Jan Kybic
Czech Technical University in Prague, Czech Republic
Biomedical image analysis competitions: The state of current participation practice
Dec 16, 2022Abstract:The number of international benchmarking competitions is steadily increasing in various fields of machine learning (ML) research and practice. So far, however, little is known about the common practice as well as bottlenecks faced by the community in tackling the research questions posed. To shed light on the status quo of algorithm development in the specific field of biomedical imaging analysis, we designed an international survey that was issued to all participants of challenges conducted in conjunction with the IEEE ISBI 2021 and MICCAI 2021 conferences (80 competitions in total). The survey covered participants' expertise and working environments, their chosen strategies, as well as algorithm characteristics. A median of 72% challenge participants took part in the survey. According to our results, knowledge exchange was the primary incentive (70%) for participation, while the reception of prize money played only a minor role (16%). While a median of 80 working hours was spent on method development, a large portion of participants stated that they did not have enough time for method development (32%). 25% perceived the infrastructure to be a bottleneck. Overall, 94% of all solutions were deep learning-based. Of these, 84% were based on standard architectures. 43% of the respondents reported that the data samples (e.g., images) were too large to be processed at once. This was most commonly addressed by patch-based training (69%), downsampling (37%), and solving 3D analysis tasks as a series of 2D tasks. K-fold cross-validation on the training set was performed by only 37% of the participants and only 50% of the participants performed ensembling based on multiple identical models (61%) or heterogeneous models (39%). 48% of the respondents applied postprocessing steps.
Learning to segment from object sizes
Jul 01, 2022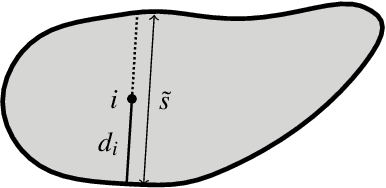
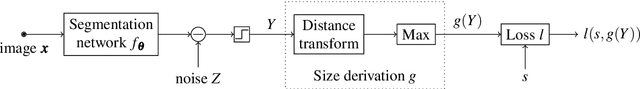
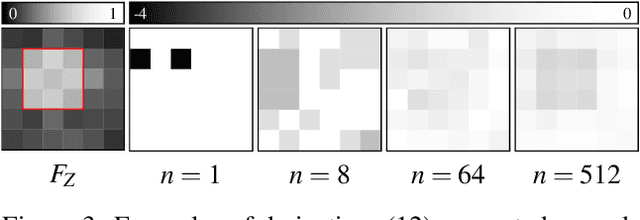

Abstract:Deep learning has proved particularly useful for semantic segmentation, a fundamental image analysis task. However, the standard deep learning methods need many training images with ground-truth pixel-wise annotations, which are usually laborious to obtain and, in some cases (e.g., medical images), require domain expertise. Therefore, instead of pixel-wise annotations, we focus on image annotations that are significantly easier to acquire but still informative, namely the size of foreground objects. We define the object size as the maximum distance between a foreground pixel and the background. We propose an algorithm for training a deep segmentation network from a dataset of a few pixel-wise annotated images and many images with known object sizes. The algorithm minimizes a discrete (non-differentiable) loss function defined over the object sizes by sampling the gradient and then using the standard back-propagation algorithm. We study the performance of our approach in terms of training time and generalization error.
Fast learning from label proportions with small bags
Oct 08, 2021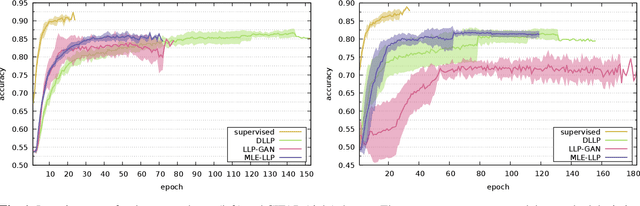
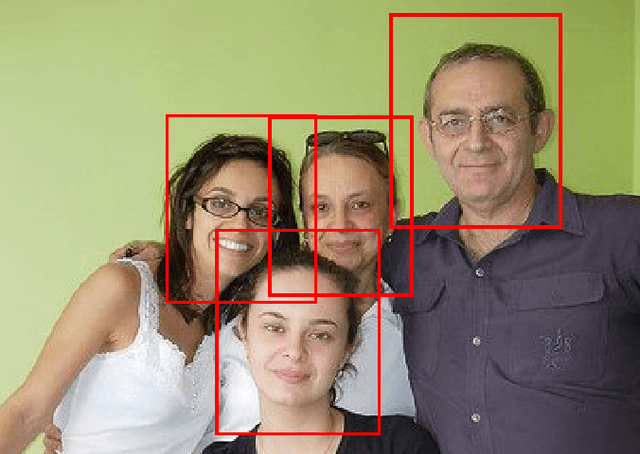
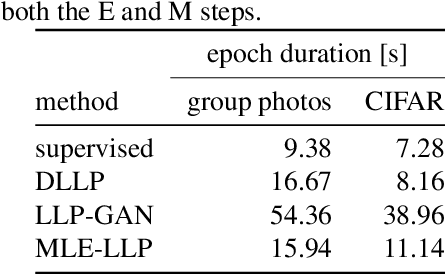

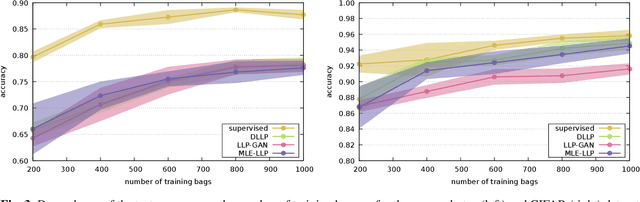
Abstract:In learning from label proportions (LLP), the instances are grouped into bags, and the task is to learn an instance classifier given relative class proportions in training bags. LLP is useful when obtaining individual instance labels is impossible or costly. In this work, we focus on the case of small bags, which allows designing more efficient algorithms by explicitly considering all consistent label combinations. In particular, we propose an EM algorithm alternating between optimizing a general neural network instance classifier and incorporating bag-level annotations. In comparison to existing deep LLP methods, our approach converges faster to a comparable or better solution. Several experiments were performed on two different datasets.
Automatic evaluation of human oocyte developmental potential from microscopy images
Feb 27, 2021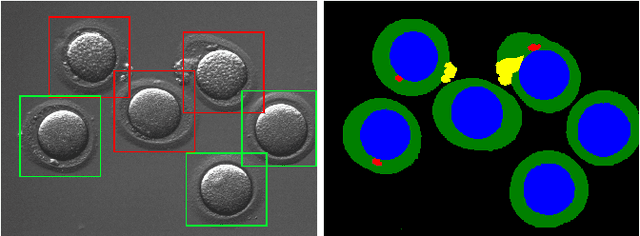
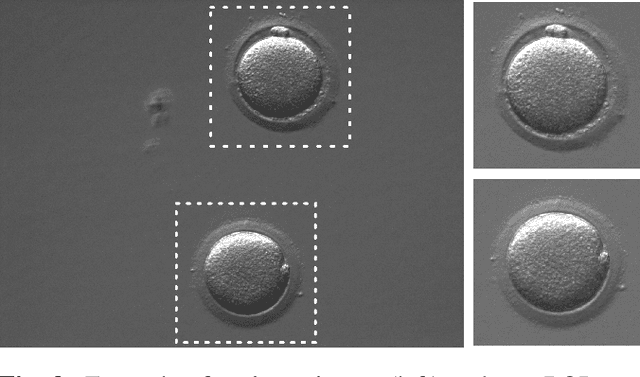
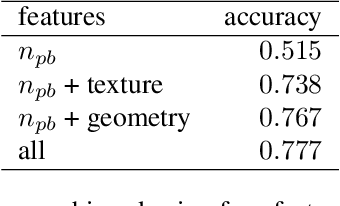
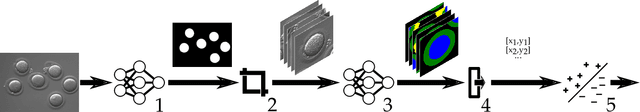

Abstract:Infertility is becoming an issue for an increasing number of couples. The most common solution, in vitro fertilization, requires embryologists to carefully examine light microscopy images of human oocytes to determine their developmental potential. We propose an automatic system to improve the speed, repeatability, and accuracy of this process. We first localize individual oocytes and identify their principal components using CNN (U-Net) segmentation. We calculate several descriptors based on geometry and texture. The final step is an SVM classifier. Both the segmentation and classification training are based on expert annotations. The presented approach leads to the classification accuracy of 70%.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge