Hubert Banville
DANCE: Detect and Classify Events in EEG
May 11, 2026Abstract:Event identification in continuous neural recordings is a critical task in neuroscience. Decoding in EEG is dominated by classifying windows aligned to known event onsets. However, while available in controlled experiments, such onsets are absent in continuous real-world monitoring. Here, we introduce DANCE, a deep learning pipeline that frames neural decoding as a set-prediction problem and jointly detects and classifies events directly from raw, unaligned signals. Evaluated separately on ten datasets curated from the literature with a wide variety of event types (ranging from milliseconds to minutes in duration), our model outperforms existing methods on a broad range of cognitive, clinical and BCI tasks. This single architecture establishes a new state of the art in the competitive task of seizure monitoring and matches the accuracy of onset-informed models for BCI tasks. Overall, our method marks a step towards end-to-end asynchronous neural decoding models
A foundation model of vision, audition, and language for in-silico neuroscience
May 05, 2026Abstract:Cognitive neuroscience is fragmented into specialized models, each tailored to specific experimental paradigms, hence preventing a unified model of cognition in the human brain. Here, we introduce TRIBE v2, a tri-modal (video, audio and language) foundation model capable of predicting human brain activity in a variety of naturalistic and experimental conditions. Leveraging a unified dataset of over 1,000 hours of fMRI across 720 subjects, we demonstrate that our model accurately predicts high-resolution brain responses for novel stimuli, tasks and subjects, superseding traditional linear encoding models, delivering several-fold improvements in accuracy. Critically, TRIBE v2 enables in silico experimentation: tested on seminal visual and neuro-linguistic paradigms, it recovers a variety of results established by decades of empirical research. Finally, by extracting interpretable latent features, TRIBE v2 reveals the fine-grained topography of multisensory integration. These results establish artificial intelligence as a unifying framework for exploring the functional organization of the human brain.
From Minutes to Days: Scaling Intracranial Speech Decoding with Supervised Pretraining
Dec 17, 2025Abstract:Decoding speech from brain activity has typically relied on limited neural recordings collected during short and highly controlled experiments. Here, we introduce a framework to leverage week-long intracranial and audio recordings from patients undergoing clinical monitoring, effectively increasing the training dataset size by over two orders of magnitude. With this pretraining, our contrastive learning model substantially outperforms models trained solely on classic experimental data, with gains that scale log-linearly with dataset size. Analysis of the learned representations reveals that, while brain activity represents speech features, its global structure largely drifts across days, highlighting the need for models that explicitly account for cross-day variability. Overall, our approach opens a scalable path toward decoding and modeling brain representations in both real-life and controlled task settings.
TRIBE: TRImodal Brain Encoder for whole-brain fMRI response prediction
Jul 29, 2025Abstract:Historically, neuroscience has progressed by fragmenting into specialized domains, each focusing on isolated modalities, tasks, or brain regions. While fruitful, this approach hinders the development of a unified model of cognition. Here, we introduce TRIBE, the first deep neural network trained to predict brain responses to stimuli across multiple modalities, cortical areas and individuals. By combining the pretrained representations of text, audio and video foundational models and handling their time-evolving nature with a transformer, our model can precisely model the spatial and temporal fMRI responses to videos, achieving the first place in the Algonauts 2025 brain encoding competition with a significant margin over competitors. Ablations show that while unimodal models can reliably predict their corresponding cortical networks (e.g. visual or auditory networks), they are systematically outperformed by our multimodal model in high-level associative cortices. Currently applied to perception and comprehension, our approach paves the way towards building an integrative model of representations in the human brain. Our code is available at https://github.com/facebookresearch/algonauts-2025.
Brain-to-Text Decoding: A Non-invasive Approach via Typing
Feb 18, 2025Abstract:Modern neuroprostheses can now restore communication in patients who have lost the ability to speak or move. However, these invasive devices entail risks inherent to neurosurgery. Here, we introduce a non-invasive method to decode the production of sentences from brain activity and demonstrate its efficacy in a cohort of 35 healthy volunteers. For this, we present Brain2Qwerty, a new deep learning architecture trained to decode sentences from either electro- (EEG) or magneto-encephalography (MEG), while participants typed briefly memorized sentences on a QWERTY keyboard. With MEG, Brain2Qwerty reaches, on average, a character-error-rate (CER) of 32% and substantially outperforms EEG (CER: 67%). For the best participants, the model achieves a CER of 19%, and can perfectly decode a variety of sentences outside of the training set. While error analyses suggest that decoding depends on motor processes, the analysis of typographical errors suggests that it also involves higher-level cognitive factors. Overall, these results narrow the gap between invasive and non-invasive methods and thus open the path for developing safe brain-computer interfaces for non-communicating patients.
Scaling laws for decoding images from brain activity
Jan 25, 2025



Abstract:Generative AI has recently propelled the decoding of images from brain activity. How do these approaches scale with the amount and type of neural recordings? Here, we systematically compare image decoding from four types of non-invasive devices: electroencephalography (EEG), magnetoencephalography (MEG), high-field functional Magnetic Resonance Imaging (3T fMRI) and ultra-high field (7T) fMRI. For this, we evaluate decoding models on the largest benchmark to date, encompassing 8 public datasets, 84 volunteers, 498 hours of brain recording and 2.3 million brain responses to natural images. Unlike previous work, we focus on single-trial decoding performance to simulate real-time settings. This systematic comparison reveals three main findings. First, the most precise neuroimaging devices tend to yield the best decoding performances, when the size of the training sets are similar. However, the gain enabled by deep learning - in comparison to linear models - is obtained with the noisiest devices. Second, we do not observe any plateau of decoding performance as the amount of training data increases. Rather, decoding performance scales log-linearly with the amount of brain recording. Third, this scaling law primarily depends on the amount of data per subject. However, little decoding gain is observed by increasing the number of subjects. Overall, these findings delineate the path most suitable to scale the decoding of images from non-invasive brain recordings.
Decoding individual words from non-invasive brain recordings across 723 participants
Dec 11, 2024Abstract:Deep learning has recently enabled the decoding of language from the neural activity of a few participants with electrodes implanted inside their brain. However, reliably decoding words from non-invasive recordings remains an open challenge. To tackle this issue, we introduce a novel deep learning pipeline to decode individual words from non-invasive electro- (EEG) and magneto-encephalography (MEG) signals. We train and evaluate our approach on an unprecedentedly large number of participants (723) exposed to five million words either written or spoken in English, French or Dutch. Our model outperforms existing methods consistently across participants, devices, languages, and tasks, and can decode words absent from the training set. Our analyses highlight the importance of the recording device and experimental protocol: MEG and reading are easier to decode than EEG and listening, respectively, and it is preferable to collect a large amount of data per participant than to repeat stimuli across a large number of participants. Furthermore, decoding performance consistently increases with the amount of (i) data used for training and (ii) data used for averaging during testing. Finally, single-word predictions show that our model effectively relies on word semantics but also captures syntactic and surface properties such as part-of-speech, word length and even individual letters, especially in the reading condition. Overall, our findings delineate the path and remaining challenges towards building non-invasive brain decoders for natural language.
Aligning brain functions boosts the decoding of visual semantics in novel subjects
Dec 11, 2023Abstract:Deep learning is leading to major advances in the realm of brain decoding from functional Magnetic Resonance Imaging (fMRI). However, the large inter-subject variability in brain characteristics has limited most studies to train models on one subject at a time. Consequently, this approach hampers the training of deep learning models, which typically requires very large datasets. Here, we propose to boost brain decoding by aligning brain responses to videos and static images across subjects. Compared to the anatomically-aligned baseline, our method improves out-of-subject decoding performance by up to 75%. Moreover, it also outperforms classical single-subject approaches when fewer than 100 minutes of data is available for the tested subject. Furthermore, we propose a new multi-subject alignment method, which obtains comparable results to that of classical single-subject approaches while improving out-of-subject generalization. Finally, we show that this method aligns neural representations in accordance with brain anatomy. Overall, this study lays the foundations for leveraging extensive neuroimaging datasets and enhancing the decoding of individuals with a limited amount of brain recordings.
Brain decoding: toward real-time reconstruction of visual perception
Oct 18, 2023



Abstract:In the past five years, the use of generative and foundational AI systems has greatly improved the decoding of brain activity. Visual perception, in particular, can now be decoded from functional Magnetic Resonance Imaging (fMRI) with remarkable fidelity. This neuroimaging technique, however, suffers from a limited temporal resolution ($\approx$0.5 Hz) and thus fundamentally constrains its real-time usage. Here, we propose an alternative approach based on magnetoencephalography (MEG), a neuroimaging device capable of measuring brain activity with high temporal resolution ($\approx$5,000 Hz). For this, we develop an MEG decoding model trained with both contrastive and regression objectives and consisting of three modules: i) pretrained embeddings obtained from the image, ii) an MEG module trained end-to-end and iii) a pretrained image generator. Our results are threefold: Firstly, our MEG decoder shows a 7X improvement of image-retrieval over classic linear decoders. Second, late brain responses to images are best decoded with DINOv2, a recent foundational image model. Third, image retrievals and generations both suggest that MEG signals primarily contain high-level visual features, whereas the same approach applied to 7T fMRI also recovers low-level features. Overall, these results provide an important step towards the decoding - in real time - of the visual processes continuously unfolding within the human brain.
Robust learning from corrupted EEG with dynamic spatial filtering
May 27, 2021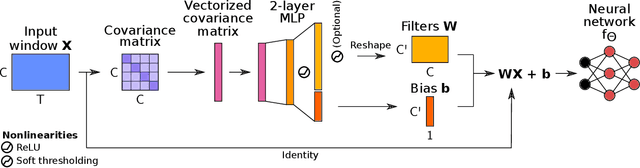
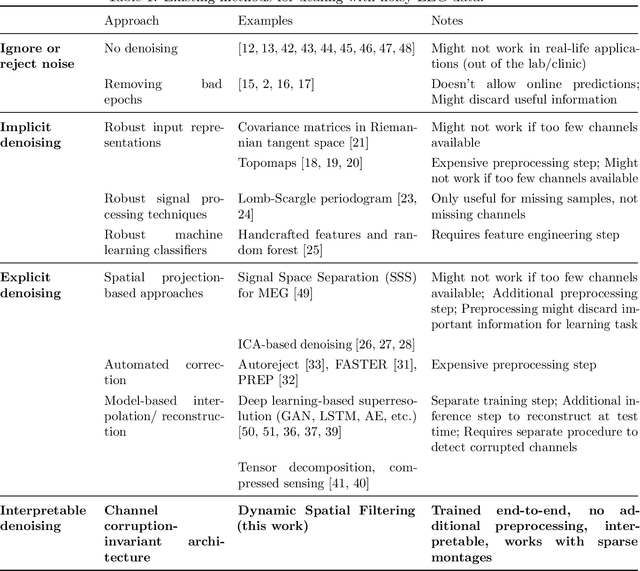
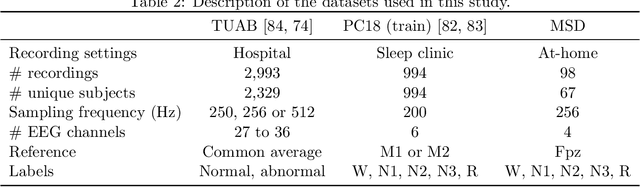
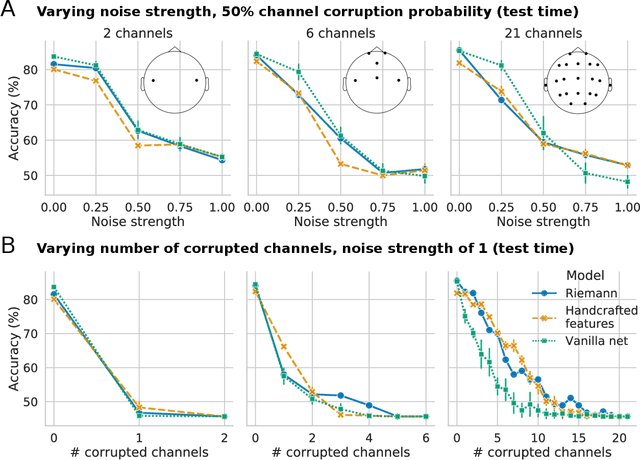
Abstract:Building machine learning models using EEG recorded outside of the laboratory setting requires methods robust to noisy data and randomly missing channels. This need is particularly great when working with sparse EEG montages (1-6 channels), often encountered in consumer-grade or mobile EEG devices. Neither classical machine learning models nor deep neural networks trained end-to-end on EEG are typically designed or tested for robustness to corruption, and especially to randomly missing channels. While some studies have proposed strategies for using data with missing channels, these approaches are not practical when sparse montages are used and computing power is limited (e.g., wearables, cell phones). To tackle this problem, we propose dynamic spatial filtering (DSF), a multi-head attention module that can be plugged in before the first layer of a neural network to handle missing EEG channels by learning to focus on good channels and to ignore bad ones. We tested DSF on public EEG data encompassing ~4,000 recordings with simulated channel corruption and on a private dataset of ~100 at-home recordings of mobile EEG with natural corruption. Our proposed approach achieves the same performance as baseline models when no noise is applied, but outperforms baselines by as much as 29.4% accuracy when significant channel corruption is present. Moreover, DSF outputs are interpretable, making it possible to monitor channel importance in real-time. This approach has the potential to enable the analysis of EEG in challenging settings where channel corruption hampers the reading of brain signals.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge