dimensional problems. We introduce Hamiltonian ABC (HABC), a set of likelihood-free algorithms that apply recent advances in scaling Bayesian learning using Hamiltonian Monte Carlo (HMC) and stochastic gradients. We find that a small number forward simulations can effectively approximate the ABC gradient, allowing Hamiltonian dynamics to efficiently traverse parameter spaces. We also describe a new simple yet general approach of incorporating random seeds into the state of the Markov chain, further reducing the random walk behavior of HABC. We demonstrate HABC on several typical ABC problems, and show that HABC samples comparably to regular Bayesian inference using true gradients on a high-dimensional problem from machine learning.
Efficient Gradient-Based Inference through Transformations between Bayes Nets and Neural Nets
Jan 22, 2015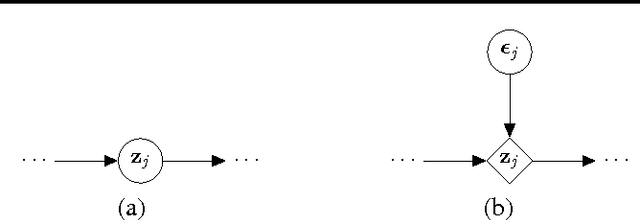
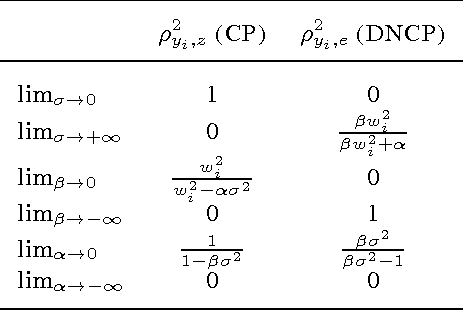
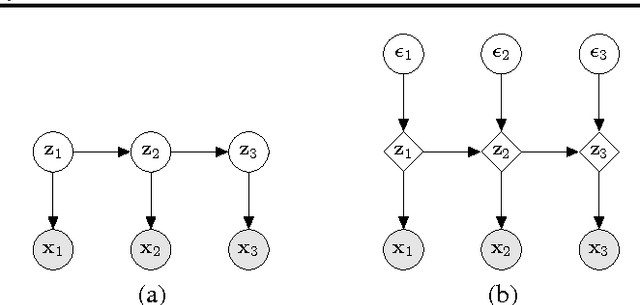
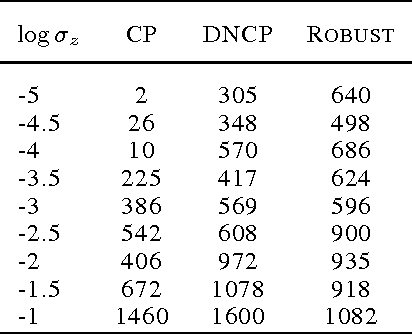
Abstract:Hierarchical Bayesian networks and neural networks with stochastic hidden units are commonly perceived as two separate types of models. We show that either of these types of models can often be transformed into an instance of the other, by switching between centered and differentiable non-centered parameterizations of the latent variables. The choice of parameterization greatly influences the efficiency of gradient-based posterior inference; we show that they are often complementary to eachother, we clarify when each parameterization is preferred and show how inference can be made robust. In the non-centered form, a simple Monte Carlo estimator of the marginal likelihood can be used for learning the parameters. Theoretical results are supported by experiments.
POPE: Post Optimization Posterior Evaluation of Likelihood Free Models
Dec 09, 2014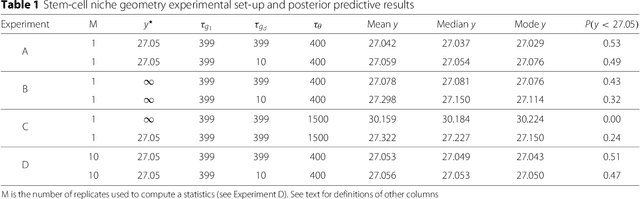
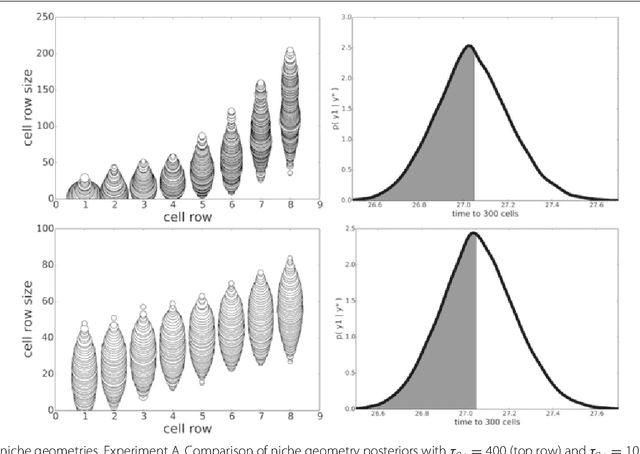
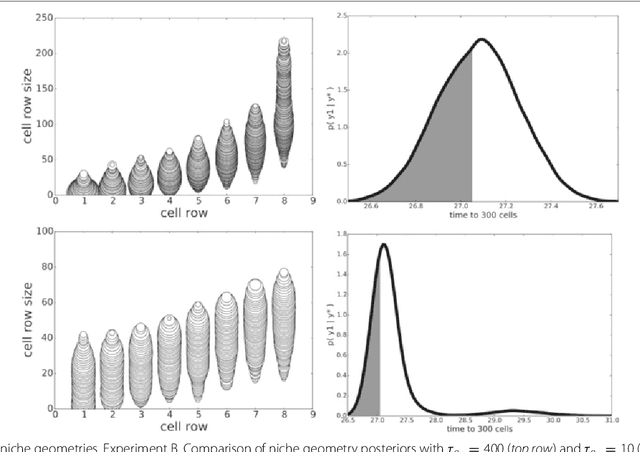
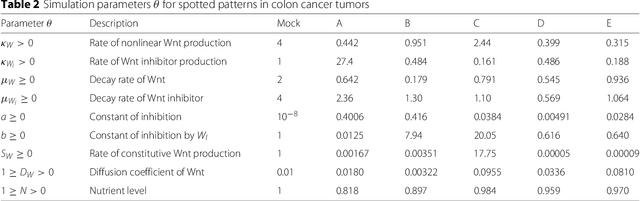
Abstract:In many domains, scientists build complex simulators of natural phenomena that encode their hypotheses about the underlying processes. These simulators can be deterministic or stochastic, fast or slow, constrained or unconstrained, and so on. Optimizing the simulators with respect to a set of parameter values is common practice, resulting in a single parameter setting that minimizes an objective subject to constraints. We propose a post optimization posterior analysis that computes and visualizes all the models that can generate equally good or better simulation results, subject to constraints. These optimization posteriors are desirable for a number of reasons among which easy interpretability, automatic parameter sensitivity and correlation analysis and posterior predictive analysis. We develop a new sampling framework based on approximate Bayesian computation (ABC) with one-sided kernels. In collaboration with two groups of scientists we applied POPE to two important biological simulators: a fast and stochastic simulator of stem-cell cycling and a slow and deterministic simulator of tumor growth patterns.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge