Juergen Hahn
Integrating AI in College Education: Positive yet Mixed Experiences with ChatGPT
Jul 08, 2024

Abstract:The integration of artificial intelligence (AI) chatbots into higher education marks a shift towards a new generation of pedagogical tools, mirroring the arrival of milestones like the internet. With the launch of ChatGPT-4 Turbo in November 2023, we developed a ChatGPT-based teaching application (https://chat.openai.com/g/g-1imx1py4K-chatge-medical-imaging) and integrated it into our undergraduate medical imaging course in the Spring 2024 semester. This study investigates the use of ChatGPT throughout a semester-long trial, providing insights into students' engagement, perception, and the overall educational effectiveness of the technology. We systematically collected and analyzed data concerning students' interaction with ChatGPT, focusing on their attitudes, concerns, and usage patterns. The findings indicate that ChatGPT offers significant advantages such as improved information access and increased interactivity, but its adoption is accompanied by concerns about the accuracy of the information provided and the necessity for well-defined guidelines to optimize its use.
Multimodal Neurodegenerative Disease Subtyping Explained by ChatGPT
Jan 31, 2024Abstract:Alzheimer's disease (AD) is the most prevalent neurodegenerative disease; yet its currently available treatments are limited to stopping disease progression. Moreover, effectiveness of these treatments is not guaranteed due to the heterogenetiy of the disease. Therefore, it is essential to be able to identify the disease subtypes at a very early stage. Current data driven approaches are able to classify the subtypes at later stages of AD or related disorders, but struggle when predicting at the asymptomatic or prodromal stage. Moreover, most existing models either lack explainability behind the classification or only use a single modality for the assessment, limiting scope of its analysis. Thus, we propose a multimodal framework that uses early-stage indicators such as imaging, genetics and clinical assessments to classify AD patients into subtypes at early stages. Similarly, we build prompts and use large language models, such as ChatGPT, to interpret the findings of our model. In our framework, we propose a tri-modal co-attention mechanism (Tri-COAT) to explicitly learn the cross-modal feature associations. Our proposed model outperforms baseline models and provides insight into key cross-modal feature associations supported by known biological mechanisms.
Noise Entangled GAN For Low-Dose CT Simulation
Feb 18, 2021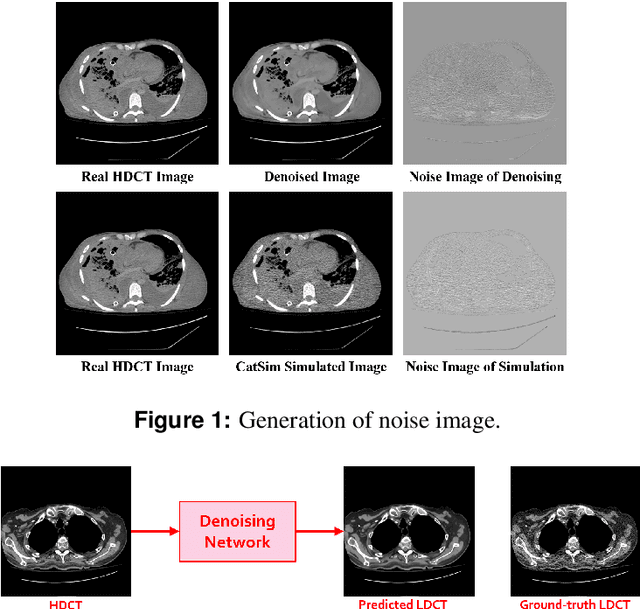
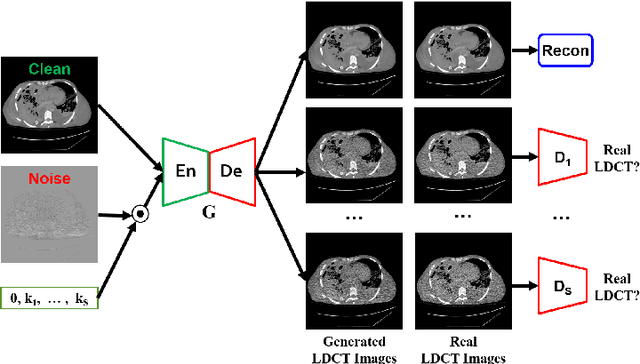
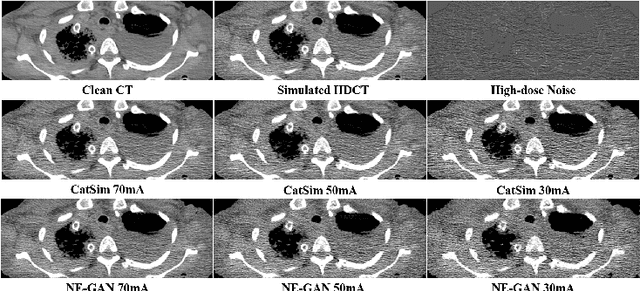
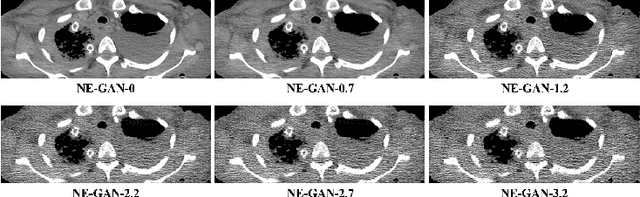
Abstract:We propose a Noise Entangled GAN (NE-GAN) for simulating low-dose computed tomography (CT) images from a higher dose CT image. First, we present two schemes to generate a clean CT image and a noise image from the high-dose CT image. Then, given these generated images, an NE-GAN is proposed to simulate different levels of low-dose CT images, where the level of generated noise can be continuously controlled by a noise factor. NE-GAN consists of a generator and a set of discriminators, and the number of discriminators is determined by the number of noise levels during training. Compared with the traditional methods based on the projection data that are usually unavailable in real applications, NE-GAN can directly learn from the real and/or simulated CT images and may create low-dose CT images quickly without the need of raw data or other proprietary CT scanner information. The experimental results show that the proposed method has the potential to simulate realistic low-dose CT images.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge