Lucas Zimmer
TumorFlow: Physics-Guided Longitudinal MRI Synthesis of Glioblastoma Growth
Mar 05, 2026Abstract:Glioblastoma exhibits diverse, infiltrative, and patient-specific growth patterns that are only partially visible on routine MRI, making it difficult to reliably assess true tumor extent and personalize treatment planning and follow-up. We present a biophysically-conditioned generative framework that synthesizes biologically realistic 3D brain MRI volumes from estimated, spatially continuous tumor-concentration fields. Our approach combines a generative model with tumor-infiltration maps that can be propagated through time using a biophysical growth model, enabling fine-grained control over tumor shape and growth while preserving patient anatomy. This enables us to synthesize consistent tumor growth trajectories directly in the space of real patients, providing interpretable, controllable estimation of tumor infiltration and progression beyond what is explicitly observed in imaging. We evaluate the framework on longitudinal glioblastoma cases and demonstrate that it can generate temporally coherent sequences with realistic changes in tumor appearance and surrounding tissue response. These results suggest that integrating mechanistic tumor growth priors with modern generative modeling can provide a practical tool for patient-specific progression visualization and for generating controlled synthetic data to support downstream neuro-oncology workflows. In longitudinal extrapolation, we achieve a consistent 75% Dice overlap with the biophysical model while maintaining a constant PSNR of 25 in the surrounding tissue. Our code is available at: https://github.com/valentin-biller/lgm.git
Learn-Morph-Infer: a new way of solving the inverse problem for brain tumor modeling
Nov 07, 2021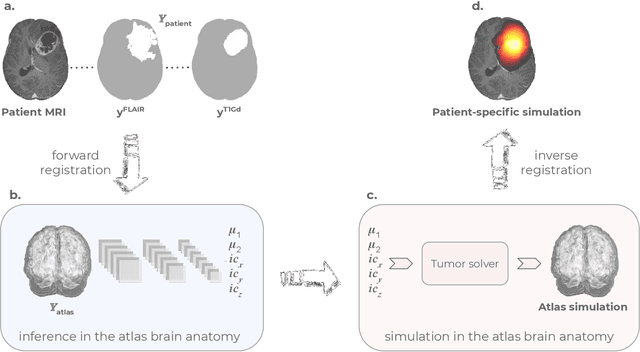
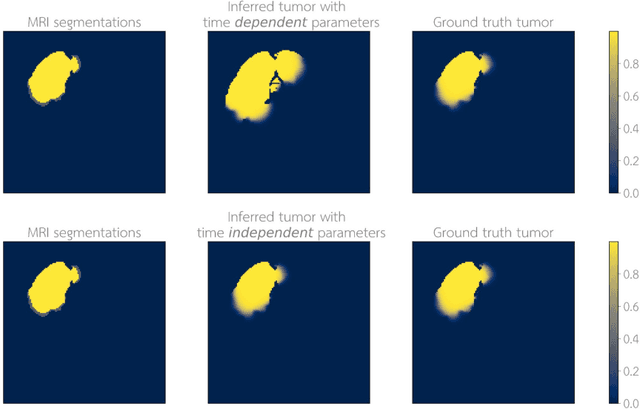
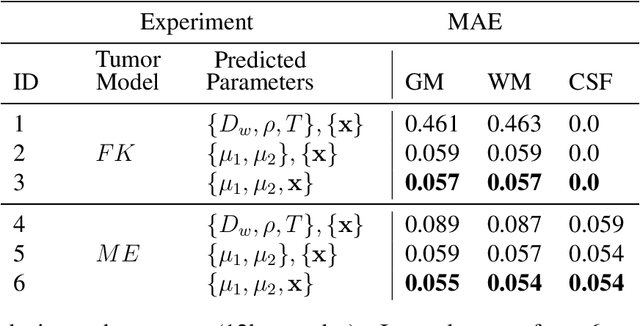
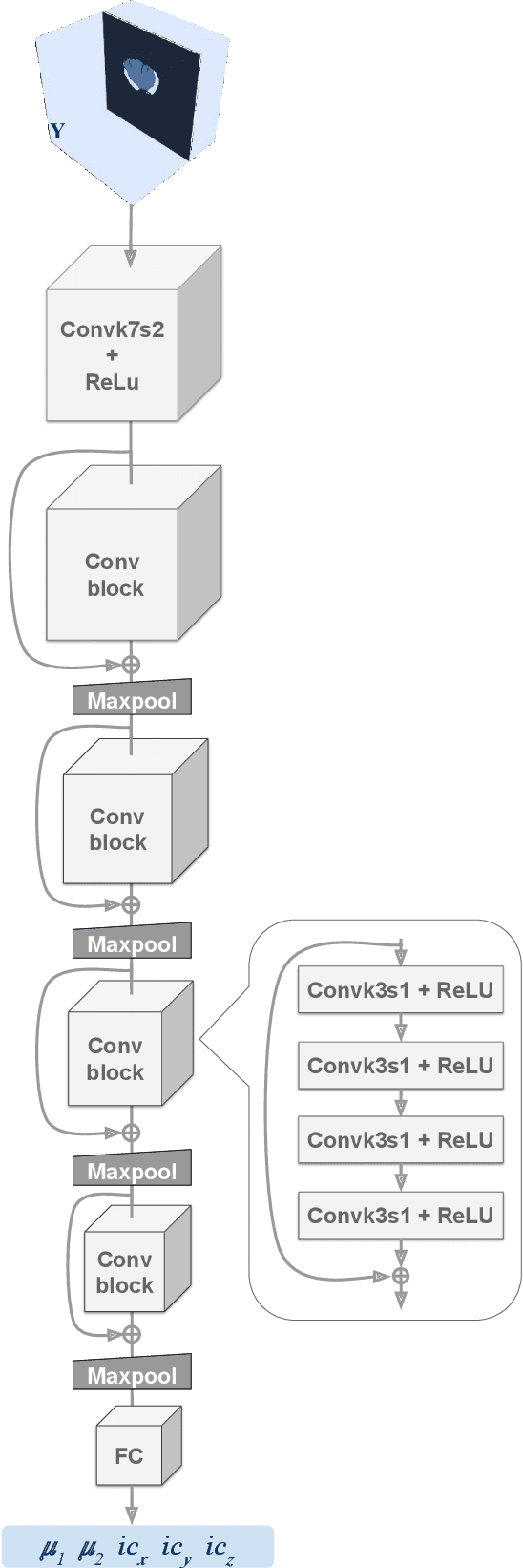

Abstract:Current treatment planning of patients diagnosed with brain tumor could significantly benefit by accessing the spatial distribution of tumor cell concentration. Existing diagnostic modalities, such as magnetic-resonance imaging (MRI), contrast sufficiently well areas of high cell density. However, they do not portray areas of low concentration, which can often serve as a source for the secondary appearance of the tumor after treatment. Numerical simulations of tumor growth could complement imaging information by providing estimates of full spatial distributions of tumor cells. Over recent years a corpus of literature on medical image-based tumor modeling was published. It includes different mathematical formalisms describing the forward tumor growth model. Alongside, various parametric inference schemes were developed to perform an efficient tumor model personalization, i.e. solving the inverse problem. However, the unifying drawback of all existing approaches is the time complexity of the model personalization that prohibits a potential integration of the modeling into clinical settings. In this work, we introduce a methodology for inferring patient-specific spatial distribution of brain tumor from T1Gd and FLAIR MRI medical scans. Coined as \textit{Learn-Morph-Infer} the method achieves real-time performance in the order of minutes on widely available hardware and the compute time is stable across tumor models of different complexity, such as reaction-diffusion and reaction-advection-diffusion models. We believe the proposed inverse solution approach not only bridges the way for clinical translation of brain tumor personalization but can also be adopted to other scientific and engineering domains.
NAS-Bench-301 and the Case for Surrogate Benchmarks for Neural Architecture Search
Aug 22, 2020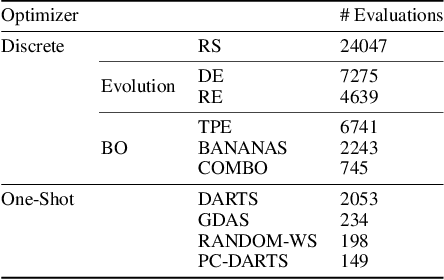
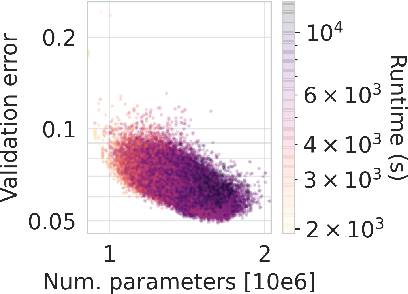
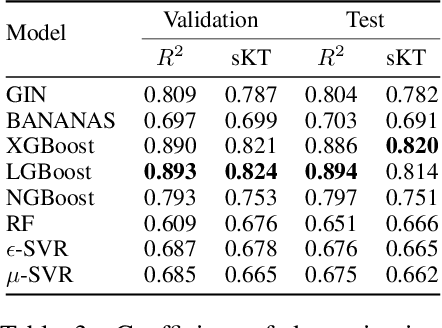
Abstract:Neural Architecture Search (NAS) is a logical next step in the automatic learning of representations, but the development of NAS methods is slowed by high computational demands. As a remedy, several tabular NAS benchmarks were proposed to simulate runs of NAS methods in seconds. However, all existing NAS benchmarks are limited to extremely small architectural spaces since they rely on exhaustive evaluations of the space. This leads to unrealistic results, such as a strong performance of local search and random search, that do not transfer to larger search spaces. To overcome this fundamental limitation, we propose NAS-Bench-301, the first model-based surrogate NAS benchmark, using a search space containing $10^{18}$ architectures, orders of magnitude larger than any previous NAS benchmark. We first motivate the benefits of using such a surrogate benchmark compared to a tabular one by smoothing out the noise stemming from the stochasticity of single SGD runs in a tabular benchmark. Then, we analyze our new dataset consisting of architecture evaluations and comprehensively evaluate various regression models as surrogates to demonstrate their capability to model the architecture space, also using deep ensembles to model uncertainty. Finally, we benchmark a wide range of NAS algorithms using NAS-Bench-301 allowing us to obtain comparable results to the true benchmark at a fraction of the cost.
Auto-PyTorch Tabular: Multi-Fidelity MetaLearning for Efficient and Robust AutoDL
Jun 24, 2020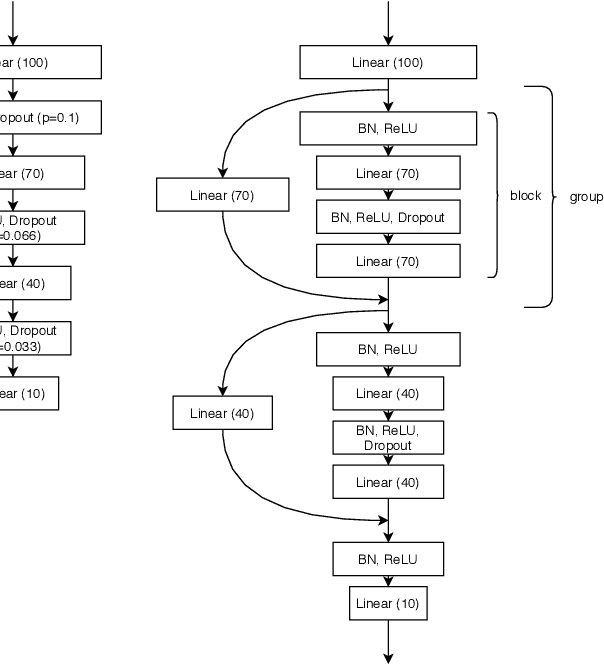
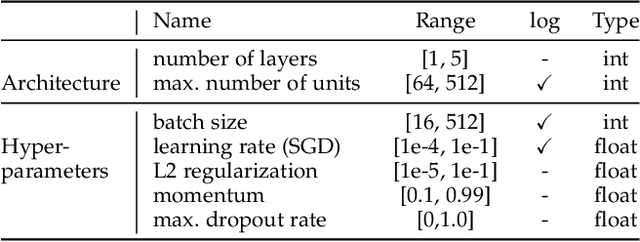
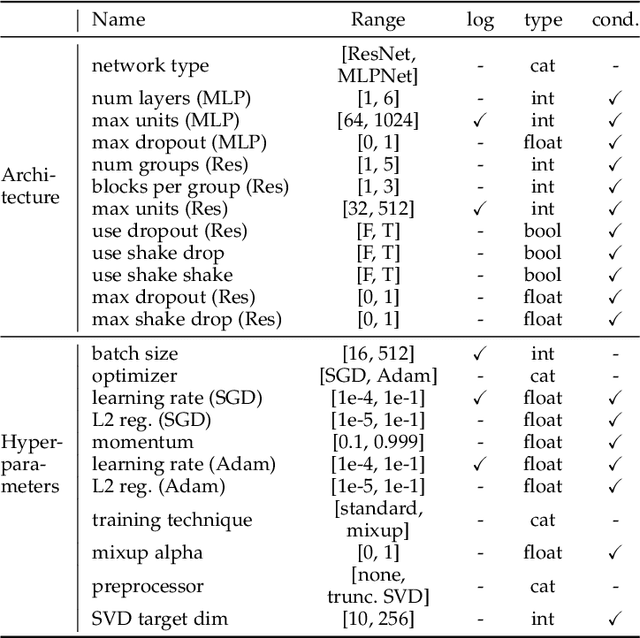
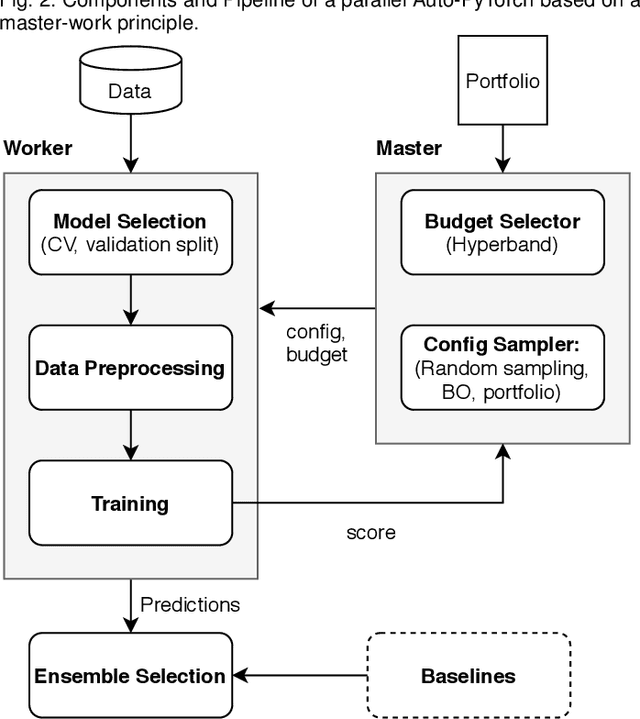
Abstract:While early AutoML frameworks focused on optimizing traditional ML pipelines and their hyperparameters, a recent trend in AutoML is to focus on neural architecture search. In this paper, we introduce Auto-PyTorch, which brings the best of these two worlds together by jointly and robustly optimizing the architecture of networks and the training hyperparameters to enable fully automated deep learning (AutoDL). Auto-PyTorch achieves state-of-the-art performance on several tabular benchmarks by combining multi-fidelity optimization with portfolio construction for warmstarting and ensembling of deep neural networks (DNNs) and common baselines for tabular data. To thoroughly study our assumptions on how to design such an AutoDL system, we additionally introduce a new benchmark on learning curves for DNNs, dubbed LCBench, and run extensive ablation studies of the full Auto-PyTorch on typical AutoML benchmarks, eventually showing that Auto-PyTorch performs better than several state-of-the-art competitors on average.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge