Jose Dolz
École de Technologie Supérieure, Montreal, Canada
Weakly supervised segmentation with cross-modality equivariant constraints
Apr 06, 2021
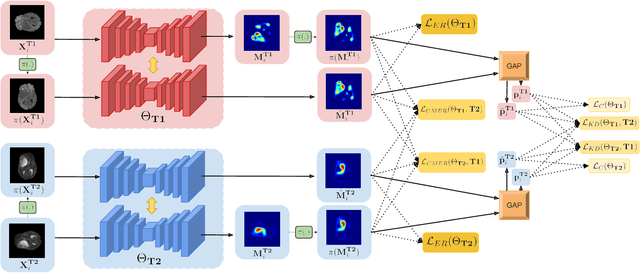
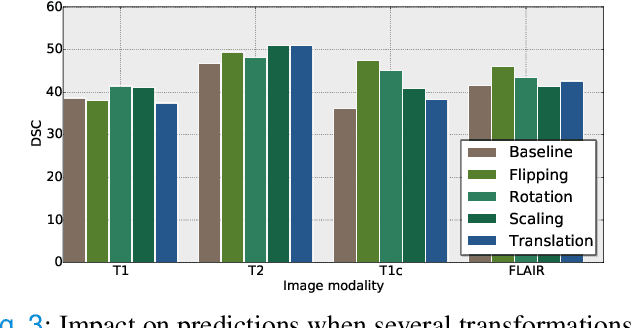
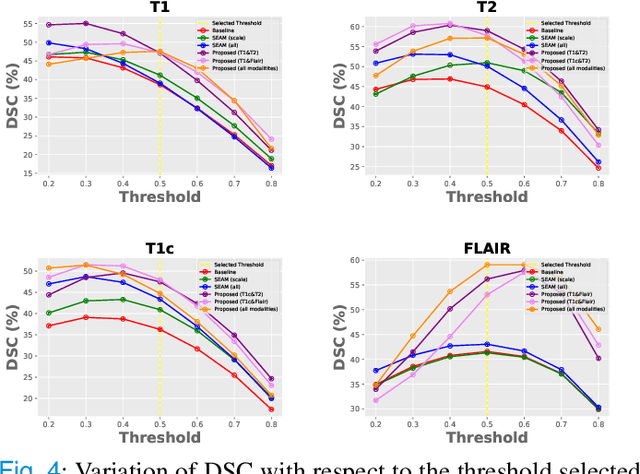
Abstract:Weakly supervised learning has emerged as an appealing alternative to alleviate the need for large labeled datasets in semantic segmentation. Most current approaches exploit class activation maps (CAMs), which can be generated from image-level annotations. Nevertheless, resulting maps have been demonstrated to be highly discriminant, failing to serve as optimal proxy pixel-level labels. We present a novel learning strategy that leverages self-supervision in a multi-modal image scenario to significantly enhance original CAMs. In particular, the proposed method is based on two observations. First, the learning of fully-supervised segmentation networks implicitly imposes equivariance by means of data augmentation, whereas this implicit constraint disappears on CAMs generated with image tags. And second, the commonalities between image modalities can be employed as an efficient self-supervisory signal, correcting the inconsistency shown by CAMs obtained across multiple modalities. To effectively train our model, we integrate a novel loss function that includes a within-modality and a cross-modality equivariant term to explicitly impose these constraints during training. In addition, we add a KL-divergence on the class prediction distributions to facilitate the information exchange between modalities, which, combined with the equivariant regularizers further improves the performance of our model. Exhaustive experiments on the popular multi-modal BRATS dataset demonstrate that our approach outperforms relevant recent literature under the same learning conditions.
Knowledge Distillation Methods for Efficient Unsupervised Adaptation Across Multiple Domains
Jan 18, 2021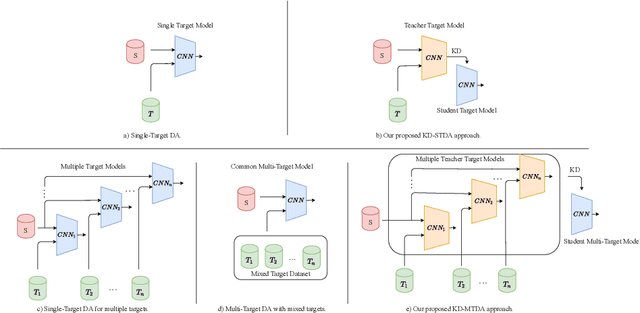
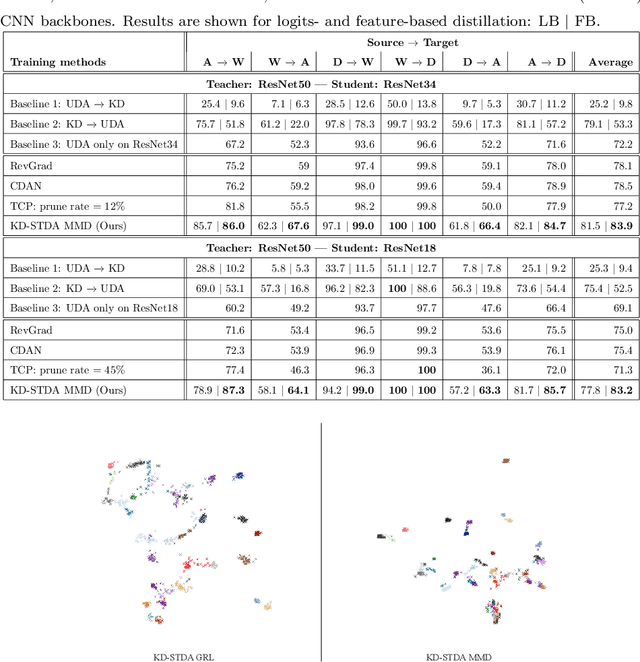
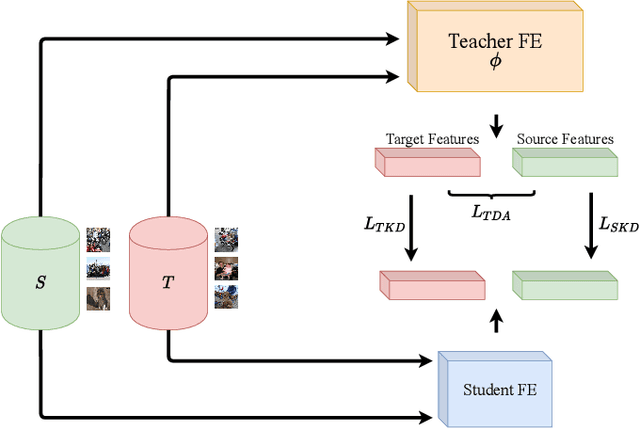
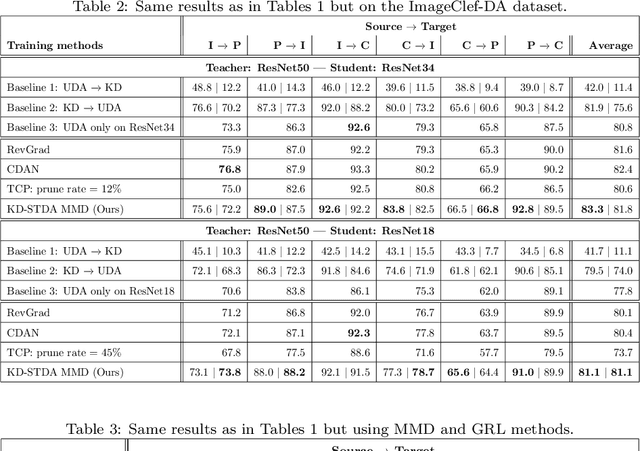
Abstract:Beyond the complexity of CNNs that require training on large annotated datasets, the domain shift between design and operational data has limited the adoption of CNNs in many real-world applications. For instance, in person re-identification, videos are captured over a distributed set of cameras with non-overlapping viewpoints. The shift between the source (e.g. lab setting) and target (e.g. cameras) domains may lead to a significant decline in recognition accuracy. Additionally, state-of-the-art CNNs may not be suitable for such real-time applications given their computational requirements. Although several techniques have recently been proposed to address domain shift problems through unsupervised domain adaptation (UDA), or to accelerate/compress CNNs through knowledge distillation (KD), we seek to simultaneously adapt and compress CNNs to generalize well across multiple target domains. In this paper, we propose a progressive KD approach for unsupervised single-target DA (STDA) and multi-target DA (MTDA) of CNNs. Our method for KD-STDA adapts a CNN to a single target domain by distilling from a larger teacher CNN, trained on both target and source domain data in order to maintain its consistency with a common representation. Our proposed approach is compared against state-of-the-art methods for compression and STDA of CNNs on the Office31 and ImageClef-DA image classification datasets. It is also compared against state-of-the-art methods for MTDA on Digits, Office31, and OfficeHome. In both settings -- KD-STDA and KD-MTDA -- results indicate that our approach can achieve the highest level of accuracy across target domains, while requiring a comparable or lower CNN complexity.
Bladder segmentation based on deep learning approaches: current limitations and lessons
Jan 16, 2021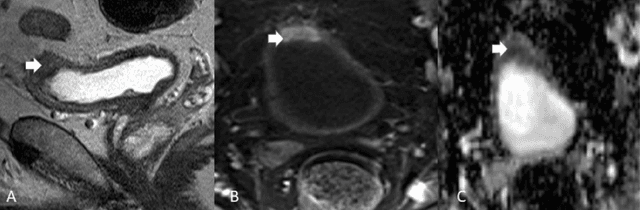
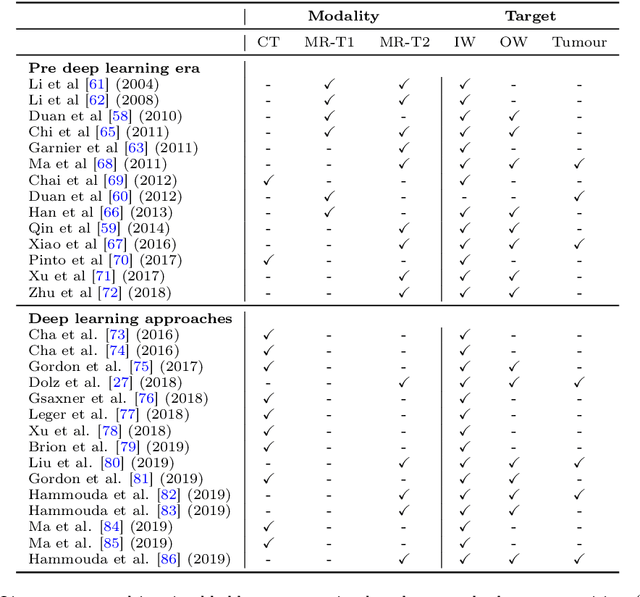
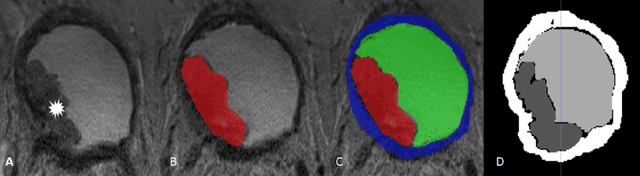
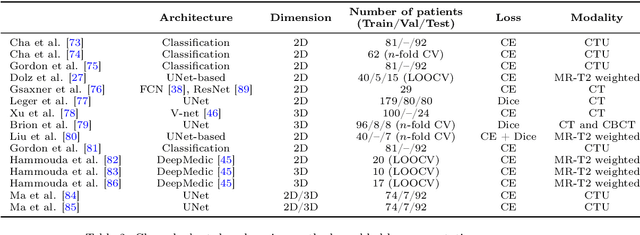
Abstract:Precise determination and assessment of bladder cancer (BC) extent of muscle invasion involvement guides proper risk stratification and personalized therapy selection. In this context, segmentation of both bladder walls and cancer are of pivotal importance, as it provides invaluable information to stage the primary tumour. Hence, multi region segmentation on patients presenting with symptoms of bladder tumours using deep learning heralds a new level of staging accuracy and prediction of the biologic behaviour of the tumour. Nevertheless, despite the success of these models in other medical problems, progress in multi region bladder segmentation is still at a nascent stage, with just a handful of works tackling a multi region scenario. Furthermore, most existing approaches systematically follow prior literature in other clinical problems, without casting a doubt on the validity of these methods on bladder segmentation, which may present different challenges. Inspired by this, we provide an in-depth look at bladder cancer segmentation using deep learning models. The critical determinants for accurate differentiation of muscle invasive disease, current status of deep learning based bladder segmentation, lessons and limitations of prior work are highlighted.
Teach me to segment with mixed supervision: Confident students become masters
Dec 15, 2020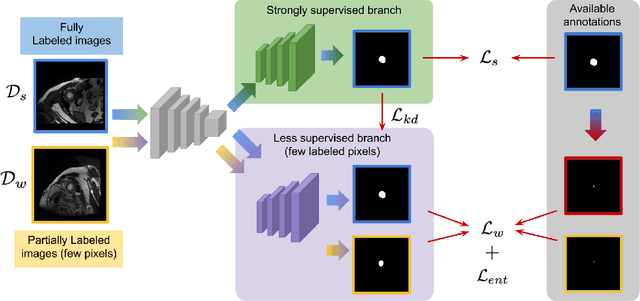
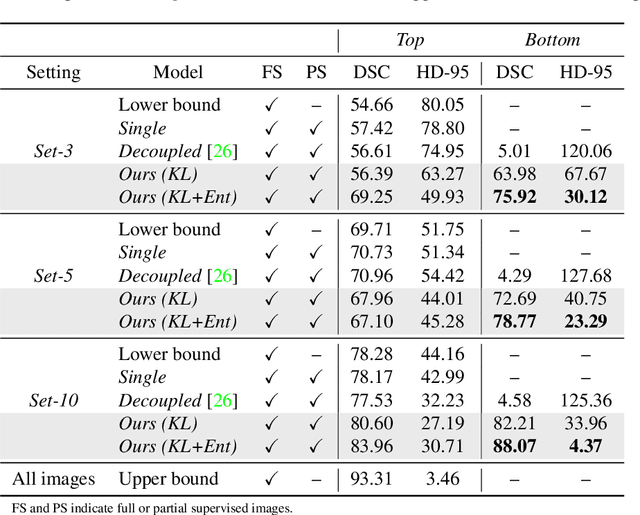
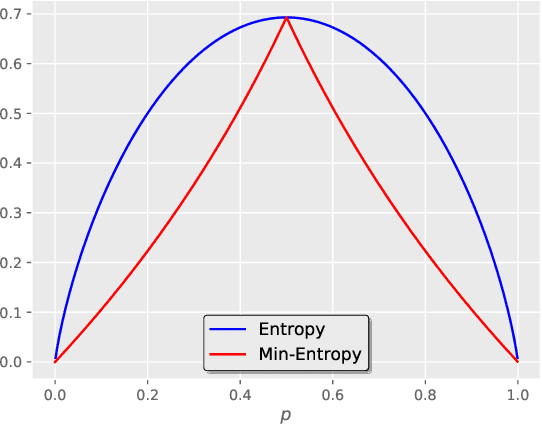
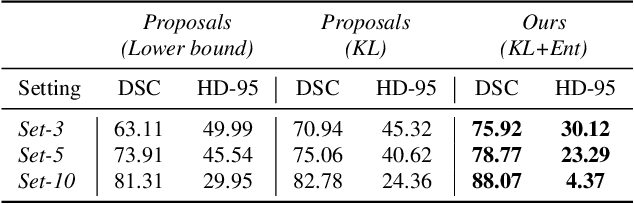
Abstract:Deep segmentation neural networks require large training datasets with pixel-wise segmentations, which are expensive to obtain in practice. Mixed supervision could mitigate this difficulty, with a small fraction of the data containing complete pixel-wise annotations, while the rest being less supervised, e.g., only a handful of pixels are labeled. In this work, we propose a dual-branch architecture, where the upper branch (teacher) receives strong annotations, while the bottom one (student) is driven by limited supervision and guided by the upper branch. In conjunction with a standard cross-entropy over the labeled pixels, our novel formulation integrates two important terms: (i) a Shannon entropy loss defined over the less-supervised images, which encourages confident student predictions at the bottom branch; and (ii) a Kullback-Leibler (KL) divergence, which transfers the knowledge from the predictions generated by the strongly supervised branch to the less-supervised branch, and guides the entropy (student-confidence) term to avoid trivial solutions. Very interestingly, we show that the synergy between the entropy and KL divergence yields substantial improvements in performances. Furthermore, we discuss an interesting link between Shannon-entropy minimization and standard pseudo-mask generation and argue that the former should be preferred over the latter for leveraging information from unlabeled pixels. Through a series of quantitative and qualitative experiments, we show the effectiveness of the proposed formulation in segmenting the left-ventricle endocardium in MRI images. We demonstrate that our method significantly outperforms other strategies to tackle semantic segmentation within a mixed-supervision framework. More interestingly, and in line with recent observations in classification, we show that the branch trained with reduced supervision largely outperforms the teacher.
Few-Shot Segmentation Without Meta-Learning: A Good Transductive Inference Is All You Need?
Dec 11, 2020
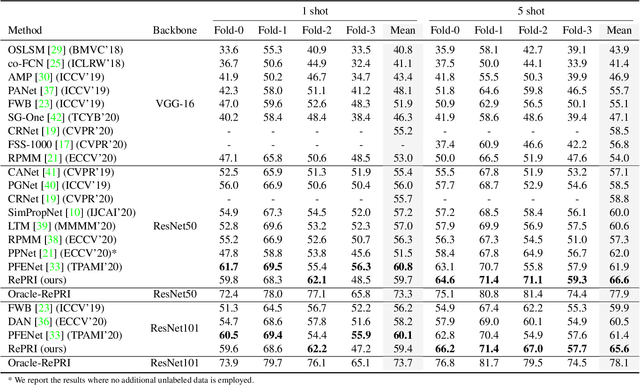
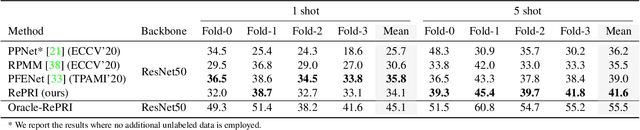
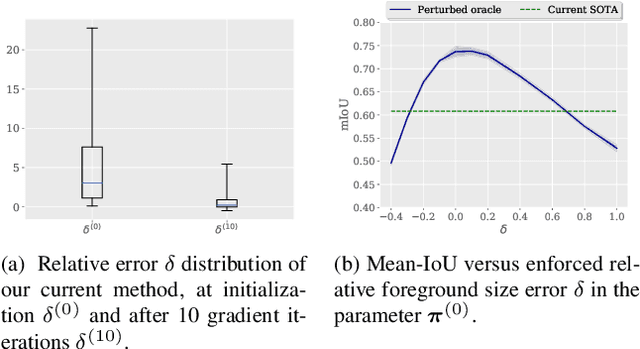
Abstract:Few-shot segmentation has recently attracted substantial interest, with the popular meta-learning paradigm widely dominating the literature. We show that the way inference is performed for a given few-shot segmentation task has a substantial effect on performances, an aspect that has been overlooked in the literature. We introduce a transductive inference, which leverages the statistics of the unlabeled pixels of a task by optimizing a new loss containing three complementary terms: (i) a standard cross-entropy on the labeled pixels; (ii) the entropy of posteriors on the unlabeled query pixels; and (iii) a global KL-divergence regularizer based on the proportion of the predicted foreground region. Our inference uses a simple linear classifier of the extracted features, has a computational load comparable to inductive inference and can be used on top of any base training. Using standard cross-entropy training on the base classes, our inference yields highly competitive performances on well-known few-shot segmentation benchmarks. On PASCAL-5i, it brings about 5% improvement over the best performing state-of-the-art method in the 5-shot scenario, while being on par in the 1-shot setting. Even more surprisingly, this gap widens as the number of support samples increases, reaching up to 6% in the 10-shot scenario. Furthermore, we introduce a more realistic setting with domain shift, where the base and novel classes are drawn from different datasets. In this setting, we found that our method achieves the best performances.
Privacy Preserving for Medical Image Analysis via Non-Linear Deformation Proxy
Nov 26, 2020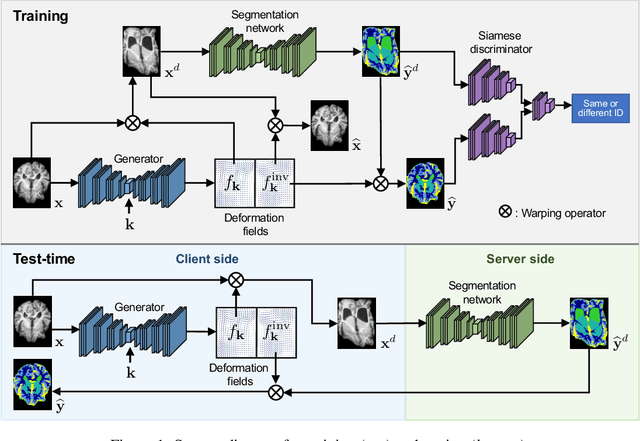
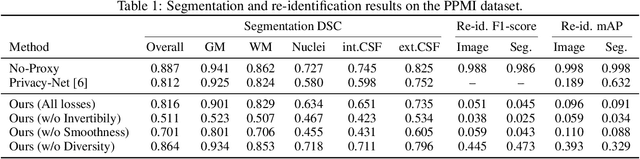
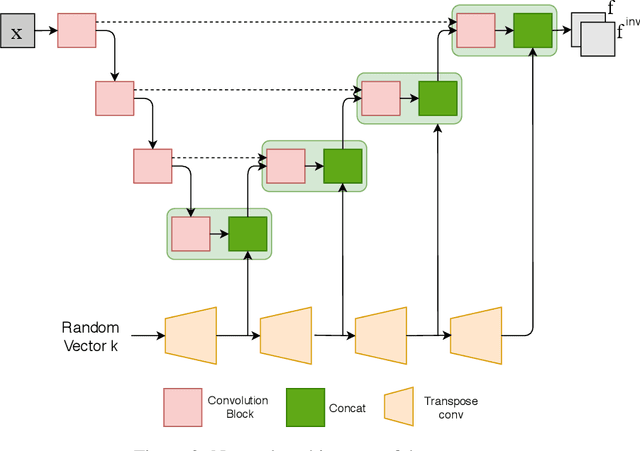
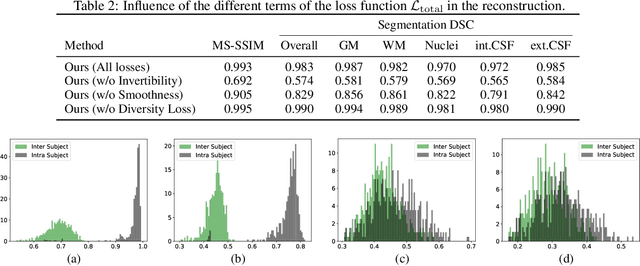
Abstract:We propose a client-server system which allows for the analysis of multi-centric medical images while preserving patient identity. In our approach, the client protects the patient identity by applying a pseudo-random non-linear deformation to the input image. This results into a proxy image which is sent to the server for processing. The server then returns back the deformed processed image which the client reverts to a canonical form. Our system has three components: 1) a flow-field generator which produces a pseudo-random deformation function, 2) a Siamese discriminator that learns the patient identity from the processed image, 3) a medical image processing network that analyzes the content of the proxy images. The system is trained end-to-end in an adversarial manner. By fooling the discriminator, the flow-field generator learns to produce a bi-directional non-linear deformation which allows to remove and recover the identity of the subject from both the input image and output result. After end-to-end training, the flow-field generator is deployed on the client side and the segmentation network is deployed on the server side. The proposed method is validated on the task of MRI brain segmentation using images from two different datasets. Results show that the segmentation accuracy of our method is similar to a system trained on non-encoded images, while considerably reducing the ability to recover subject identity.
Deep Interpretable Classification and Weakly-Supervised Segmentation of Histology Images via Max-Min Uncertainty
Nov 14, 2020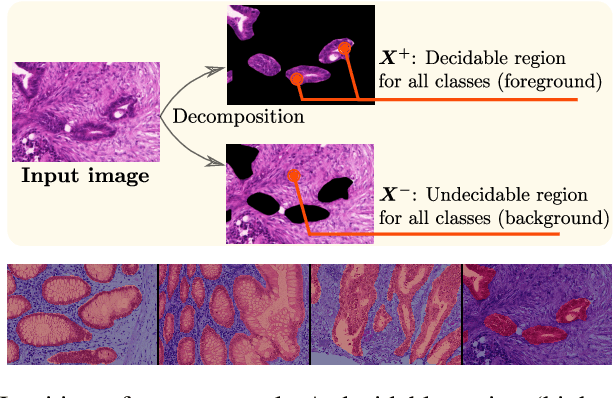
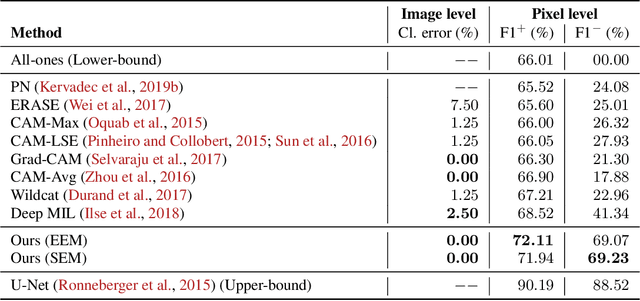
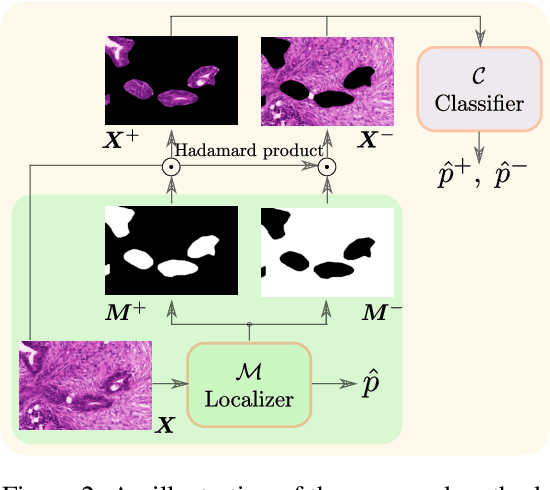
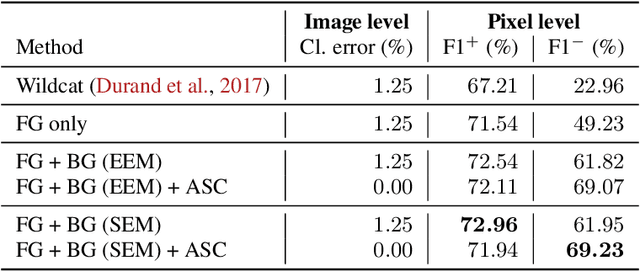
Abstract:Weakly supervised learning (WSL) has recently triggered substantial interest as it mitigates the lack of pixel-wise annotations, while enabling interpretable models. Given global image labels, WSL methods yield pixel-level predictions (segmentations). Despite their recent success, mostly with natural images, such methods could be seriously challenged when the foreground and background regions have similar visual cues, yielding high false-positive rates in segmentations, as is the case of challenging histology images. WSL training is commonly driven by standard classification losses, which implicitly maximize model confidence and find the discriminative regions linked to classification decisions. Therefore, they lack mechanisms for modeling explicitly non-discriminative regions and reducing false-positive rates. We propose new regularization terms, which enable the model to seek both non-discriminative and discriminative regions, while discouraging unbalanced segmentations. We introduce high uncertainty as a criterion to localize non-discriminative regions that do not affect classifier decision, and describe it with original Kullback-Leibler (KL) divergence losses evaluating the deviation of posterior predictions from the uniform distribution. Our KL terms encourage high uncertainty of the model when the latter takes the latent non-discriminative regions as input. Our loss integrates: (i) a cross-entropy seeking a foreground, where model confidence about class prediction is high; (ii) a KL regularizer seeking a background, where model uncertainty is high; and (iii) log-barrier terms discouraging unbalanced segmentations. Comprehensive experiments and ablation studies over the public GlaS colon cancer data show substantial improvements over state-of-the-art WSL methods, and confirm the effect of our new regularizers. Our code is publicly available.
Unsupervised Multi-Target Domain Adaptation Through Knowledge Distillation
Jul 20, 2020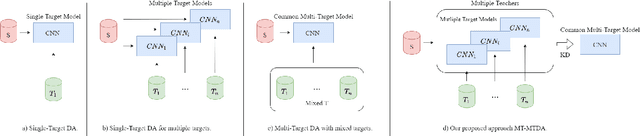
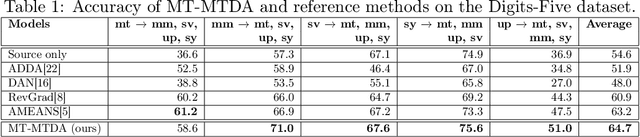
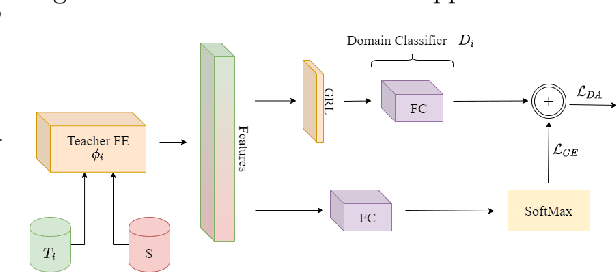
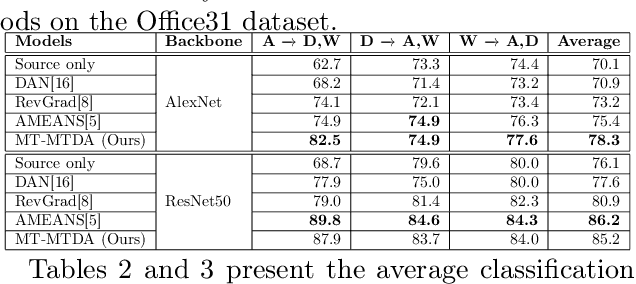
Abstract:Unsupervised domain adaptation (UDA) seeks to alleviate the problem of domain shift between the distribution of unlabeled data from the target domain w.r.t labeled data from source domain. While the single-target domain scenario is well studied in UDA literature, the Multi-Target Domain Adaptation (MTDA) setting remains largely unexplored despite its importance. For instance, in video surveillance, each camera can corresponds to a different viewpoint (target domain). MTDA problem can be addressed by adapting one specialized model per target domain, although this solution is too costly in many applications. It has also been addressed by blending target data for multi-domain adaptation to train a common model, yet this may lead to a reduction in performance. In this paper, we propose a new unsupervised MTDA approach to train a common CNN that can generalize across multiple target domains. Our approach the Multi-Teacher MTDA (MT-MTDA) relies on multi-teacher knowledge distillation (KD) in order to distill target domain knowledge from multiple teachers to a common student. Inspired by a common education scenario, a different target domain is assigned to each teacher model for UDA, and these teachers alternatively distill their knowledge to one common student model. The KD process is performed in a progressive manner, where the student is trained by each teacher on how to perform UDA, instead of directly learning domain adapted features. Finally, instead of directly combining the knowledge from each teacher, MT-MTDA alternates between teachers that distill knowledge in order to preserve the specificity of each target (teacher) when learning to adapt the student. MT-MTDA is compared against state-of-the-art methods on OfficeHome, Office31 and Digits-5 datasets, and empirical results show that our proposed model can provide a considerably higher level of accuracy across multiple target domains.
Laplacian Regularized Few-Shot Learning
Jun 28, 2020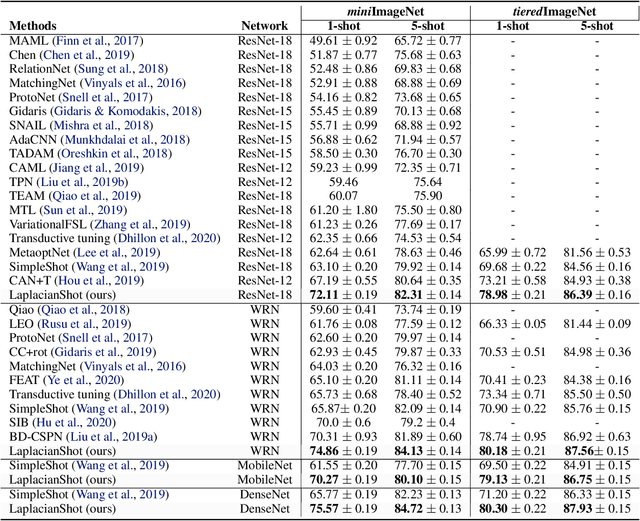
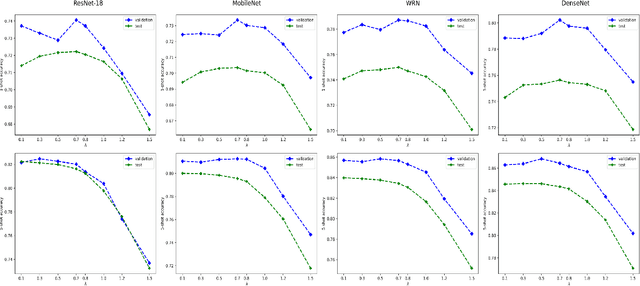
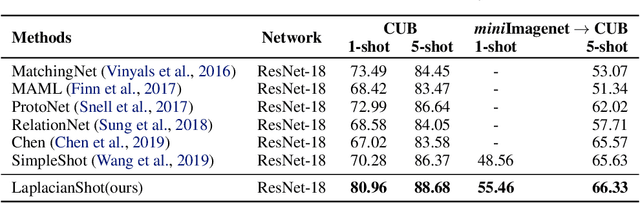
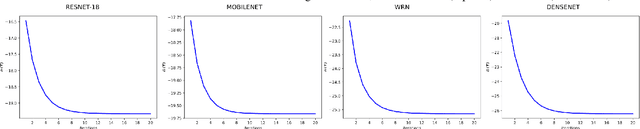
Abstract:We propose a transductive Laplacian-regularized inference for few-shot tasks. Given any feature embedding learned from the base classes, we minimize a quadratic binary-assignment function containing two terms: (1) a unary term assigning query samples to the nearest class prototype, and (2) a pairwise Laplacian term encouraging nearby query samples to have consistent label assignments. Our transductive inference does not re-train the base model, and can be viewed as a graph clustering of the query set, subject to supervision constraints from the support set. We derive a computationally efficient bound optimizer of a relaxation of our function, which computes independent (parallel) updates for each query sample, while guaranteeing convergence. Following a simple cross-entropy training on the base classes, and without complex meta-learning strategies, we conducted comprehensive experiments over five few-shot learning benchmarks. Our LaplacianShot consistently outperforms state-of-the-art methods by significant margins across different models, settings, and data sets. Furthermore, our transductive inference is very fast, with computational times that are close to inductive inference, and can be used for large-scale few-shot tasks.
Joint Progressive Knowledge Distillation and Unsupervised Domain Adaptation
May 16, 2020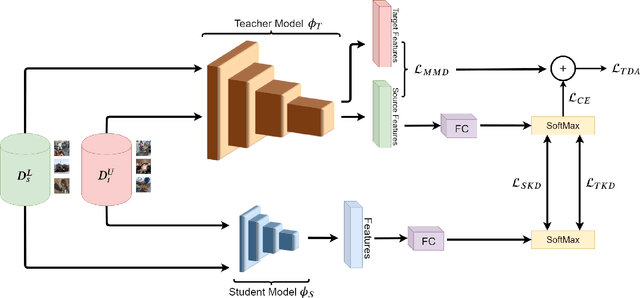

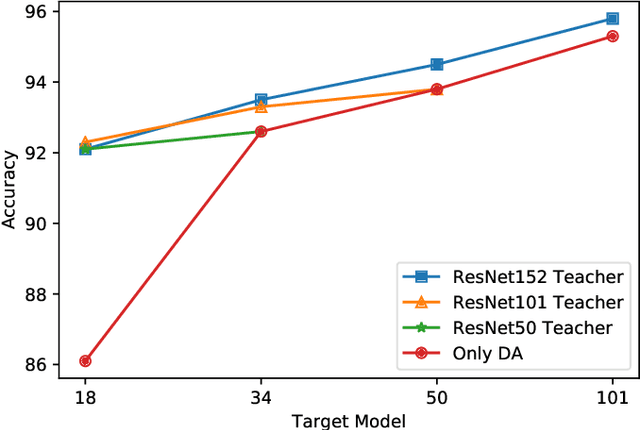
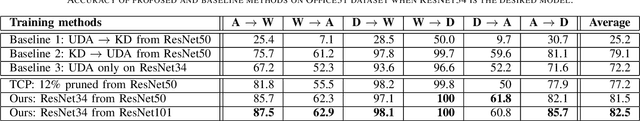
Abstract:Currently, the divergence in distributions of design and operational data, and large computational complexity are limiting factors in the adoption of CNNs in real-world applications. For instance, person re-identification systems typically rely on a distributed set of cameras, where each camera has different capture conditions. This can translate to a considerable shift between source (e.g. lab setting) and target (e.g. operational camera) domains. Given the cost of annotating image data captured for fine-tuning in each target domain, unsupervised domain adaptation (UDA) has become a popular approach to adapt CNNs. Moreover, state-of-the-art deep learning models that provide a high level of accuracy often rely on architectures that are too complex for real-time applications. Although several compression and UDA approaches have recently been proposed to overcome these limitations, they do not allow optimizing a CNN to simultaneously address both. In this paper, we propose an unexplored direction -- the joint optimization of CNNs to provide a compressed model that is adapted to perform well for a given target domain. In particular, the proposed approach performs unsupervised knowledge distillation (KD) from a complex teacher model to a compact student model, by leveraging both source and target data. It also improves upon existing UDA techniques by progressively teaching the student about domain-invariant features, instead of directly adapting a compact model on target domain data. Our method is compared against state-of-the-art compression and UDA techniques, using two popular classification datasets for UDA -- Office31 and ImageClef-DA. In both datasets, results indicate that our method can achieve the highest level of accuracy while requiring a comparable or lower time complexity.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge