Hung Q. Vo
MIRAM: Masked Image Reconstruction Across Multiple Scales for Breast Lesion Risk Prediction
Mar 10, 2025Abstract:Self-supervised learning (SSL) has garnered substantial interest within the machine learning and computer vision communities. Two prominent approaches in SSL include contrastive-based learning and self-distillation utilizing cropping augmentation. Lately, masked image modeling (MIM) has emerged as a more potent SSL technique, employing image inpainting as a pretext task. MIM creates a strong inductive bias toward meaningful spatial and semantic understanding. This has opened up new opportunities for SSL to contribute not only to classification tasks but also to more complex applications like object detection and image segmentation. Building upon this progress, our research paper introduces a scalable and practical SSL approach centered around more challenging pretext tasks that facilitate the acquisition of robust features. Specifically, we leverage multi-scale image reconstruction from randomly masked input images as the foundation for feature learning. Our hypothesis posits that reconstructing high-resolution images enables the model to attend to finer spatial details, particularly beneficial for discerning subtle intricacies within medical images. The proposed SSL features help improve classification performance on the Curated Breast Imaging Subset of Digital Database for Screening Mammography (CBIS-DDSM) dataset. In pathology classification, our method demonstrates a 3\% increase in average precision (AP) and a 1\% increase in the area under the receiver operating characteristic curve (AUC) when compared to state-of-the-art (SOTA) algorithms. Moreover, in mass margins classification, our approach achieves a 4\% increase in AP and a 2\% increase in AUC.
Revisiting Invariant Learning for Out-of-Domain Generalization on Multi-Site Mammogram Datasets
Mar 09, 2025Abstract:Despite significant progress in robust deep learning techniques for mammogram breast cancer classification, their reliability in real-world clinical development settings remains uncertain. The translation of these models to clinical practice faces challenges due to variations in medical centers, imaging protocols, and patient populations. To enhance their robustness, invariant learning methods have been proposed, prioritizing causal factors over misleading features. However, their effectiveness in clinical development and impact on mammogram classification require investigation. This paper reassesses the application of invariant learning for breast cancer risk estimation based on mammograms. Utilizing diverse multi-site public datasets, it represents the first study in this area. The objective is to evaluate invariant learning's benefits in developing robust models. Invariant learning methods, including Invariant Risk Minimization and Variance Risk Extrapolation, are compared quantitatively against Empirical Risk Minimization. Evaluation metrics include accuracy, average precision, and area under the curve. Additionally, interpretability is examined through class activation maps and visualization of learned representations. This research examines the advantages, limitations, and challenges of invariant learning for mammogram classification, guiding future studies to develop generalized methods for breast cancer prediction on whole mammograms in out-of-domain scenarios.
Student Collaboration Improves Self-Supervised Learning: Dual-Loss Adaptive Masked Autoencoder for Brain Cell Image Analysis
May 10, 2022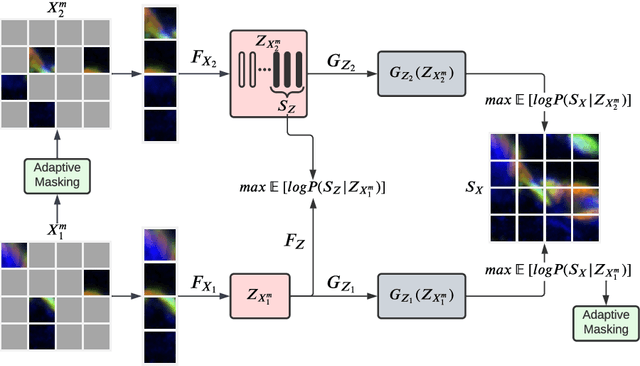
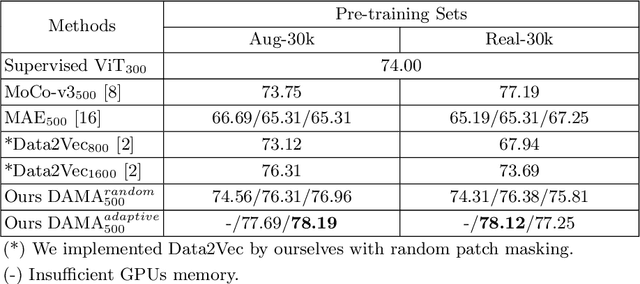
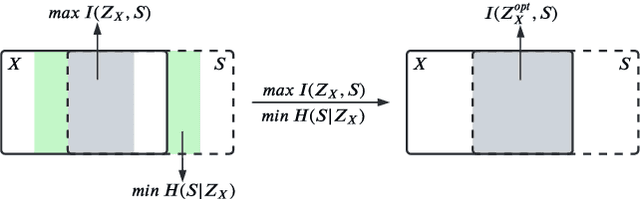
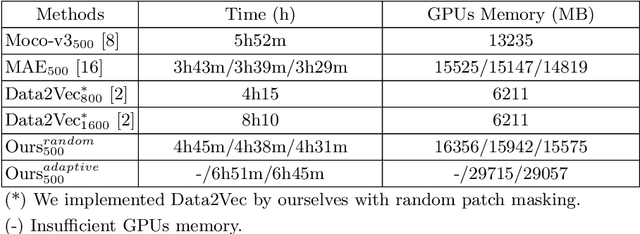
Abstract:Self-supervised learning leverages the underlying data structure as the source of the supervisory signal without the need for human annotation effort. This approach offers a practical solution to learning with a large amount of biomedical data and limited annotation. Unlike other studies exploiting data via multi-view (e.g., augmented images), this study presents a self-supervised Dual-Loss Adaptive Masked Autoencoder (DAMA) algorithm established from the viewpoint of the information theory. Specifically, our objective function maximizes the mutual information by minimizing the conditional entropy in pixel-level reconstruction and feature-level regression. We further introduce an adaptive mask sampling strategy to maximize mutual information. We conduct extensive experiments on brain cell images to validate the proposed method. DAMA significantly outperforms both state-of-the-art self-supervised and supervised methods on brain cells data and demonstrates competitive result on ImageNet-1k. Code: https://github.com/hula-ai/DAMA
Multimodal Breast Lesion Classification Using Cross-Attention Deep Networks
Aug 21, 2021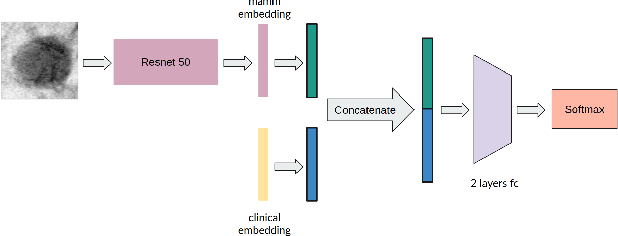
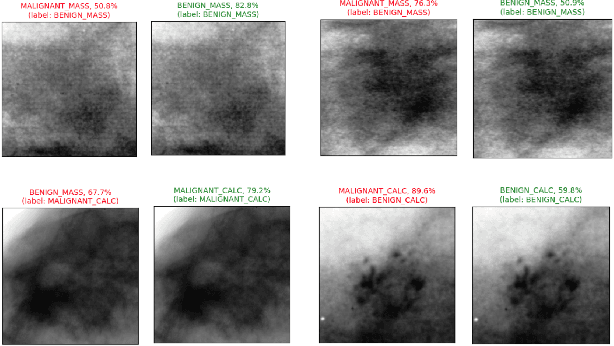
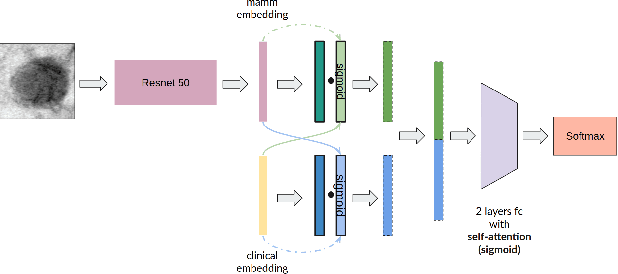
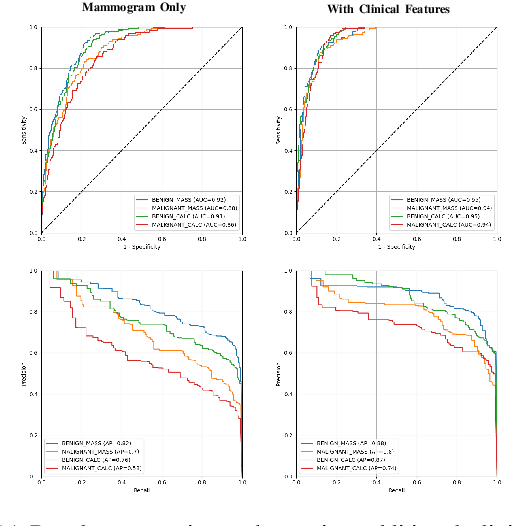

Abstract:Accurate breast lesion risk estimation can significantly reduce unnecessary biopsies and help doctors decide optimal treatment plans. Most existing computer-aided systems rely solely on mammogram features to classify breast lesions. While this approach is convenient, it does not fully exploit useful information in clinical reports to achieve the optimal performance. Would clinical features significantly improve breast lesion classification compared to using mammograms alone? How to handle missing clinical information caused by variation in medical practice? What is the best way to combine mammograms and clinical features? There is a compelling need for a systematic study to address these fundamental questions. This paper investigates several multimodal deep networks based on feature concatenation, cross-attention, and co-attention to combine mammograms and categorical clinical variables. We show that the proposed architectures significantly increase the lesion classification performance (average area under ROC curves from 0.89 to 0.94). We also evaluate the model when clinical variables are missing.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge