Jiang Shaohua
Bottleneck Tokens for Unified Multimodal Retrieval
Apr 13, 2026Abstract:Adapting decoder-only multimodal large language models (MLLMs) for unified multimodal retrieval faces two structural gaps. First, existing methods rely on implicit pooling, which overloads the hidden state of a standard vocabulary token (e.g., <EOS>) as the sequence-level representation, a mechanism never designed for information aggregation. Second, contrastive fine-tuning specifies what the embedding should match but provides no token-level guidance on how information should be compressed into it. We address both gaps with two complementary components. Architecturally, we introduce Bottleneck Tokens (BToks), a small set of learnable tokens that serve as a fixed-capacity explicit pooling mechanism. For training, we propose Generative Information Condensation: a next-token prediction objective coupled with a Condensation Mask that severs the direct attention path from target tokens to query tokens. All predictive signals are thereby forced through the BToks, converting the generative loss into dense, token-level supervision for semantic compression. At inference time, only the input and BToks are processed in a single forward pass with negligible overhead over conventional last-token pooling. On MMEB-V2 (78 datasets, 3 modalities, 9 meta-tasks), our approach achieves state-of-the-art among 2B-scale methods under comparable data conditions, attaining an Overall score of 59.0 (+3.6 over VLM2Vec-V2) with substantial gains on semantically demanding tasks (e.g., +12.6 on Video-QA).
GDGRU-DTA: Predicting Drug-Target Binding Affinity Based on GNN and Double GRU
Apr 25, 2022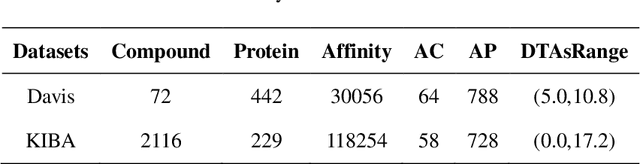
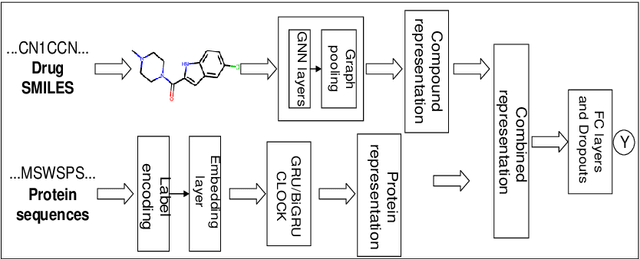
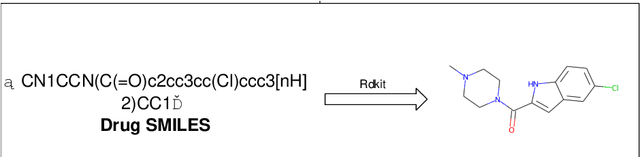
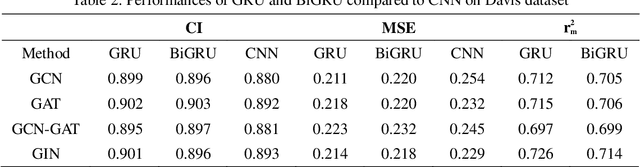
Abstract:The work for predicting drug and target affinity(DTA) is crucial for drug development and repurposing. In this work, we propose a novel method called GDGRU-DTA to predict the binding affinity between drugs and targets, which is based on GraphDTA, but we consider that protein sequences are long sequences, so simple CNN cannot capture the context dependencies in protein sequences well. Therefore, we improve it by interpreting the protein sequences as time series and extracting their features using Gate Recurrent Unit(GRU) and Bidirectional Gate Recurrent Unit(BiGRU). For the drug, our processing method is similar to that of GraphDTA, but uses two different graph convolution methods. Subsequently, the representation of drugs and proteins are concatenated for final prediction. We evaluate the proposed model on two benchmark datasets. Our model outperforms some state-of-the-art deep learning methods, and the results demonstrate the feasibility and excellent feature capture ability of our model.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge