Anke Meyer-Baese
TumorFlow: Physics-Guided Longitudinal MRI Synthesis of Glioblastoma Growth
Mar 05, 2026Abstract:Glioblastoma exhibits diverse, infiltrative, and patient-specific growth patterns that are only partially visible on routine MRI, making it difficult to reliably assess true tumor extent and personalize treatment planning and follow-up. We present a biophysically-conditioned generative framework that synthesizes biologically realistic 3D brain MRI volumes from estimated, spatially continuous tumor-concentration fields. Our approach combines a generative model with tumor-infiltration maps that can be propagated through time using a biophysical growth model, enabling fine-grained control over tumor shape and growth while preserving patient anatomy. This enables us to synthesize consistent tumor growth trajectories directly in the space of real patients, providing interpretable, controllable estimation of tumor infiltration and progression beyond what is explicitly observed in imaging. We evaluate the framework on longitudinal glioblastoma cases and demonstrate that it can generate temporally coherent sequences with realistic changes in tumor appearance and surrounding tissue response. These results suggest that integrating mechanistic tumor growth priors with modern generative modeling can provide a practical tool for patient-specific progression visualization and for generating controlled synthetic data to support downstream neuro-oncology workflows. In longitudinal extrapolation, we achieve a consistent 75% Dice overlap with the biophysical model while maintaining a constant PSNR of 25 in the surrounding tissue. Our code is available at: https://github.com/valentin-biller/lgm.git
Automated detection and segmentation of non-mass enhancing breast tumors with dynamic contrast-enhanced magnetic resonance imaging
Sep 26, 2018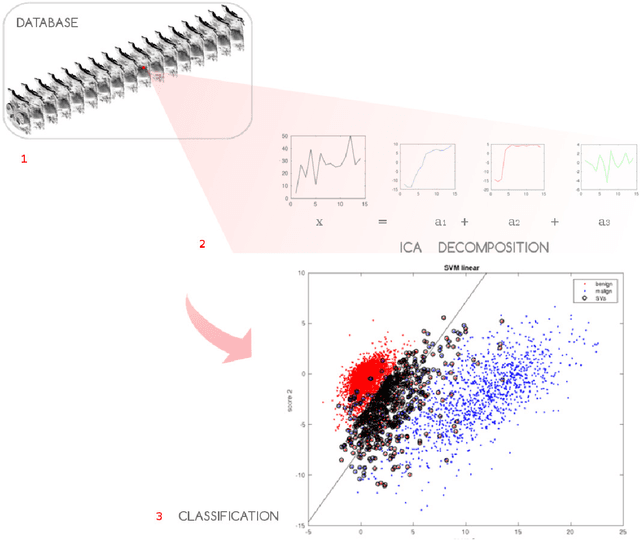
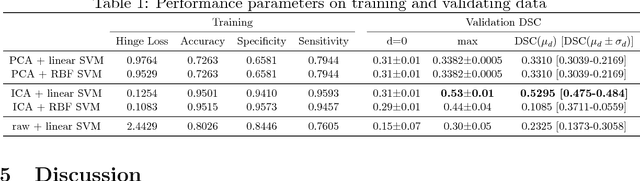
Abstract:Non-mass enhancing lesions (NME) constitute a diagnostic challenge in dynamic contrast enhanced magnetic resonance imaging (DCE-MRI) of the breast. Computer Aided Diagnosis (CAD) systems provide physicians with advanced tools for analysis, assessment and evaluation that have a significant impact on the diagnostic performance. Here, we propose a new approach to address the challenge of NME detection and segmentation, taking advantage of independent component analysis (ICA) to extract data-driven dynamic lesion characterizations. A set of independent sources was obtained from DCE-MRI dataset of breast patients, and the dynamic behavior of the different tissues was described by multiple dynamic curves, together with a set of eigenimages describing the scores for each voxel. A new test image is projected onto the independent source space using the unmixing matrix, and each voxel is classified by a support vector machine (SVM) that has already been trained with manually delineated data. A solution to the high false positive rate problem is proposed by controlling the SVM hyperplane location, outperforming previously published approaches.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge