Iain Murray
Markov Chain Truncation for Doubly-Intractable Inference
Mar 11, 2017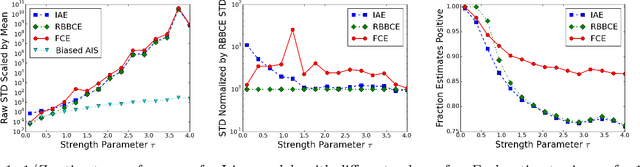
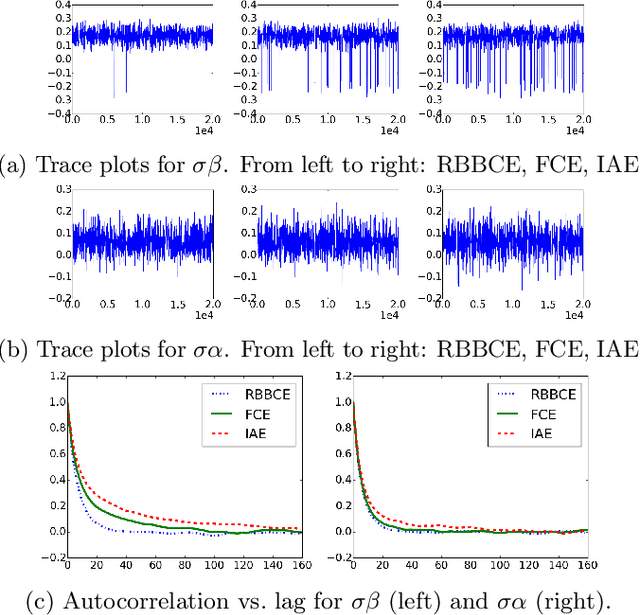
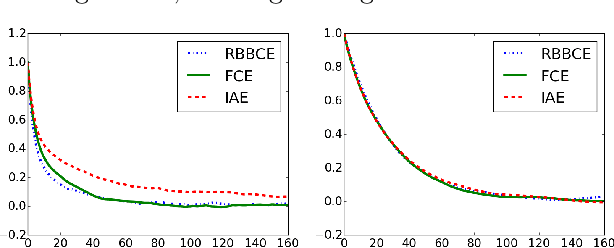
Abstract:Computing partition functions, the normalizing constants of probability distributions, is often hard. Variants of importance sampling give unbiased estimates of a normalizer Z, however, unbiased estimates of the reciprocal 1/Z are harder to obtain. Unbiased estimates of 1/Z allow Markov chain Monte Carlo sampling of "doubly-intractable" distributions, such as the parameter posterior for Markov Random Fields or Exponential Random Graphs. We demonstrate how to construct unbiased estimates for 1/Z given access to black-box importance sampling estimators for Z. We adapt recent work on random series truncation and Markov chain coupling, producing estimators with lower variance and a higher percentage of positive estimates than before. Our debiasing algorithms are simple to implement, and have some theoretical and empirical advantages over existing methods.
Neural Autoregressive Distribution Estimation
May 27, 2016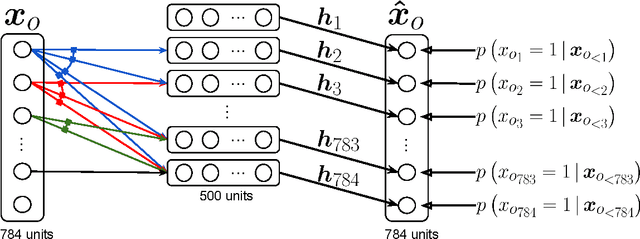
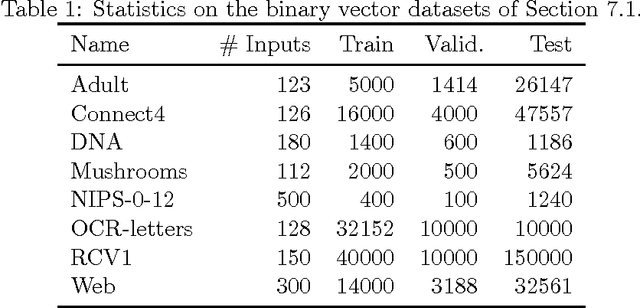
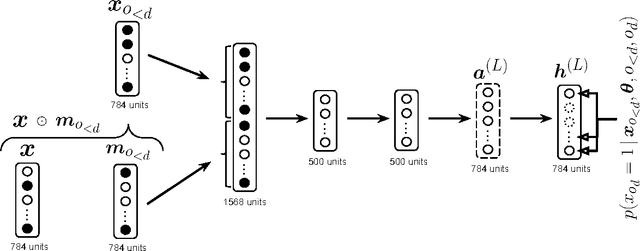
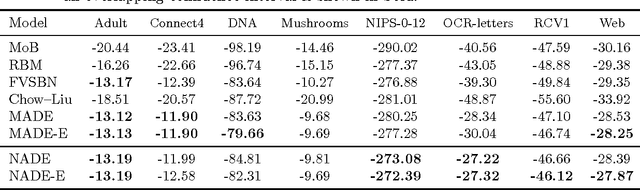
Abstract:We present Neural Autoregressive Distribution Estimation (NADE) models, which are neural network architectures applied to the problem of unsupervised distribution and density estimation. They leverage the probability product rule and a weight sharing scheme inspired from restricted Boltzmann machines, to yield an estimator that is both tractable and has good generalization performance. We discuss how they achieve competitive performance in modeling both binary and real-valued observations. We also present how deep NADE models can be trained to be agnostic to the ordering of input dimensions used by the autoregressive product rule decomposition. Finally, we also show how to exploit the topological structure of pixels in images using a deep convolutional architecture for NADE.
MADE: Masked Autoencoder for Distribution Estimation
Jun 05, 2015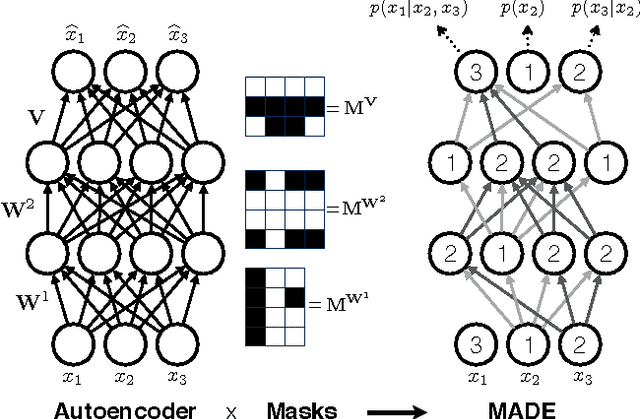
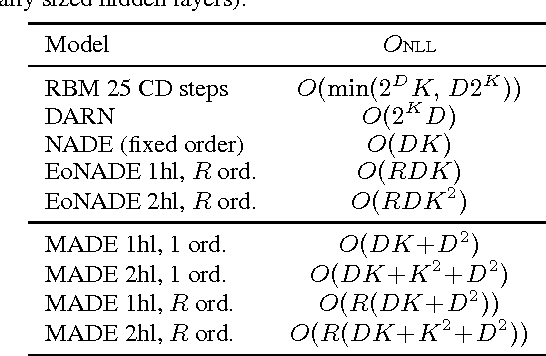
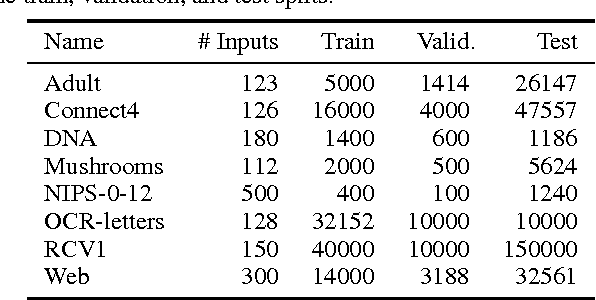
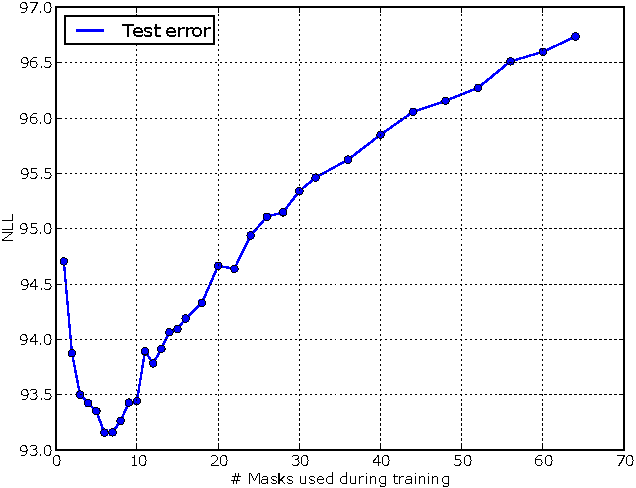
Abstract:There has been a lot of recent interest in designing neural network models to estimate a distribution from a set of examples. We introduce a simple modification for autoencoder neural networks that yields powerful generative models. Our method masks the autoencoder's parameters to respect autoregressive constraints: each input is reconstructed only from previous inputs in a given ordering. Constrained this way, the autoencoder outputs can be interpreted as a set of conditional probabilities, and their product, the full joint probability. We can also train a single network that can decompose the joint probability in multiple different orderings. Our simple framework can be applied to multiple architectures, including deep ones. Vectorized implementations, such as on GPUs, are simple and fast. Experiments demonstrate that this approach is competitive with state-of-the-art tractable distribution estimators. At test time, the method is significantly faster and scales better than other autoregressive estimators.
* 9 pages and 1 page of supplementary material. Updated to match published version
Incorporating Side Information in Probabilistic Matrix Factorization with Gaussian Processes
Aug 09, 2014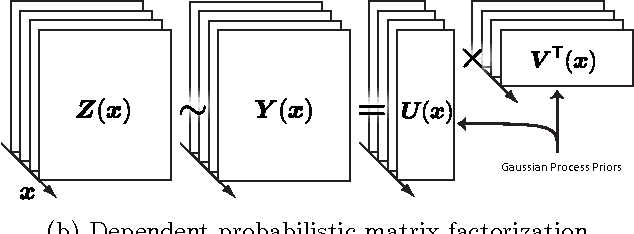
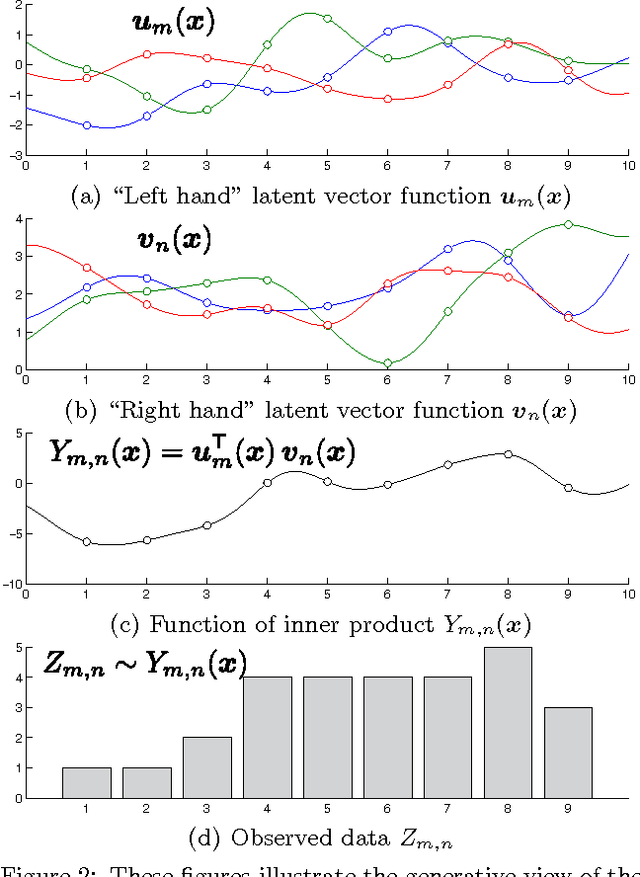
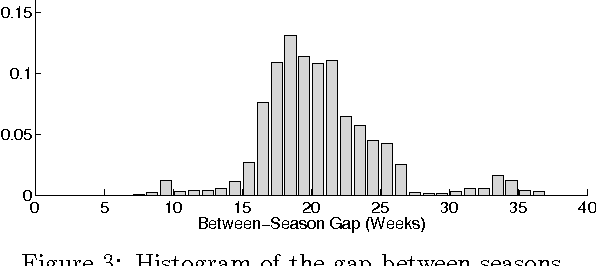
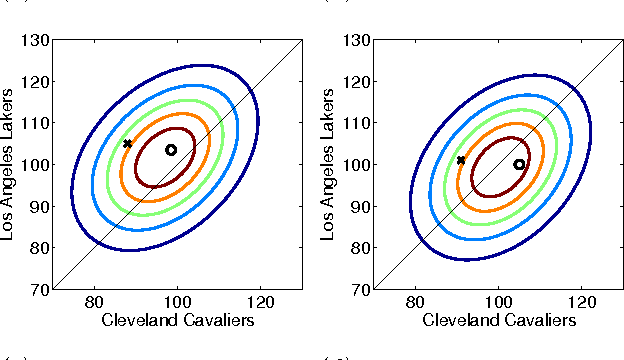
Abstract:Probabilistic matrix factorization (PMF) is a powerful method for modeling data associ- ated with pairwise relationships, Finding use in collaborative Filtering, computational bi- ology, and document analysis, among other areas. In many domains, there are additional covariates that can assist in prediction. For example, when modeling movie ratings, we might know when the rating occurred, where the user lives, or what actors appear in the movie. It is difficult, however, to incorporate this side information into the PMF model. We propose a framework for incorporating side information by coupling together multi- ple PMF problems via Gaussian process priors. We replace scalar latent features with func- tions that vary over the covariate space. The GP priors on these functions require them to vary smoothly and share information. We apply this new method to predict the scores of professional basketball games, where side information about the venue and date of the game are relevant for the outcome.
Parallel MCMC with Generalized Elliptical Slice Sampling
Jul 24, 2014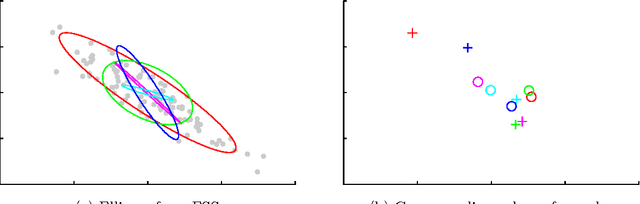
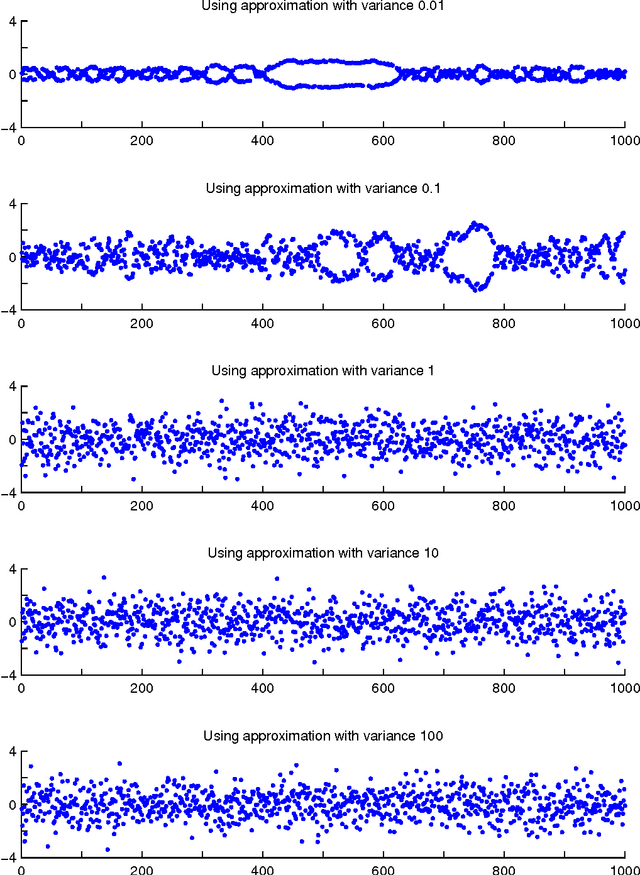
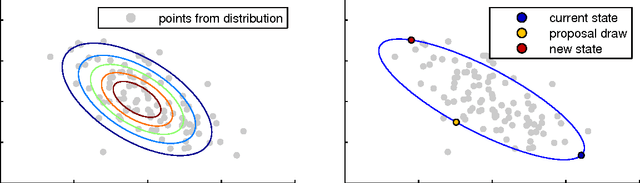
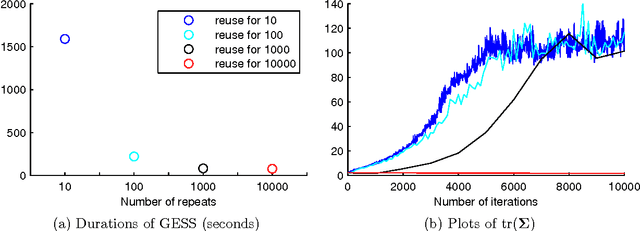
Abstract:Probabilistic models are conceptually powerful tools for finding structure in data, but their practical effectiveness is often limited by our ability to perform inference in them. Exact inference is frequently intractable, so approximate inference is often performed using Markov chain Monte Carlo (MCMC). To achieve the best possible results from MCMC, we want to efficiently simulate many steps of a rapidly mixing Markov chain which leaves the target distribution invariant. Of particular interest in this regard is how to take advantage of multi-core computing to speed up MCMC-based inference, both to improve mixing and to distribute the computational load. In this paper, we present a parallelizable Markov chain Monte Carlo algorithm for efficiently sampling from continuous probability distributions that can take advantage of hundreds of cores. This method shares information between parallel Markov chains to build a scale-mixture of Gaussians approximation to the density function of the target distribution. We combine this approximation with a recent method known as elliptical slice sampling to create a Markov chain with no step-size parameters that can mix rapidly without requiring gradient or curvature computations.
* 19 pages, 8 figures, 3 algorithms
A Deep and Tractable Density Estimator
Jan 11, 2014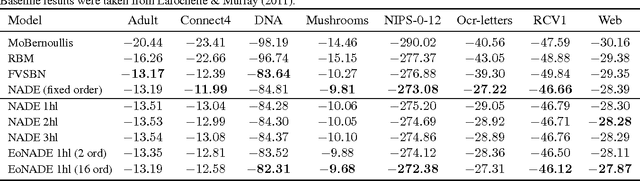
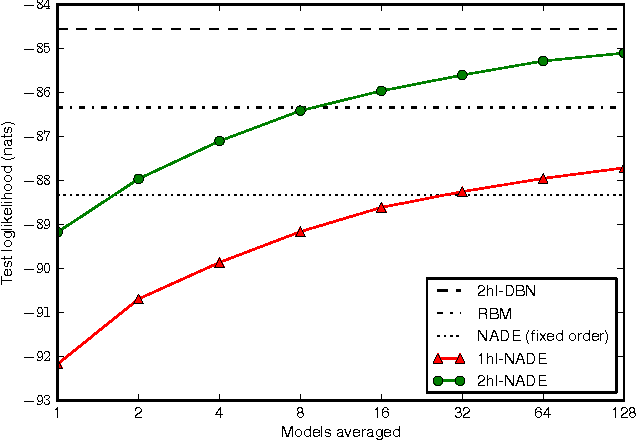

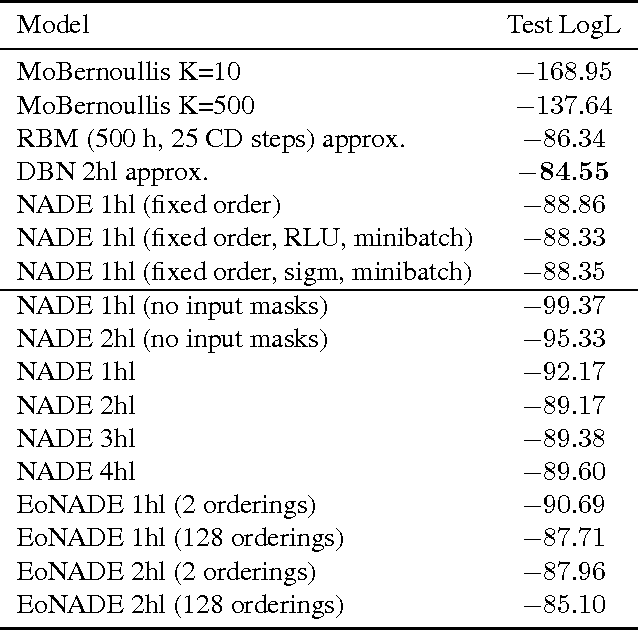
Abstract:The Neural Autoregressive Distribution Estimator (NADE) and its real-valued version RNADE are competitive density models of multidimensional data across a variety of domains. These models use a fixed, arbitrary ordering of the data dimensions. One can easily condition on variables at the beginning of the ordering, and marginalize out variables at the end of the ordering, however other inference tasks require approximate inference. In this work we introduce an efficient procedure to simultaneously train a NADE model for each possible ordering of the variables, by sharing parameters across all these models. We can thus use the most convenient model for each inference task at hand, and ensembles of such models with different orderings are immediately available. Moreover, unlike the original NADE, our training procedure scales to deep models. Empirically, ensembles of Deep NADE models obtain state of the art density estimation performance.
RNADE: The real-valued neural autoregressive density-estimator
Jan 09, 2014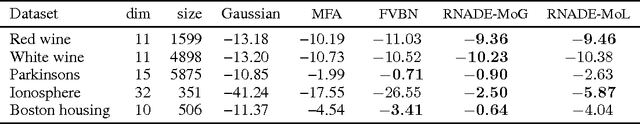

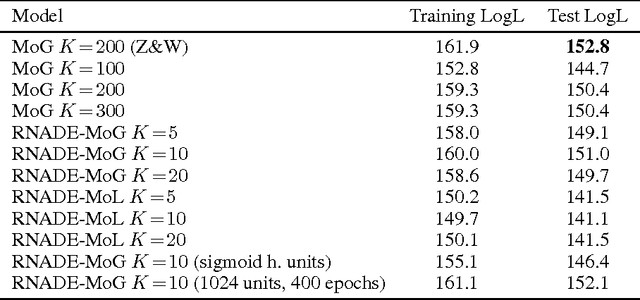

Abstract:We introduce RNADE, a new model for joint density estimation of real-valued vectors. Our model calculates the density of a datapoint as the product of one-dimensional conditionals modeled using mixture density networks with shared parameters. RNADE learns a distributed representation of the data, while having a tractable expression for the calculation of densities. A tractable likelihood allows direct comparison with other methods and training by standard gradient-based optimizers. We compare the performance of RNADE on several datasets of heterogeneous and perceptual data, finding it outperforms mixture models in all but one case.
* 12 pages, 3 figures, 3 tables, 2 algorithms. Merges the published paper and supplementary material into one document
A Framework for Evaluating Approximation Methods for Gaussian Process Regression
Nov 05, 2012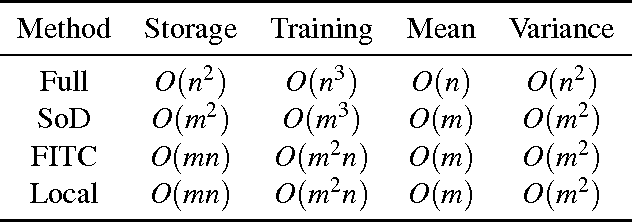
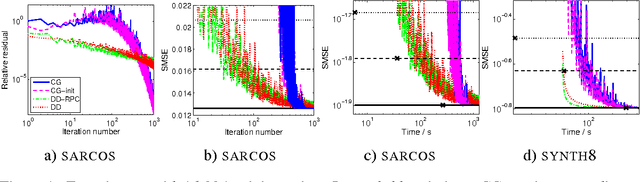
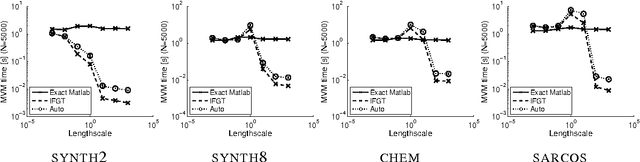
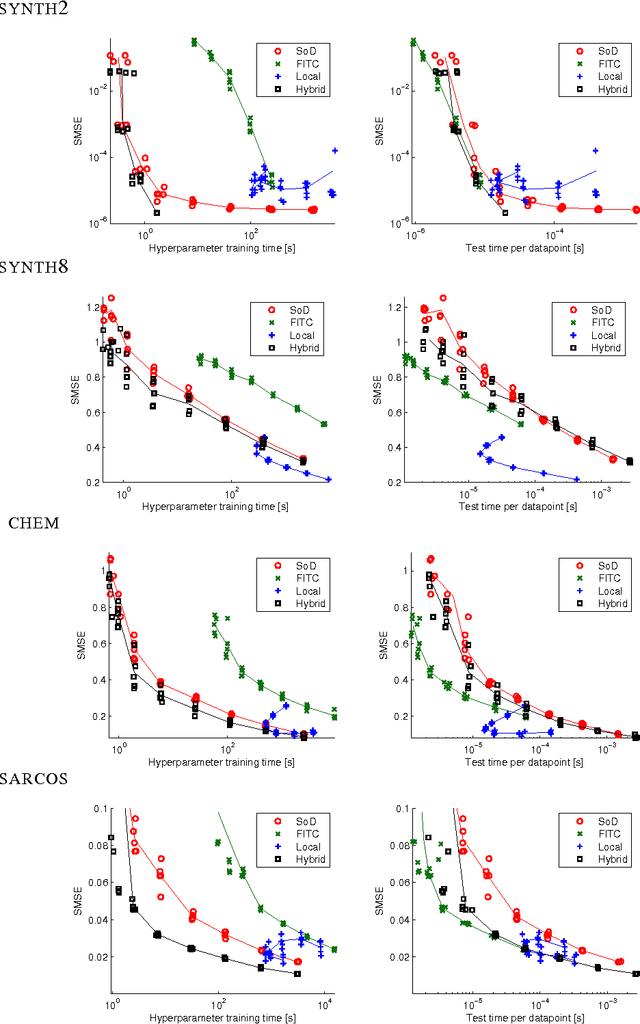
Abstract:Gaussian process (GP) predictors are an important component of many Bayesian approaches to machine learning. However, even a straightforward implementation of Gaussian process regression (GPR) requires O(n^2) space and O(n^3) time for a dataset of n examples. Several approximation methods have been proposed, but there is a lack of understanding of the relative merits of the different approximations, and in what situations they are most useful. We recommend assessing the quality of the predictions obtained as a function of the compute time taken, and comparing to standard baselines (e.g., Subset of Data and FITC). We empirically investigate four different approximation algorithms on four different prediction problems, and make our code available to encourage future comparisons.
Bayesian Learning in Undirected Graphical Models: Approximate MCMC algorithms
Jul 11, 2012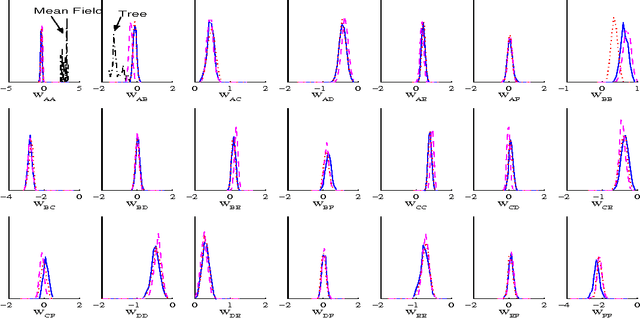
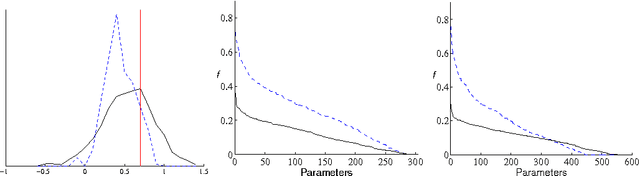
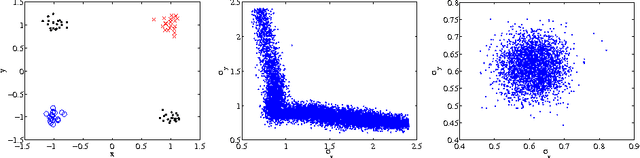
Abstract:Bayesian learning in undirected graphical models|computing posterior distributions over parameters and predictive quantities is exceptionally difficult. We conjecture that for general undirected models, there are no tractable MCMC (Markov Chain Monte Carlo) schemes giving the correct equilibrium distribution over parameters. While this intractability, due to the partition function, is familiar to those performing parameter optimisation, Bayesian learning of posterior distributions over undirected model parameters has been unexplored and poses novel challenges. we propose several approximate MCMC schemes and test on fully observed binary models (Boltzmann machines) for a small coronary heart disease data set and larger artificial systems. While approximations must perform well on the model, their interaction with the sampling scheme is also important. Samplers based on variational mean- field approximations generally performed poorly, more advanced methods using loopy propagation, brief sampling and stochastic dynamics lead to acceptable parameter posteriors. Finally, we demonstrate these techniques on a Markov random field with hidden variables.
Slice sampling covariance hyperparameters of latent Gaussian models
Oct 28, 2010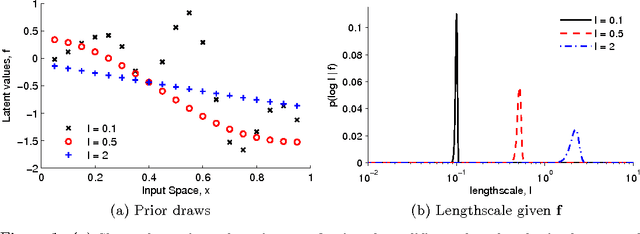
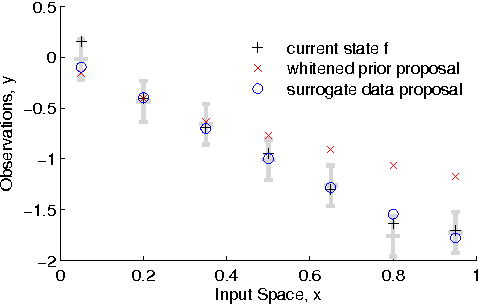
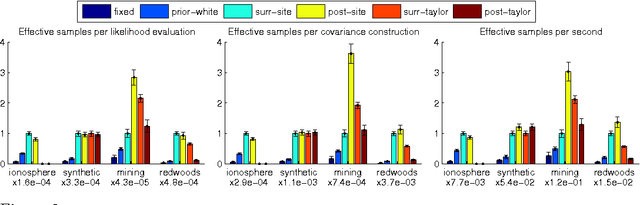
Abstract:The Gaussian process (GP) is a popular way to specify dependencies between random variables in a probabilistic model. In the Bayesian framework the covariance structure can be specified using unknown hyperparameters. Integrating over these hyperparameters considers different possible explanations for the data when making predictions. This integration is often performed using Markov chain Monte Carlo (MCMC) sampling. However, with non-Gaussian observations standard hyperparameter sampling approaches require careful tuning and may converge slowly. In this paper we present a slice sampling approach that requires little tuning while mixing well in both strong- and weak-data regimes.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge