Jun Lian
SinoSynth: A Physics-based Domain Randomization Approach for Generalizable CBCT Image Enhancement
Sep 27, 2024



Abstract:Cone Beam Computed Tomography (CBCT) finds diverse applications in medicine. Ensuring high image quality in CBCT scans is essential for accurate diagnosis and treatment delivery. Yet, the susceptibility of CBCT images to noise and artifacts undermines both their usefulness and reliability. Existing methods typically address CBCT artifacts through image-to-image translation approaches. These methods, however, are limited by the artifact types present in the training data, which may not cover the complete spectrum of CBCT degradations stemming from variations in imaging protocols. Gathering additional data to encompass all possible scenarios can often pose a challenge. To address this, we present SinoSynth, a physics-based degradation model that simulates various CBCT-specific artifacts to generate a diverse set of synthetic CBCT images from high-quality CT images without requiring pre-aligned data. Through extensive experiments, we demonstrate that several different generative networks trained on our synthesized data achieve remarkable results on heterogeneous multi-institutional datasets, outperforming even the same networks trained on actual data. We further show that our degradation model conveniently provides an avenue to enforce anatomical constraints in conditional generative models, yielding high-quality and structure-preserving synthetic CT images.
OpenKBP-Opt: An international and reproducible evaluation of 76 knowledge-based planning pipelines
Feb 16, 2022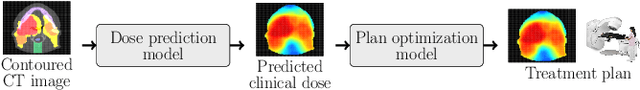

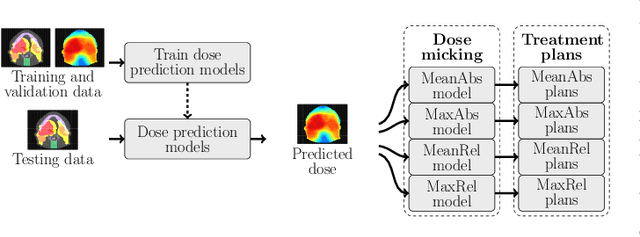
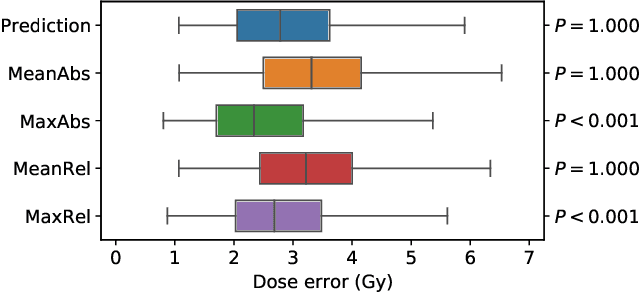
Abstract:We establish an open framework for developing plan optimization models for knowledge-based planning (KBP) in radiotherapy. Our framework includes reference plans for 100 patients with head-and-neck cancer and high-quality dose predictions from 19 KBP models that were developed by different research groups during the OpenKBP Grand Challenge. The dose predictions were input to four optimization models to form 76 unique KBP pipelines that generated 7600 plans. The predictions and plans were compared to the reference plans via: dose score, which is the average mean absolute voxel-by-voxel difference in dose a model achieved; the deviation in dose-volume histogram (DVH) criterion; and the frequency of clinical planning criteria satisfaction. We also performed a theoretical investigation to justify our dose mimicking models. The range in rank order correlation of the dose score between predictions and their KBP pipelines was 0.50 to 0.62, which indicates that the quality of the predictions is generally positively correlated with the quality of the plans. Additionally, compared to the input predictions, the KBP-generated plans performed significantly better (P<0.05; one-sided Wilcoxon test) on 18 of 23 DVH criteria. Similarly, each optimization model generated plans that satisfied a higher percentage of criteria than the reference plans. Lastly, our theoretical investigation demonstrated that the dose mimicking models generated plans that are also optimal for a conventional planning model. This was the largest international effort to date for evaluating the combination of KBP prediction and optimization models. In the interest of reproducibility, our data and code is freely available at https://github.com/ababier/open-kbp-opt.
Dual Shape Guided Segmentation Network for Organs-at-Risk in Head and Neck CT Images
Oct 23, 2021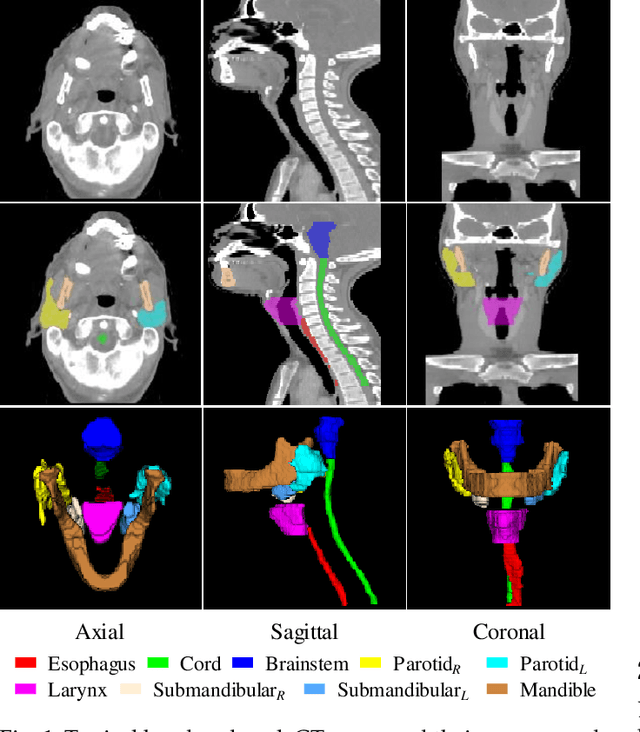
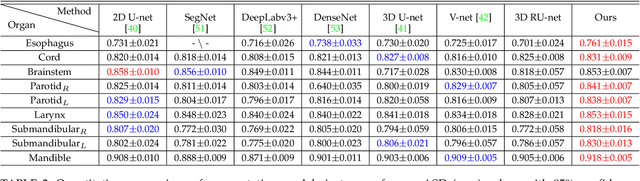
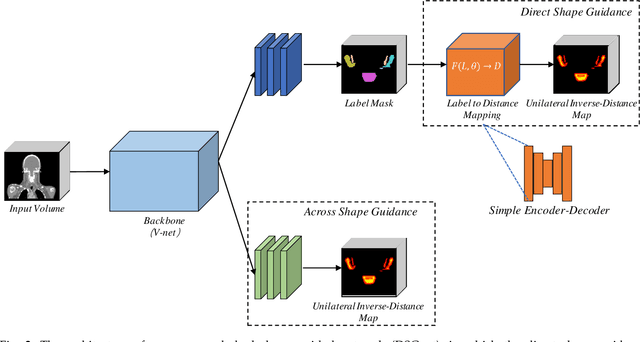
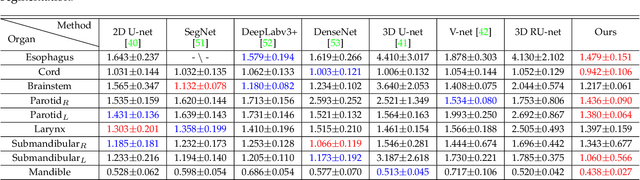
Abstract:The accurate segmentation of organs-at-risk (OARs) in head and neck CT images is a critical step for radiation therapy of head and neck cancer patients. However, manual delineation for numerous OARs is time-consuming and laborious, even for expert oncologists. Moreover, manual delineation results are susceptible to high intra- and inter-variability. To this end, we propose a novel dual shape guided network (DSGnet) to automatically delineate nine important OARs in head and neck CT images. To deal with the large shape variation and unclear boundary of OARs in CT images, we represent the organ shape using an organ-specific unilateral inverse-distance map (UIDM) and guide the segmentation task from two different perspectives: direct shape guidance by following the segmentation prediction and across shape guidance by sharing the segmentation feature. In the direct shape guidance, the segmentation prediction is not only supervised by the true label mask, but also by the true UIDM, which is implemented through a simple yet effective encoder-decoder mapping from the label space to the distance space. In the across shape guidance, UIDM is used to facilitate the segmentation by optimizing the shared feature maps. For the experiments, we build a large head and neck CT dataset with a total of 699 images from different volunteers, and conduct comprehensive experiments and comparisons with other state-of-the-art methods to justify the effectiveness and efficiency of our proposed method. The overall Dice Similarity Coefficient (DSC) value of 0.842 across the nine important OARs demonstrates great potential applications in improving the delineation quality and reducing the time cost.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge