Kathy Lee
Large Scale Automated Reading of Frontal and Lateral Chest X-Rays using Dual Convolutional Neural Networks
Apr 24, 2018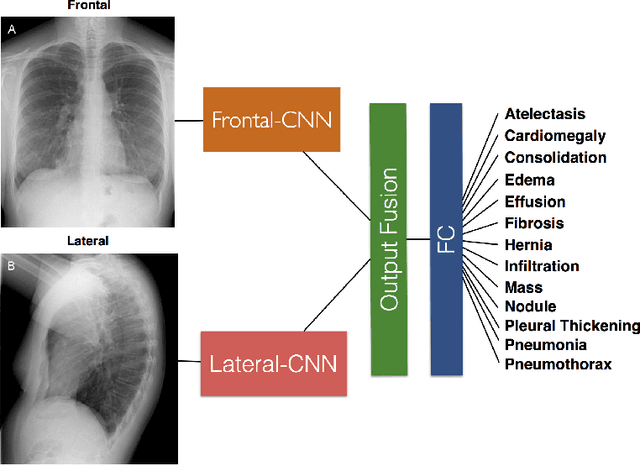
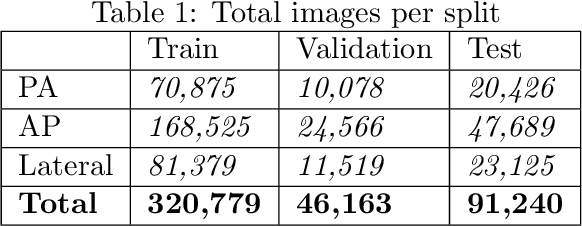
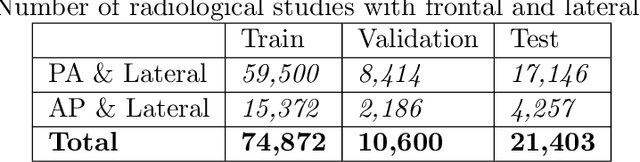

Abstract:The MIMIC-CXR dataset is (to date) the largest released chest x-ray dataset consisting of 473,064 chest x-rays and 206,574 radiology reports collected from 63,478 patients. We present the results of training and evaluating a collection of deep convolutional neural networks on this dataset to recognize multiple common thorax diseases. To the best of our knowledge, this is the first work that trains CNNs for this task on such a large collection of chest x-ray images, which is over four times the size of the largest previously released chest x-ray corpus (ChestX-Ray14). We describe and evaluate individual CNN models trained on frontal and lateral CXR view types. In addition, we present a novel DualNet architecture that emulates routine clinical practice by simultaneously processing both frontal and lateral CXR images obtained from a radiological exam. Our DualNet architecture shows improved performance in recognizing findings in CXR images when compared to applying separate baseline frontal and lateral classifiers.
DR-BiLSTM: Dependent Reading Bidirectional LSTM for Natural Language Inference
Apr 11, 2018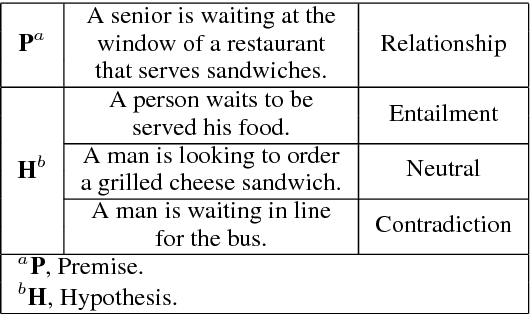
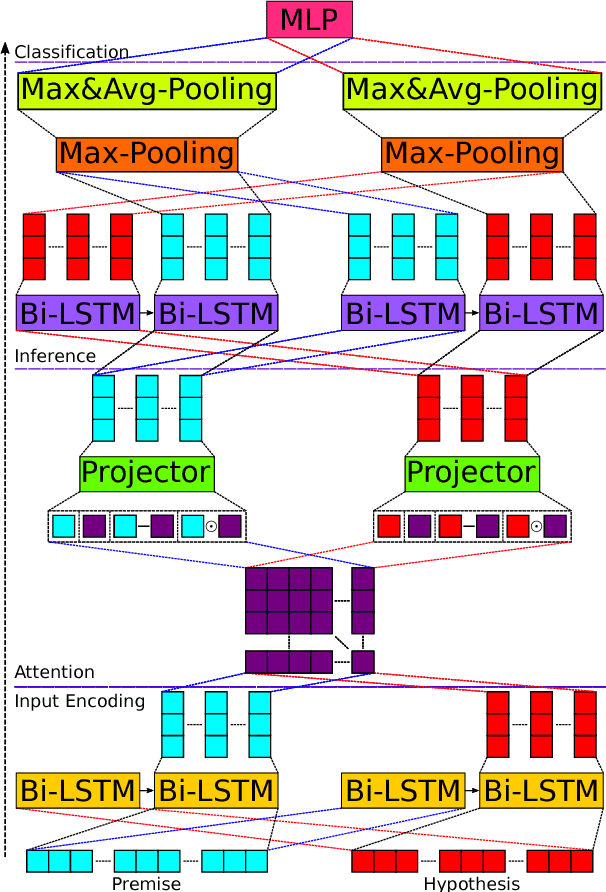
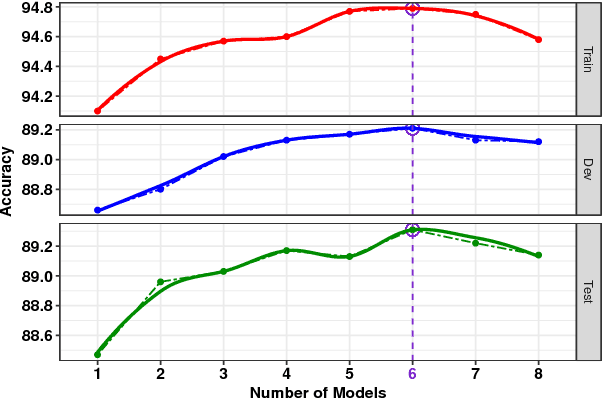
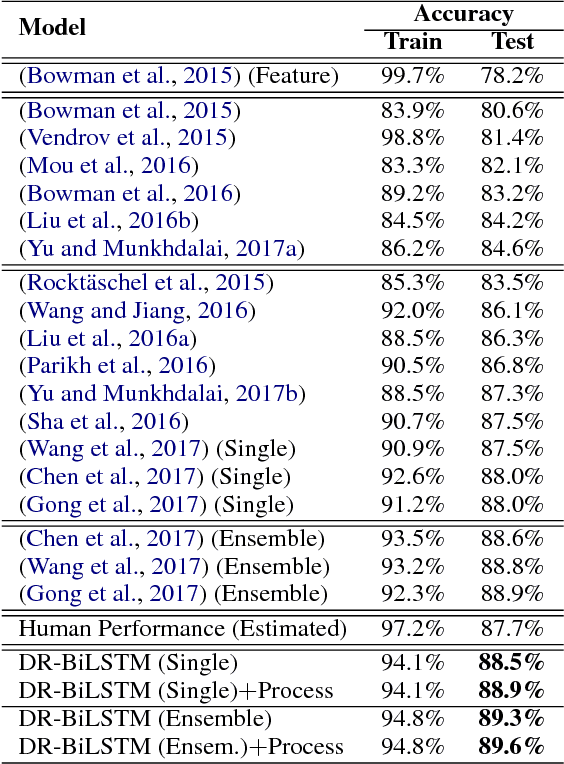
Abstract:We present a novel deep learning architecture to address the natural language inference (NLI) task. Existing approaches mostly rely on simple reading mechanisms for independent encoding of the premise and hypothesis. Instead, we propose a novel dependent reading bidirectional LSTM network (DR-BiLSTM) to efficiently model the relationship between a premise and a hypothesis during encoding and inference. We also introduce a sophisticated ensemble strategy to combine our proposed models, which noticeably improves final predictions. Finally, we demonstrate how the results can be improved further with an additional preprocessing step. Our evaluation shows that DR-BiLSTM obtains the best single model and ensemble model results achieving the new state-of-the-art scores on the Stanford NLI dataset.
Condensed Memory Networks for Clinical Diagnostic Inferencing
Jan 03, 2017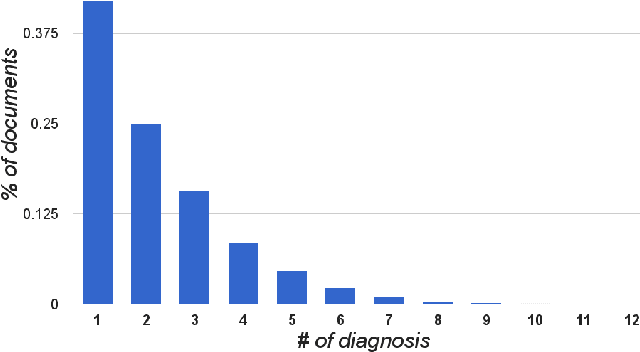
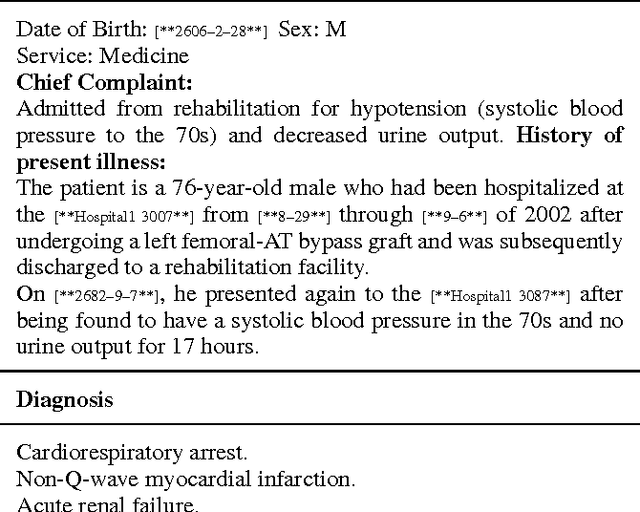
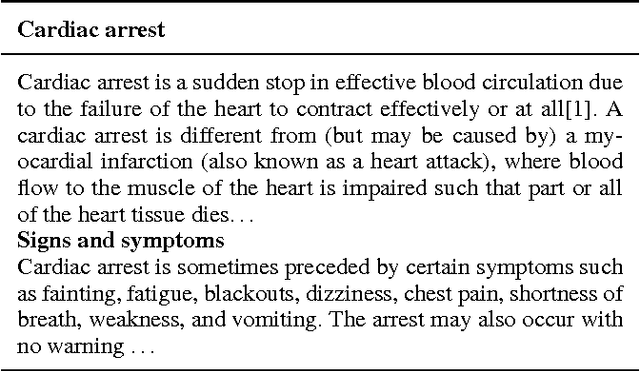


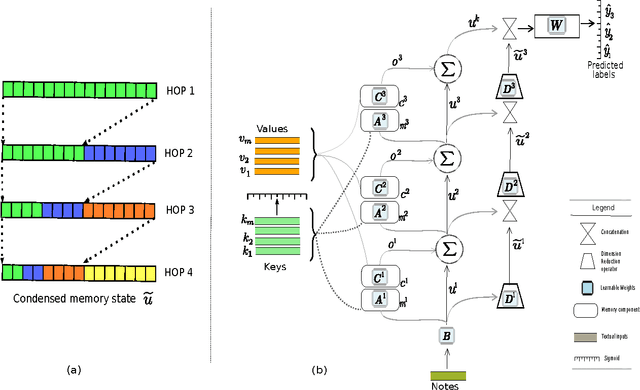
Abstract:Diagnosis of a clinical condition is a challenging task, which often requires significant medical investigation. Previous work related to diagnostic inferencing problems mostly consider multivariate observational data (e.g. physiological signals, lab tests etc.). In contrast, we explore the problem using free-text medical notes recorded in an electronic health record (EHR). Complex tasks like these can benefit from structured knowledge bases, but those are not scalable. We instead exploit raw text from Wikipedia as a knowledge source. Memory networks have been demonstrated to be effective in tasks which require comprehension of free-form text. They use the final iteration of the learned representation to predict probable classes. We introduce condensed memory neural networks (C-MemNNs), a novel model with iterative condensation of memory representations that preserves the hierarchy of features in the memory. Experiments on the MIMIC-III dataset show that the proposed model outperforms other variants of memory networks to predict the most probable diagnoses given a complex clinical scenario.
Neural Paraphrase Generation with Stacked Residual LSTM Networks
Oct 13, 2016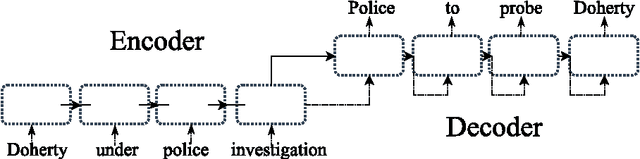

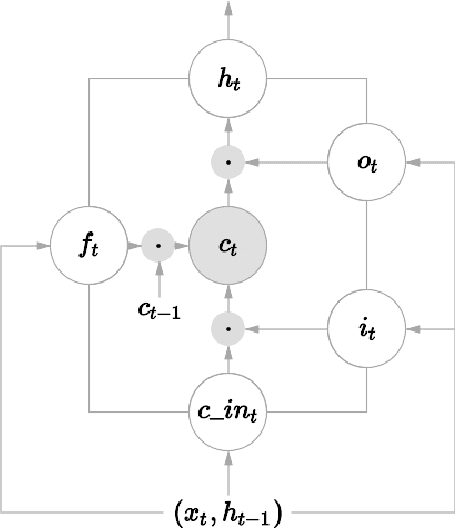
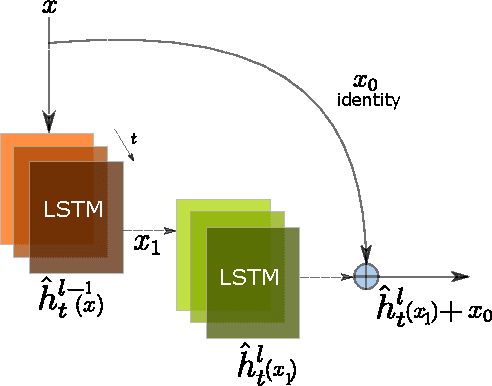
Abstract:In this paper, we propose a novel neural approach for paraphrase generation. Conventional para- phrase generation methods either leverage hand-written rules and thesauri-based alignments, or use statistical machine learning principles. To the best of our knowledge, this work is the first to explore deep learning models for paraphrase generation. Our primary contribution is a stacked residual LSTM network, where we add residual connections between LSTM layers. This allows for efficient training of deep LSTMs. We evaluate our model and other state-of-the-art deep learning models on three different datasets: PPDB, WikiAnswers and MSCOCO. Evaluation results demonstrate that our model outperforms sequence to sequence, attention-based and bi- directional LSTM models on BLEU, METEOR, TER and an embedding-based sentence similarity metric.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge