Kevin Yeh
Anatomy of a failure: When, how, and why deep vision fails in scientific domains
May 05, 2026Abstract:Mirroring its ubiquity in popular media and all human activities, the use of deep learning (DL) is rapidly growing in scientific imaging modalities. However, unlike everyday RGB pictures, pixels encode precise physicochemical properties in scientific imaging across potentially thousands of channels. While DL is well validated on human-centric RGB perceptual tasks, its effectiveness for scientific imaging remains uncertain. Here, we show that the naive application of DL frameworks to scientific images can lead to critical failures. We evaluate the use of DL for pathology, comparing RGB images of stained tissue with the quantitative and information-rich biochemical signatures of infrared (IR) imaging. Despite this informational advantage, DL models trained on IR data paradoxically underperform. We investigate this discrepancy to find that IR data priors interact poorly with the simplicity bias of DL, causing models to collapse to one-dimensional predictions. This constitutes a catastrophic DL failure because the model's representational capacity remains largely unused, while furthermore raising AI safety concerns and undermining the advantages of such scientific modalities. Notably, this problem persists even with state-of-the-art DL robustification strategies, which are primarily designed and validated for RGB imagery and thus inherit the same prior-bias mismatch. This work establishes a framework for understanding the limitations of generic DL in science and advocates for the study of modality-specific failure modes to guide the development of specialized, safe AI algorithms.
Deepfake histological images for enhancing digital pathology
Jun 16, 2022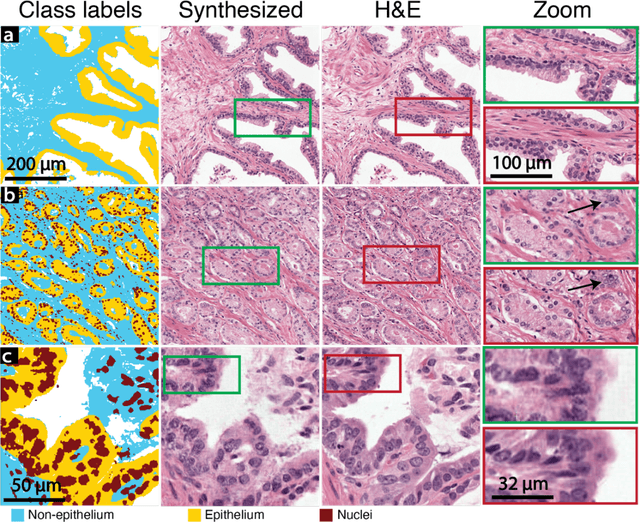
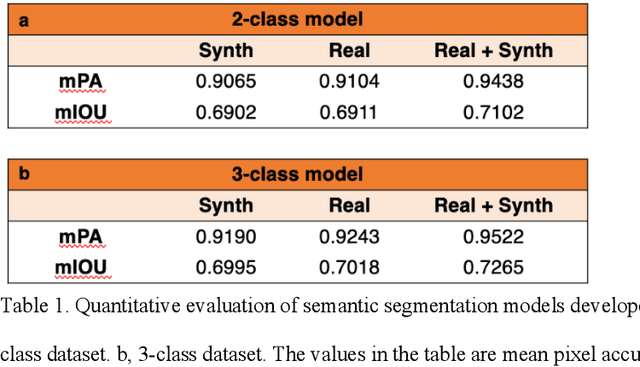
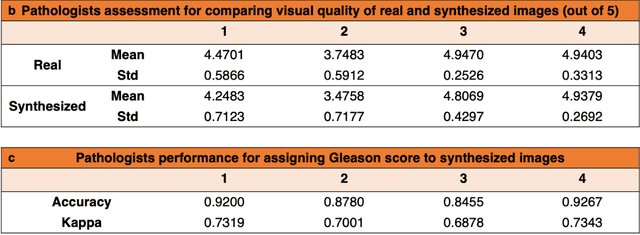
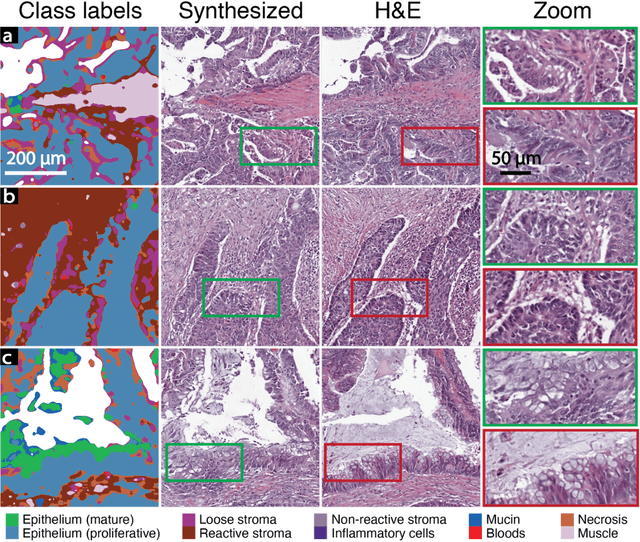
Abstract:An optical microscopic examination of thinly cut stained tissue on glass slides prepared from a FFPE tissue blocks is the gold standard for tissue diagnostics. In addition, the diagnostic abilities and expertise of any pathologist is dependent on their direct experience with common as well as rarer variant morphologies. Recently, deep learning approaches have been used to successfully show a high level of accuracy for such tasks. However, obtaining expert-level annotated images is an expensive and time-consuming task and artificially synthesized histological images can prove greatly beneficial. Here, we present an approach to not only generate histological images that reproduce the diagnostic morphologic features of common disease but also provide a user ability to generate new and rare morphologies. Our approach involves developing a generative adversarial network model that synthesizes pathology images constrained by class labels. We investigated the ability of this framework in synthesizing realistic prostate and colon tissue images and assessed the utility of these images in augmenting diagnostic ability of machine learning methods as well as their usability by a panel of experienced anatomic pathologists. Synthetic data generated by our framework performed similar to real data in training a deep learning model for diagnosis. Pathologists were not able to distinguish between real and synthetic images and showed a similar level of inter-observer agreement for prostate cancer grading. We extended the approach to significantly more complex images from colon biopsies and showed that the complex microenvironment in such tissues can also be reproduced. Finally, we present the ability for a user to generate deepfake histological images via a simple markup of sematic labels.
A deep learning framework for morphologic detail beyond the diffraction limit in infrared spectroscopic imaging
Dec 19, 2019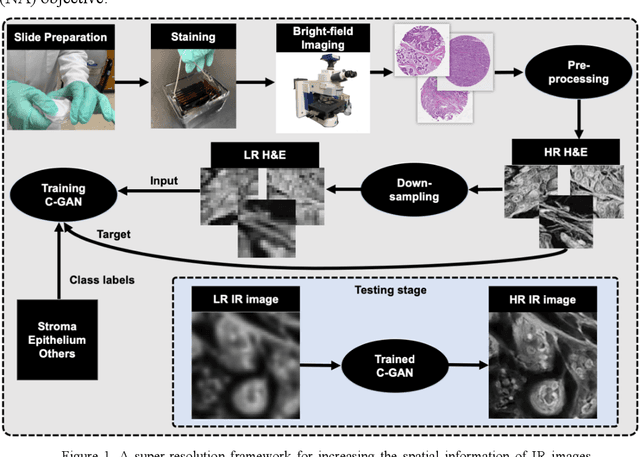
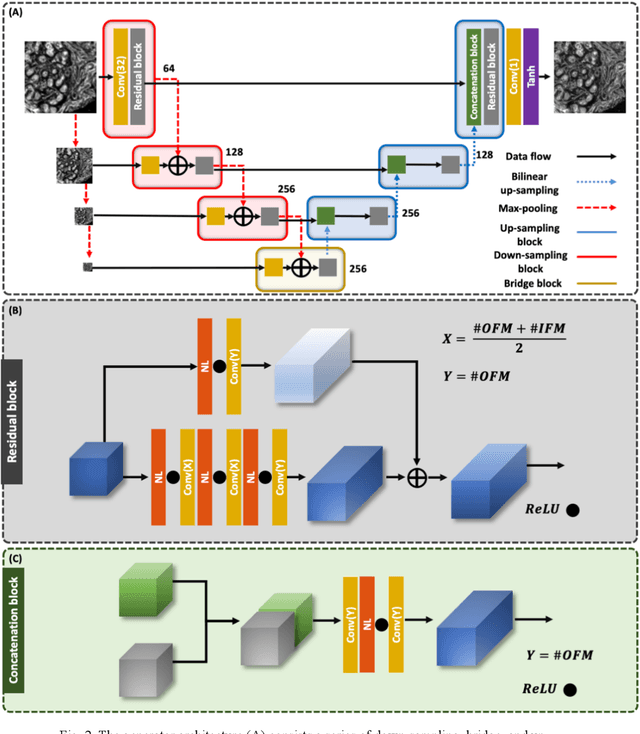
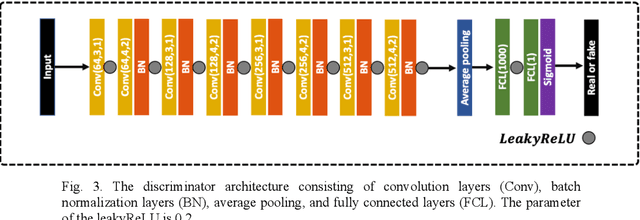
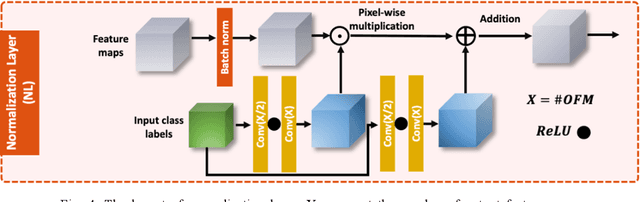
Abstract:Infrared (IR) microscopes measure spectral information that quantifies molecular content to assign the identity of biomedical cells but lack the spatial quality of optical microscopy to appreciate morphologic features. Here, we propose a method to utilize the semantic information of cellular identity from IR imaging with the morphologic detail of pathology images in a deep learning-based approach to image super-resolution. Using Generative Adversarial Networks (GANs), we enhance the spatial detail in IR imaging beyond the diffraction limit while retaining their spectral contrast. This technique can be rapidly integrated with modern IR microscopes to provide a framework useful for routine pathology.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge