Jakub Nalepa
KP Labs, Silesian University of Technology
Squeezing nnU-Nets with Knowledge Distillation for On-Board Cloud Detection
Jun 16, 2023



Abstract:Cloud detection is a pivotal satellite image pre-processing step that can be performed both on the ground and on board a satellite to tag useful images. In the latter case, it can reduce the amount of data to downlink by pruning the cloudy areas, or to make a satellite more autonomous through data-driven acquisition re-scheduling. We approach this task with nnU-Nets, a self-reconfigurable framework able to perform meta-learning of a segmentation network over various datasets. Unfortunately, such models are commonly memory-inefficient due to their (very) large architectures. To benefit from them in on-board processing, we compress nnU-Nets with knowledge distillation into much smaller and compact U-Nets. Our experiments, performed over Sentinel-2 and Landsat-8 images revealed that nnU-Nets deliver state-of-the-art performance without any manual design. Our approach was ranked within the top 7% best solutions (across 847 teams) in the On Cloud N: Cloud Cover Detection Challenge, where we reached the Jaccard index of 0.882 over more than 10k unseen Sentinel-2 images (the winners obtained 0.897, the baseline U-Net with the ResNet-34 backbone: 0.817, and the classic Sentinel-2 image thresholding: 0.652). Finally, we showed that knowledge distillation enables to elaborate dramatically smaller (almost 280x) U-Nets when compared to nnU-Nets while still maintaining their segmentation capabilities.
Detecting Clouds in Multispectral Satellite Images Using Quantum-Kernel Support Vector Machines
Feb 16, 2023Abstract:Support vector machines (SVMs) are a well-established classifier effectively deployed in an array of classification tasks. In this work, we consider extending classical SVMs with quantum kernels and applying them to satellite data analysis. The design and implementation of SVMs with quantum kernels (hybrid SVMs) are presented. Here, the pixels are mapped to the Hilbert space using a family of parameterized quantum feature maps (related to quantum kernels). The parameters are optimized to maximize the kernel target alignment. The quantum kernels have been selected such that they enabled analysis of numerous relevant properties while being able to simulate them with classical computers on a real-life large-scale dataset. Specifically, we approach the problem of cloud detection in the multispectral satellite imagery, which is one of the pivotal steps in both on-the-ground and on-board satellite image analysis processing chains. The experiments performed over the benchmark Landsat-8 multispectral dataset revealed that the simulated hybrid SVM successfully classifies satellite images with accuracy comparable to the classical SVM with the RBF kernel for large datasets. Interestingly, for large datasets, the high accuracy was also observed for the simple quantum kernels, lacking quantum entanglement.
Multitemporal and multispectral data fusion for super-resolution of Sentinel-2 images
Jan 26, 2023



Abstract:Multispectral Sentinel-2 images are a valuable source of Earth observation data, however spatial resolution of their spectral bands limited to 10 m, 20 m, and 60 m ground sampling distance remains insufficient in many cases. This problem can be addressed with super-resolution, aimed at reconstructing a high-resolution image from a low-resolution observation. For Sentinel-2, spectral information fusion allows for enhancing the 20 m and 60 m bands to the 10 m resolution. Also, there were attempts to combine multitemporal stacks of individual Sentinel-2 bands, however these two approaches have not been combined so far. In this paper, we introduce DeepSent -- a new deep network for super-resolving multitemporal series of multispectral Sentinel-2 images. It is underpinned with information fusion performed simultaneously in the spectral and temporal dimensions to generate an enlarged multispectral image. In our extensive experimental study, we demonstrate that our solution outperforms other state-of-the-art techniques that realize either multitemporal or multispectral data fusion. Furthermore, we show that the advantage of DeepSent results from how these two fusion types are combined in a single architecture, which is superior to performing such fusion in a sequential manner. Importantly, we have applied our method to super-resolve real-world Sentinel-2 images, enhancing the spatial resolution of all the spectral bands to 3.3 m nominal ground sampling distance, and we compare the outcome with very high-resolution WorldView-2 images. We will publish our implementation upon paper acceptance, and we expect it will increase the possibilities of exploiting super-resolved Sentinel-2 images in real-life applications.
Biomedical image analysis competitions: The state of current participation practice
Dec 16, 2022Abstract:The number of international benchmarking competitions is steadily increasing in various fields of machine learning (ML) research and practice. So far, however, little is known about the common practice as well as bottlenecks faced by the community in tackling the research questions posed. To shed light on the status quo of algorithm development in the specific field of biomedical imaging analysis, we designed an international survey that was issued to all participants of challenges conducted in conjunction with the IEEE ISBI 2021 and MICCAI 2021 conferences (80 competitions in total). The survey covered participants' expertise and working environments, their chosen strategies, as well as algorithm characteristics. A median of 72% challenge participants took part in the survey. According to our results, knowledge exchange was the primary incentive (70%) for participation, while the reception of prize money played only a minor role (16%). While a median of 80 working hours was spent on method development, a large portion of participants stated that they did not have enough time for method development (32%). 25% perceived the infrastructure to be a bottleneck. Overall, 94% of all solutions were deep learning-based. Of these, 84% were based on standard architectures. 43% of the respondents reported that the data samples (e.g., images) were too large to be processed at once. This was most commonly addressed by patch-based training (69%), downsampling (37%), and solving 3D analysis tasks as a series of 2D tasks. K-fold cross-validation on the training set was performed by only 37% of the participants and only 50% of the participants performed ensembling based on multiple identical models (61%) or heterogeneous models (39%). 48% of the respondents applied postprocessing steps.
Self-Configuring nnU-Nets Detect Clouds in Satellite Images
Oct 24, 2022



Abstract:Cloud detection is a pivotal satellite image pre-processing step that can be performed both on the ground and on board a satellite to tag useful images. In the latter case, it can help to reduce the amount of data to downlink by pruning the cloudy areas, or to make a satellite more autonomous through data-driven acquisition re-scheduling of the cloudy areas. We approach this important task with nnU-Nets, a self-reconfigurable framework able to perform meta-learning of a segmentation network over various datasets. Our experiments, performed over Sentinel-2 and Landsat-8 multispectral images revealed that nnU-Nets deliver state-of-the-art cloud segmentation performance without any manual design. Our approach was ranked within the top 7% best solutions (across 847 participating teams) in the On Cloud N: Cloud Cover Detection Challenge, where we reached the Jaccard index of 0.882 over more than 10k unseen Sentinel-2 image patches (the winners obtained 0.897, whereas the baseline U-Net with the ResNet-34 backbone used as an encoder: 0.817, and the classic Sentinel-2 image thresholding: 0.652).
MuS2: A Benchmark for Sentinel-2 Multi-Image Super-Resolution
Oct 06, 2022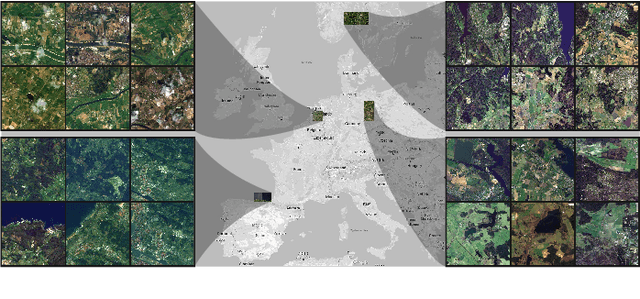
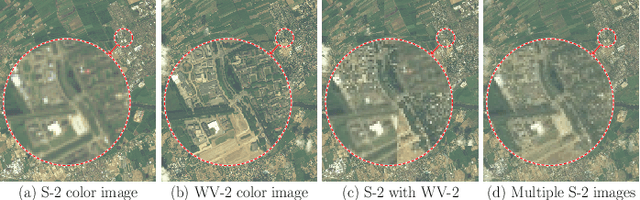
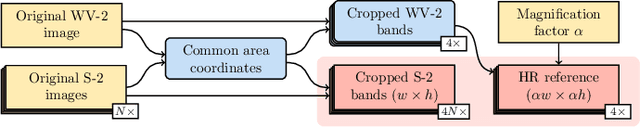


Abstract:Insufficient spatial resolution of satellite imagery, including Sentinel-2 data, is a serious limitation in many practical use cases. To mitigate this problem, super-resolution reconstruction is receiving considerable attention from the remote sensing community. When it is performed from multiple images captured at subsequent revisits, it may benefit from information fusion, leading to enhanced reconstruction accuracy. One of the obstacles in multi-image super-resolution consists in the scarcity of real-life benchmark datasets -- most of the research was performed for simulated data which do not fully reflect the operating conditions. In this letter, we introduce a new MuS2 benchmark for multi-image super-resolution reconstruction of Sentinel-2 images, with WorldView-2 imagery used as the high-resolution reference. Within MuS2, we publish the first end-to-end evaluation procedure for this problem which we expect to help the researchers in advancing the state of the art in multi-image super-resolution for Sentinel-2 imagery.
Deep learning automates bidimensional and volumetric tumor burden measurement from MRI in pre- and post-operative glioblastoma patients
Sep 03, 2022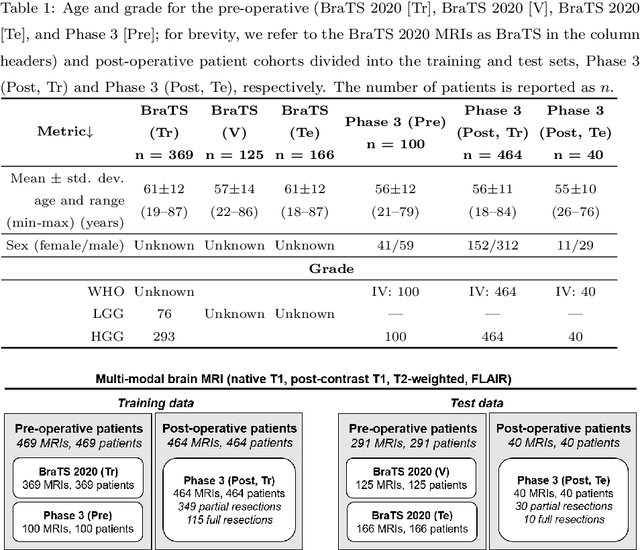
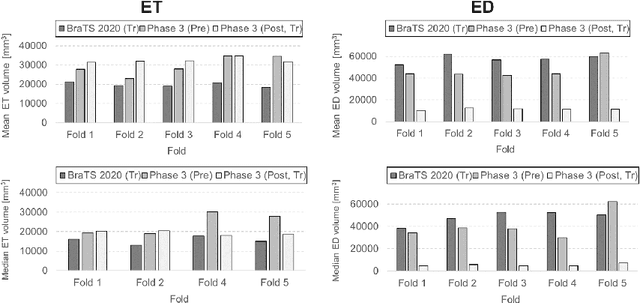
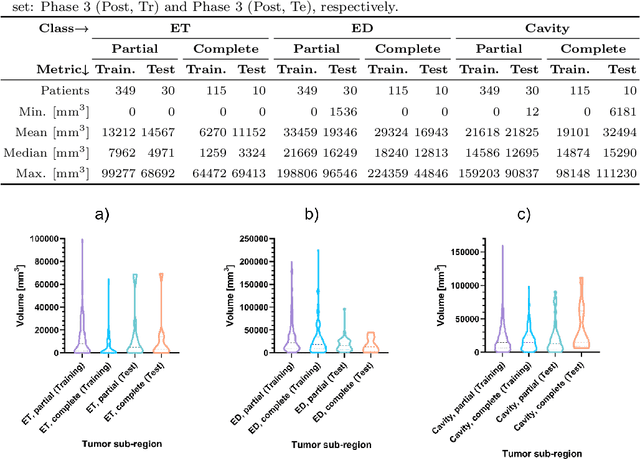
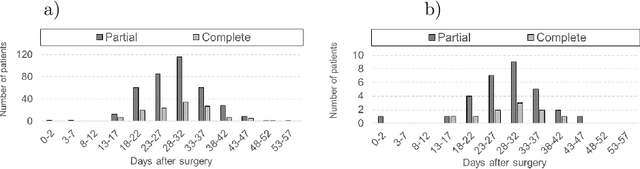
Abstract:Tumor burden assessment by magnetic resonance imaging (MRI) is central to the evaluation of treatment response for glioblastoma. This assessment is complex to perform and associated with high variability due to the high heterogeneity and complexity of the disease. In this work, we tackle this issue and propose a deep learning pipeline for the fully automated end-to-end analysis of glioblastoma patients. Our approach simultaneously identifies tumor sub-regions, including the enhancing tumor, peritumoral edema and surgical cavity in the first step, and then calculates the volumetric and bidimensional measurements that follow the current Response Assessment in Neuro-Oncology (RANO) criteria. Also, we introduce a rigorous manual annotation process which was followed to delineate the tumor sub-regions by the human experts, and to capture their segmentation confidences that are later used while training the deep learning models. The results of our extensive experimental study performed over 760 pre-operative and 504 post-operative adult patients with glioma obtained from the public database (acquired at 19 sites in years 2021-2020) and from a clinical treatment trial (47 and 69 sites for pre-/post-operative patients, 2009-2011) and backed up with thorough quantitative, qualitative and statistical analysis revealed that our pipeline performs accurate segmentation of pre- and post-operative MRIs in a fraction of the manual delineation time (up to 20 times faster than humans). The bidimensional and volumetric measurements were in strong agreement with expert radiologists, and we showed that RANO measurements are not always sufficient to quantify tumor burden.
A Multibranch Convolutional Neural Network for Hyperspectral Unmixing
Aug 03, 2022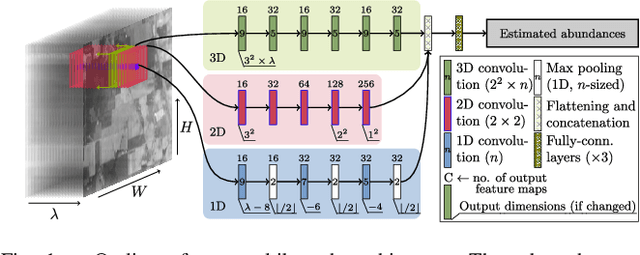
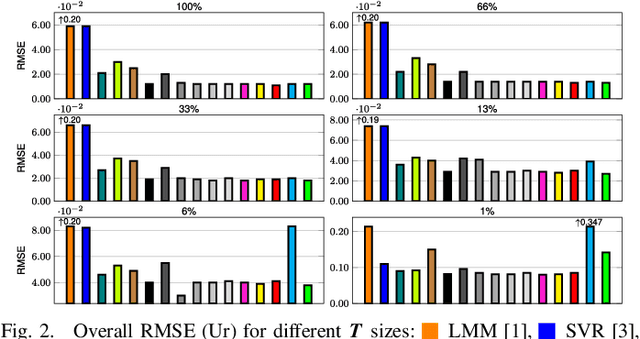
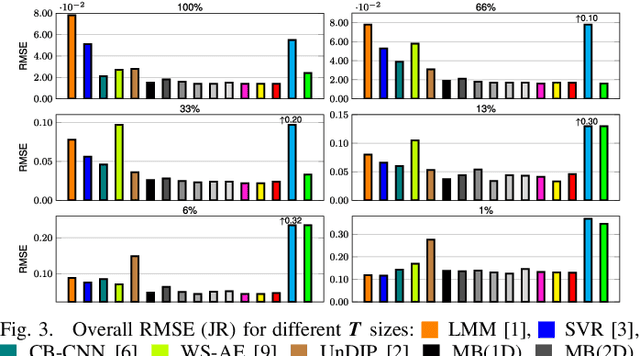
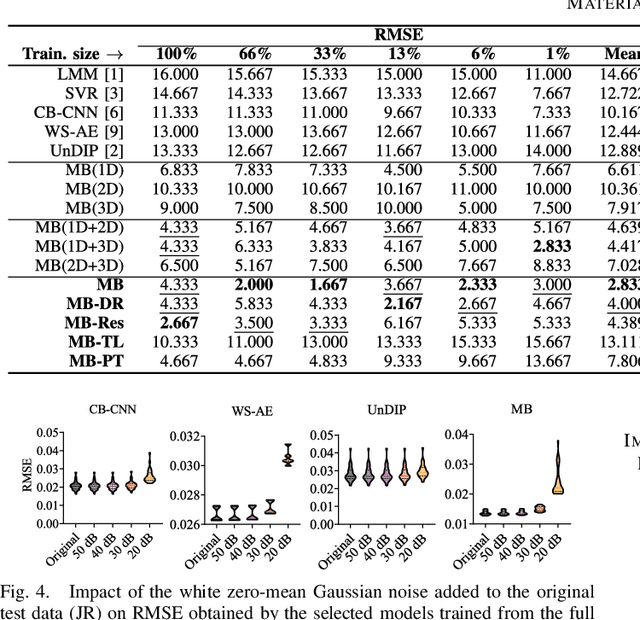
Abstract:Hyperspectral unmixing remains one of the most challenging tasks in the analysis of such data. Deep learning has been blooming in the field and proved to outperform other classic unmixing techniques, and can be effectively deployed onboard Earth observation satellites equipped with hyperspectral imagers. In this letter, we follow this research pathway and propose a multi-branch convolutional neural network that benefits from fusing spectral, spatial, and spectral-spatial features in the unmixing process. The results of our experiments, backed up with the ablation study, revealed that our techniques outperform others from the literature and lead to higher-quality fractional abundance estimation. Also, we investigated the influence of reducing the training sets on the capabilities of all algorithms and their robustness against noise, as capturing large and representative ground-truth sets is time-consuming and costly in practice, especially in emerging Earth observation scenarios.
* 14 pages (including supplementary material), published in IEEE Geoscience and Remote Sensing Letters
Graph Neural Networks Extract High-Resolution Cultivated Land Maps from Sentinel-2 Image Series
Aug 03, 2022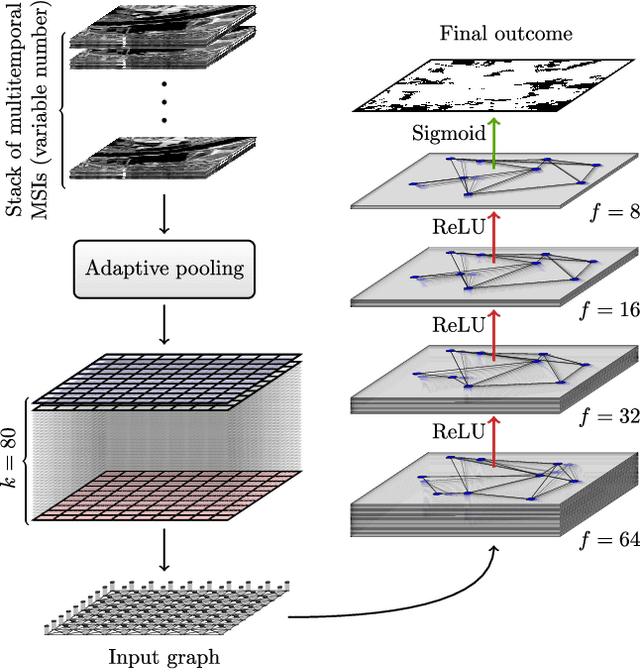

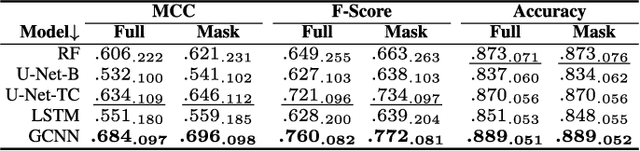
Abstract:Maintaining farm sustainability through optimizing the agricultural management practices helps build more planet-friendly environment. The emerging satellite missions can acquire multi- and hyperspectral imagery which captures more detailed spectral information concerning the scanned area, hence allows us to benefit from subtle spectral features during the analysis process in agricultural applications. We introduce an approach for extracting 2.5 m cultivated land maps from 10 m Sentinel-2 multispectral image series which benefits from a compact graph convolutional neural network. The experiments indicate that our models not only outperform classical and deep machine learning techniques through delivering higher-quality segmentation maps, but also dramatically reduce the memory footprint when compared to U-Nets (almost 8k trainable parameters of our models, with up to 31M parameters of U-Nets). Such memory frugality is pivotal in the missions which allow us to uplink a model to the AI-powered satellite once it is in orbit, as sending large nets is impossible due to the time constraints.
* 7 pages (including supplementary material), published in IEEE Geoscience and Remote Sensing Letters
Segmenting Hyperspectral Images Using Spectral-Spatial Convolutional Neural Networks With Training-Time Data Augmentation
Jul 27, 2019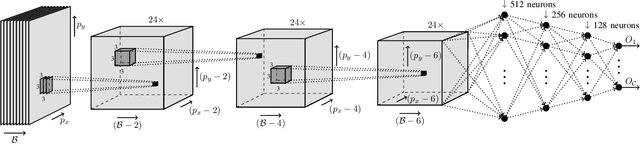
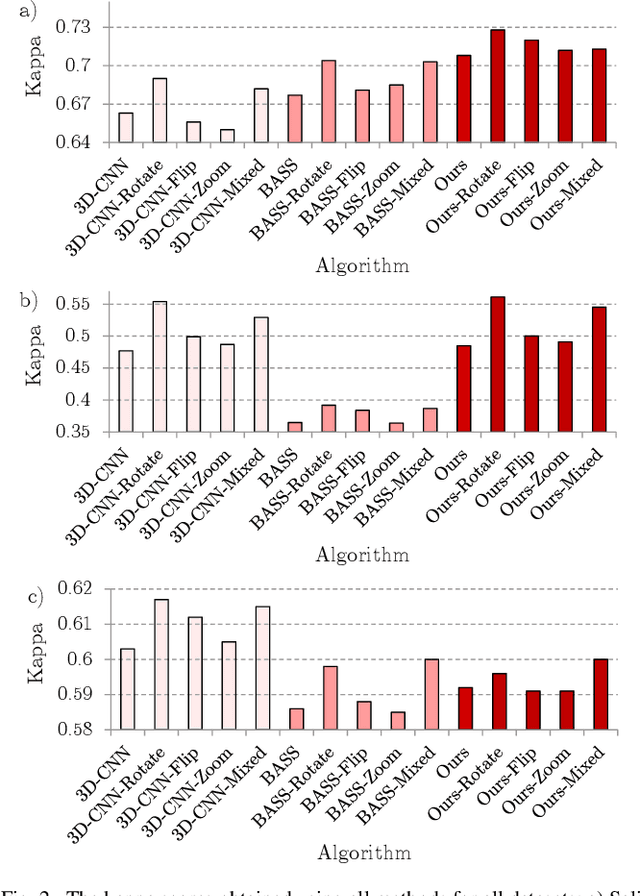
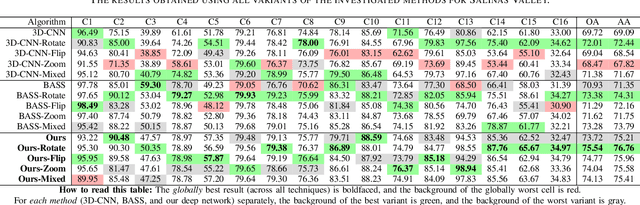
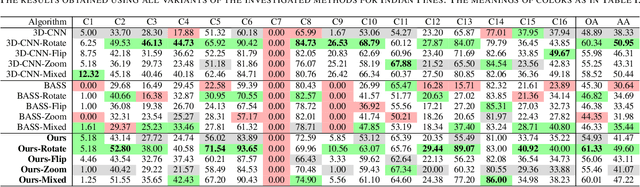
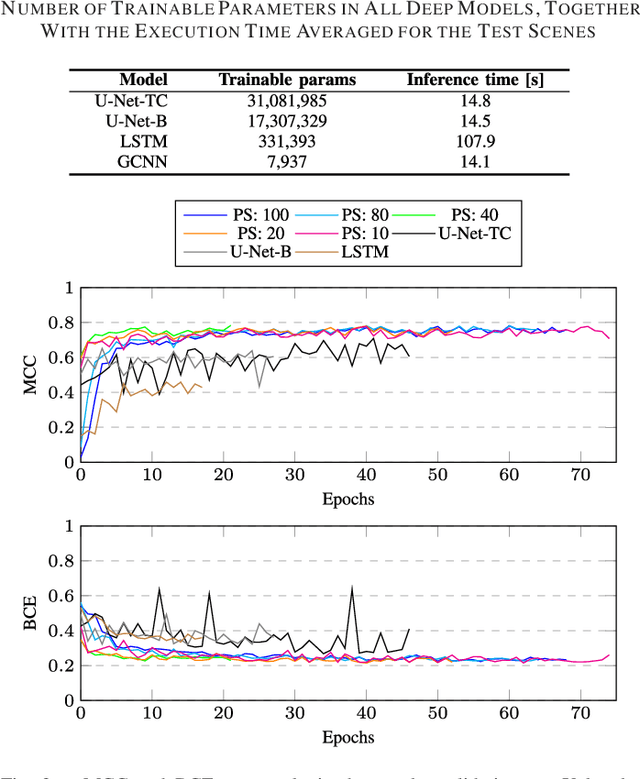
Abstract:Hyperspectral imaging provides detailed information about the scanned objects, as it captures their spectral characteristics within a large number of wavelength bands. Classification of such data has become an active research topic due to its wide applicability in a variety of fields. Deep learning has established the state of the art in the area, and it constitutes the current research mainstream. In this letter, we introduce a new spectral-spatial convolutional neural network, benefitting from a battery of data augmentation techniques which help deal with a real-life problem of lacking ground-truth training data. Our rigorous experiments showed that the proposed method outperforms other spectral-spatial techniques from the literature, and delivers precise hyperspectral classification in real time.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge