Holger R. Roth
NVIDIA, USA
Multi-scale Image Fusion Between Pre-operative Clinical CT and X-ray Microtomography of Lung Pathology
Feb 27, 2017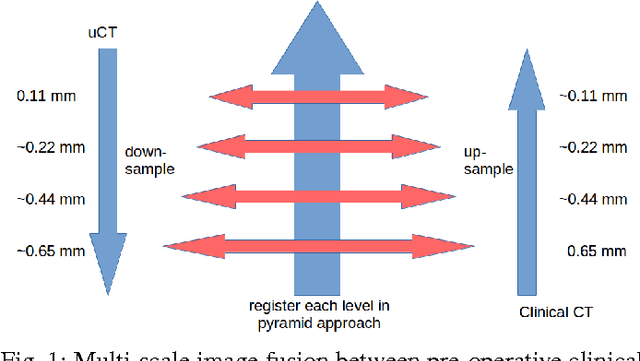
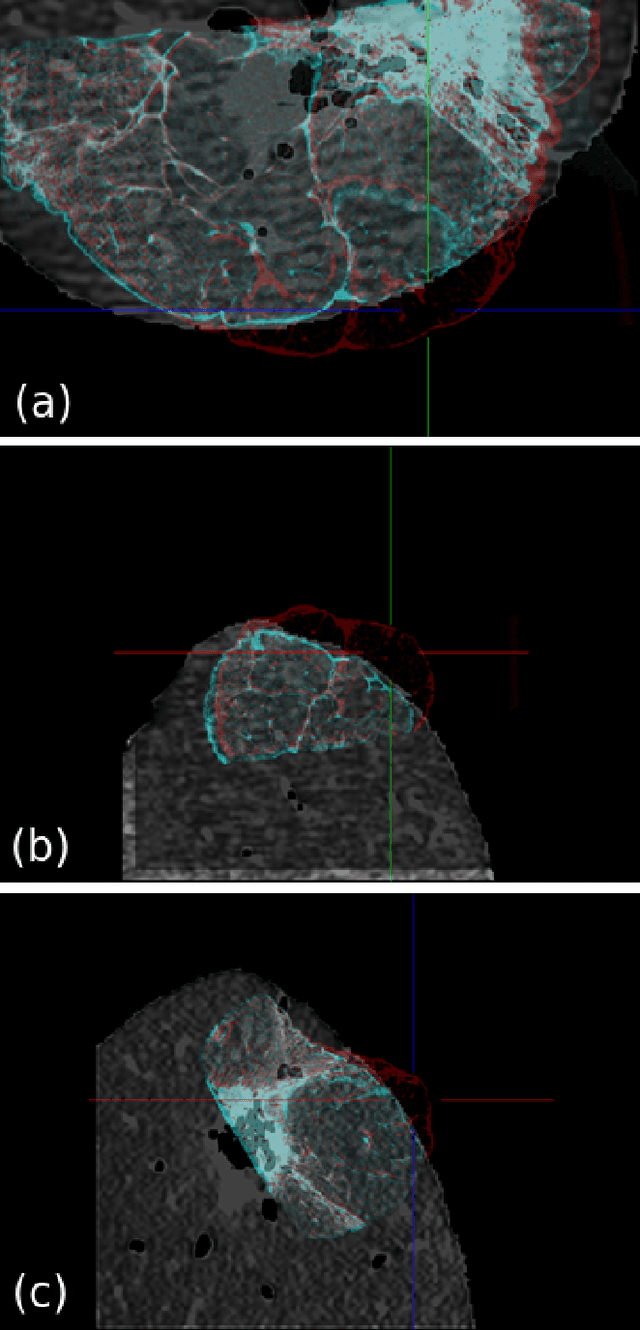
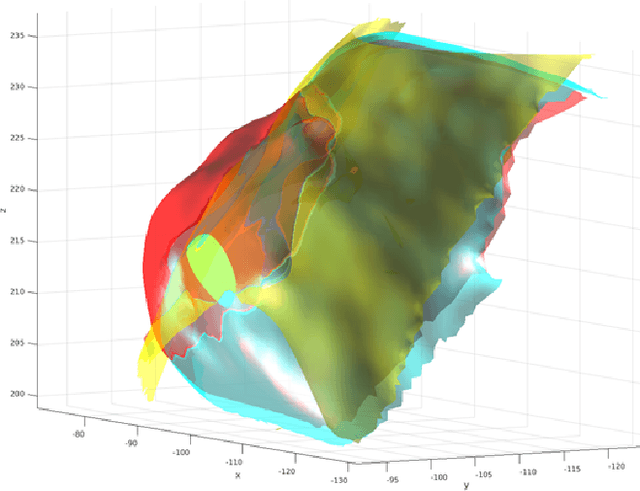
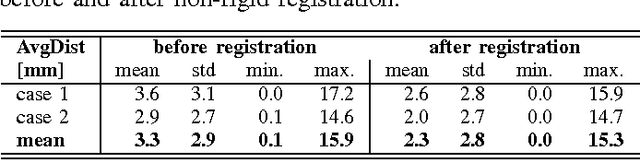
Abstract:Computational anatomy allows the quantitative analysis of organs in medical images. However, most analysis is constrained to the millimeter scale because of the limited resolution of clinical computed tomography (CT). X-ray microtomography ($\mu$CT) on the other hand allows imaging of ex-vivo tissues at a resolution of tens of microns. In this work, we use clinical CT to image lung cancer patients before partial pneumonectomy (resection of pathological lung tissue). The resected specimen is prepared for $\mu$CT imaging at a voxel resolution of 50 $\mu$m (0.05 mm). This high-resolution image of the lung cancer tissue allows further insides into understanding of tumor growth and categorization. For making full use of this additional information, image fusion (registration) needs to be performed in order to re-align the $\mu$CT image with clinical CT. We developed a multi-scale non-rigid registration approach. After manual initialization using a few landmark points and rigid alignment, several levels of non-rigid registration between down-sampled (in the case of $\mu$CT) and up-sampled (in the case of clinical CT) representations of the image are performed. Any non-lung tissue is ignored during the computation of the similarity measure used to guide the registration during optimization. We are able to recover the volume differences introduced by the resection and preparation of the lung specimen. The average ($\pm$ std. dev.) minimum surface distance between $\mu$CT and clinical CT at the resected lung surface is reduced from 3.3 $\pm$ 2.9 (range: [0.1, 15.9]) to 2.3 mm $\pm$ 2.8 (range: [0.0, 15.3]) mm. The alignment of clinical CT with $\mu$CT will allow further registration with even finer resolutions of $\mu$CT (up to 10 $\mu$m resolution) and ultimately with histopathological microscopy images for further macro to micro image fusion that can aid medical image analysis.
Spatial Aggregation of Holistically-Nested Convolutional Neural Networks for Automated Pancreas Localization and Segmentation
Jan 31, 2017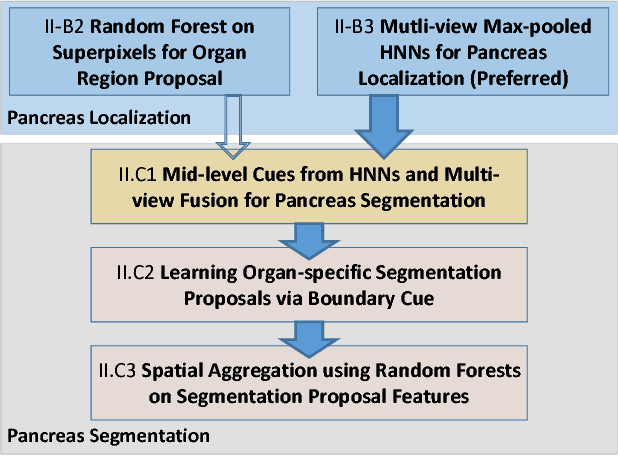
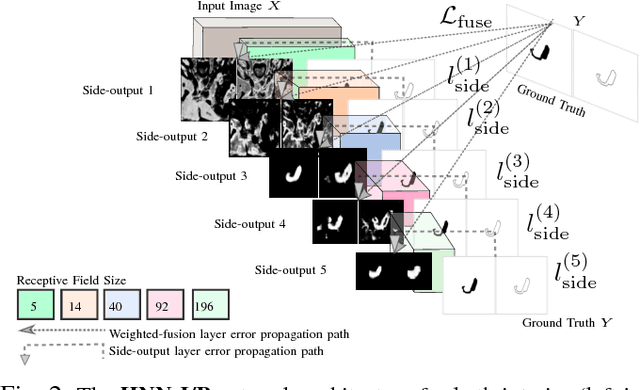
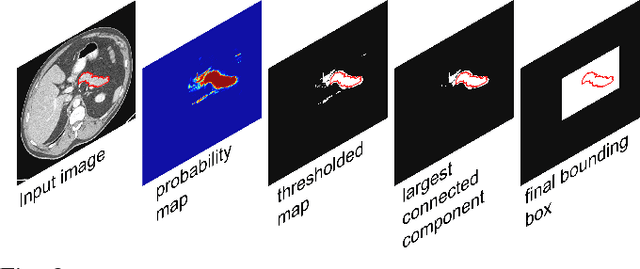
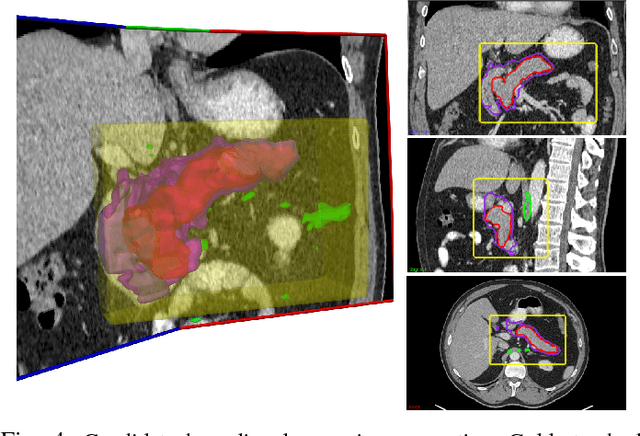
Abstract:Accurate and automatic organ segmentation from 3D radiological scans is an important yet challenging problem for medical image analysis. Specifically, the pancreas demonstrates very high inter-patient anatomical variability in both its shape and volume. In this paper, we present an automated system using 3D computed tomography (CT) volumes via a two-stage cascaded approach: pancreas localization and segmentation. For the first step, we localize the pancreas from the entire 3D CT scan, providing a reliable bounding box for the more refined segmentation step. We introduce a fully deep-learning approach, based on an efficient application of holistically-nested convolutional networks (HNNs) on the three orthogonal axial, sagittal, and coronal views. The resulting HNN per-pixel probability maps are then fused using pooling to reliably produce a 3D bounding box of the pancreas that maximizes the recall. We show that our introduced localizer compares favorably to both a conventional non-deep-learning method and a recent hybrid approach based on spatial aggregation of superpixels using random forest classification. The second, segmentation, phase operates within the computed bounding box and integrates semantic mid-level cues of deeply-learned organ interior and boundary maps, obtained by two additional and separate realizations of HNNs. By integrating these two mid-level cues, our method is capable of generating boundary-preserving pixel-wise class label maps that result in the final pancreas segmentation. Quantitative evaluation is performed on a publicly available dataset of 82 patient CT scans using 4-fold cross-validation (CV). We achieve a Dice similarity coefficient (DSC) of 81.27+/-6.27% in validation, which significantly outperforms previous state-of-the art methods that report DSCs of 71.80+/-10.70% and 78.01+/-8.20%, respectively, using the same dataset.
Spatial Aggregation of Holistically-Nested Networks for Automated Pancreas Segmentation
Jun 24, 2016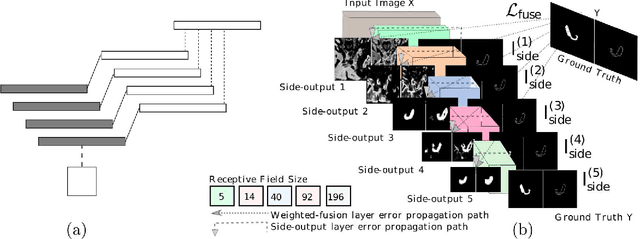
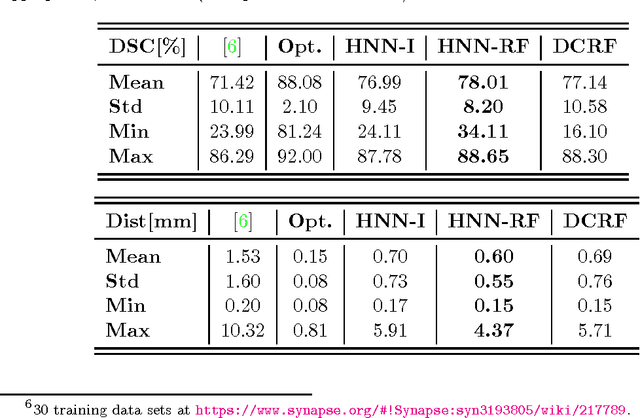
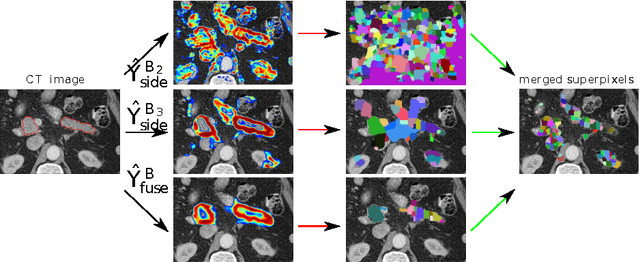
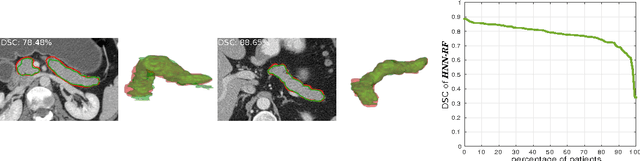
Abstract:Accurate automatic organ segmentation is an important yet challenging problem for medical image analysis. The pancreas is an abdominal organ with very high anatomical variability. This inhibits traditional segmentation methods from achieving high accuracies, especially compared to other organs such as the liver, heart or kidneys. In this paper, we present a holistic learning approach that integrates semantic mid-level cues of deeply-learned organ interior and boundary maps via robust spatial aggregation using random forest. Our method generates boundary preserving pixel-wise class labels for pancreas segmentation. Quantitative evaluation is performed on CT scans of 82 patients in 4-fold cross-validation. We achieve a (mean $\pm$ std. dev.) Dice Similarity Coefficient of 78.01% $\pm$ 8.2% in testing which significantly outperforms the previous state-of-the-art approach of 71.8% $\pm$ 10.7% under the same evaluation criterion.
A Bottom-up Approach for Pancreas Segmentation using Cascaded Superpixels and (Deep) Image Patch Labeling
Mar 07, 2016
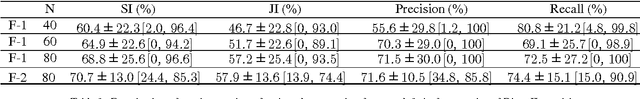

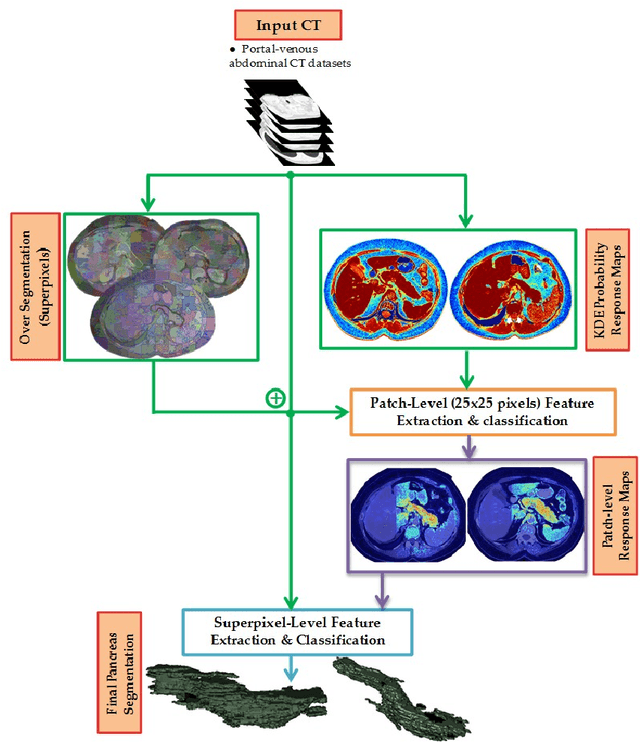
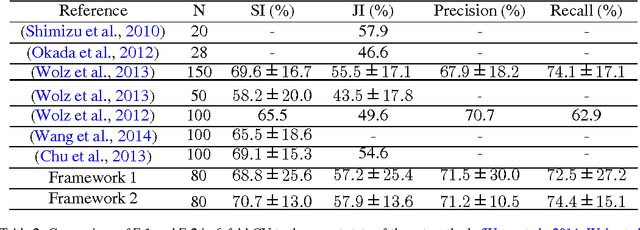
Abstract:Robust automated organ segmentation is a prerequisite for computer-aided diagnosis (CAD), quantitative imaging analysis and surgical assistance. For high-variability organs such as the pancreas, previous approaches report undesirably low accuracies. We present a bottom-up approach for pancreas segmentation in abdominal CT scans that is based on a hierarchy of information propagation by classifying image patches at different resolutions; and cascading superpixels. There are four stages: 1) decomposing CT slice images as a set of disjoint boundary-preserving superpixels; 2) computing pancreas class probability maps via dense patch labeling; 3) classifying superpixels by pooling both intensity and probability features to form empirical statistics in cascaded random forest frameworks; and 4) simple connectivity based post-processing. The dense image patch labeling are conducted by: efficient random forest classifier on image histogram, location and texture features; and more expensive (but with better specificity) deep convolutional neural network classification on larger image windows (with more spatial contexts). Evaluation of the approach is performed on a database of 80 manually segmented CT volumes in six-fold cross-validation (CV). Our achieved results are comparable, or better than the state-of-the-art methods (evaluated by "leave-one-patient-out"), with Dice 70.7% and Jaccard 57.9%. The computational efficiency has been drastically improved in the order of 6~8 minutes, comparing with others of ~10 hours per case. Finally, we implement a multi-atlas label fusion (MALF) approach for pancreas segmentation using the same datasets. Under six-fold CV, our bottom-up segmentation method significantly outperforms its MALF counterpart: (70.7 +/- 13.0%) versus (52.5 +/- 20.8%) in Dice. Deep CNN patch labeling confidences offer more numerical stability, reflected by smaller standard deviations.
Deep Convolutional Neural Networks for Computer-Aided Detection: CNN Architectures, Dataset Characteristics and Transfer Learning
Feb 10, 2016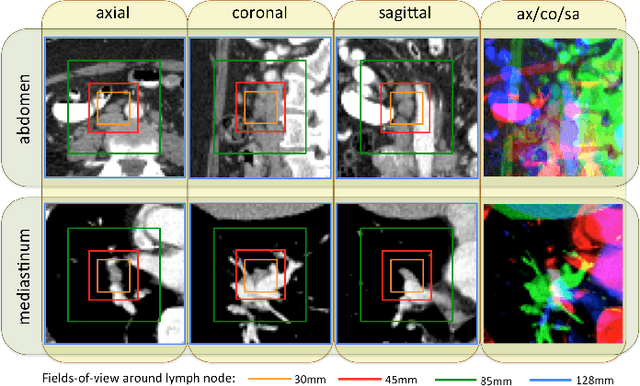
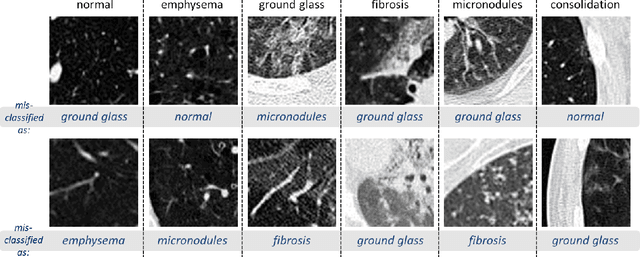
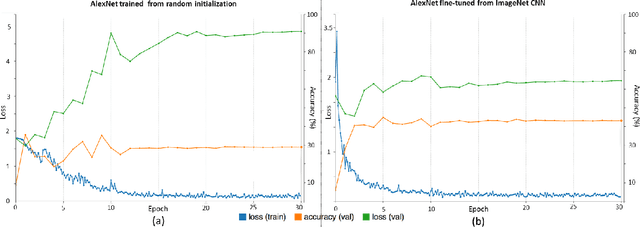

Abstract:Remarkable progress has been made in image recognition, primarily due to the availability of large-scale annotated datasets and the revival of deep CNN. CNNs enable learning data-driven, highly representative, layered hierarchical image features from sufficient training data. However, obtaining datasets as comprehensively annotated as ImageNet in the medical imaging domain remains a challenge. There are currently three major techniques that successfully employ CNNs to medical image classification: training the CNN from scratch, using off-the-shelf pre-trained CNN features, and conducting unsupervised CNN pre-training with supervised fine-tuning. Another effective method is transfer learning, i.e., fine-tuning CNN models pre-trained from natural image dataset to medical image tasks. In this paper, we exploit three important, but previously understudied factors of employing deep convolutional neural networks to computer-aided detection problems. We first explore and evaluate different CNN architectures. The studied models contain 5 thousand to 160 million parameters, and vary in numbers of layers. We then evaluate the influence of dataset scale and spatial image context on performance. Finally, we examine when and why transfer learning from pre-trained ImageNet (via fine-tuning) can be useful. We study two specific computer-aided detection (CADe) problems, namely thoraco-abdominal lymph node (LN) detection and interstitial lung disease (ILD) classification. We achieve the state-of-the-art performance on the mediastinal LN detection, with 85% sensitivity at 3 false positive per patient, and report the first five-fold cross-validation classification results on predicting axial CT slices with ILD categories. Our extensive empirical evaluation, CNN model analysis and valuable insights can be extended to the design of high performance CAD systems for other medical imaging tasks.
Improving Vertebra Segmentation through Joint Vertebra-Rib Atlases
Feb 01, 2016Abstract:Accurate spine segmentation allows for improved identification and quantitative characterization of abnormalities of the vertebra, such as vertebral fractures. However, in existing automated vertebra segmentation methods on computed tomography (CT) images, leakage into nearby bones such as ribs occurs due to the close proximity of these visibly intense structures in a 3D CT volume. To reduce this error, we propose the use of joint vertebra-rib atlases to improve the segmentation of vertebrae via multi-atlas joint label fusion. Segmentation was performed and evaluated on CTs containing 106 thoracic and lumbar vertebrae from 10 pathological and traumatic spine patients on an individual vertebra level basis. Vertebra atlases produced errors where the segmentation leaked into the ribs. The use of joint vertebra-rib atlases produced a statistically significant increase in the Dice coefficient from 92.5 $\pm$ 3.1% to 93.8 $\pm$ 2.1% for the left and right transverse processes and a decrease in the mean and max surface distance from 0.75 $\pm$ 0.60mm and 8.63 $\pm$ 4.44mm to 0.30 $\pm$ 0.27mm and 3.65 $\pm$ 2.87mm, respectively.
Deep convolutional networks for automated detection of posterior-element fractures on spine CT
Jan 29, 2016Abstract:Injuries of the spine, and its posterior elements in particular, are a common occurrence in trauma patients, with potentially devastating consequences. Computer-aided detection (CADe) could assist in the detection and classification of spine fractures. Furthermore, CAD could help assess the stability and chronicity of fractures, as well as facilitate research into optimization of treatment paradigms. In this work, we apply deep convolutional networks (ConvNets) for the automated detection of posterior element fractures of the spine. First, the vertebra bodies of the spine with its posterior elements are segmented in spine CT using multi-atlas label fusion. Then, edge maps of the posterior elements are computed. These edge maps serve as candidate regions for predicting a set of probabilities for fractures along the image edges using ConvNets in a 2.5D fashion (three orthogonal patches in axial, coronal and sagittal planes). We explore three different methods for training the ConvNet using 2.5D patches along the edge maps of 'positive', i.e. fractured posterior-elements and 'negative', i.e. non-fractured elements. An experienced radiologist retrospectively marked the location of 55 displaced posterior-element fractures in 18 trauma patients. We randomly split the data into training and testing cases. In testing, we achieve an area-under-the-curve of 0.857. This corresponds to 71% or 81% sensitivities at 5 or 10 false-positives per patient, respectively. Analysis of our set of trauma patients demonstrates the feasibility of detecting posterior-element fractures in spine CT images using computer vision techniques such as deep convolutional networks.
Multi-Atlas Segmentation with Joint Label Fusion of Osteoporotic Vertebral Compression Fractures on CT
Jan 13, 2016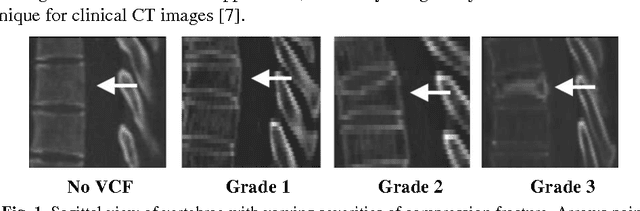
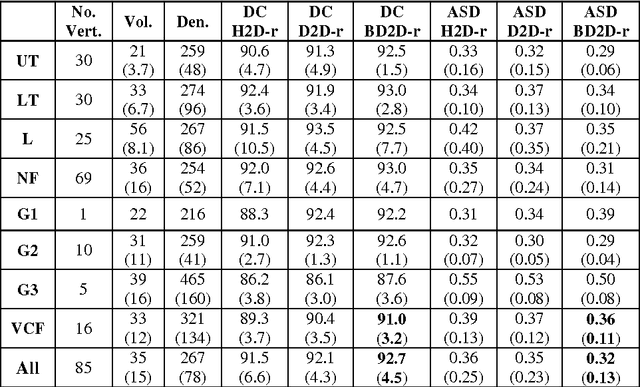
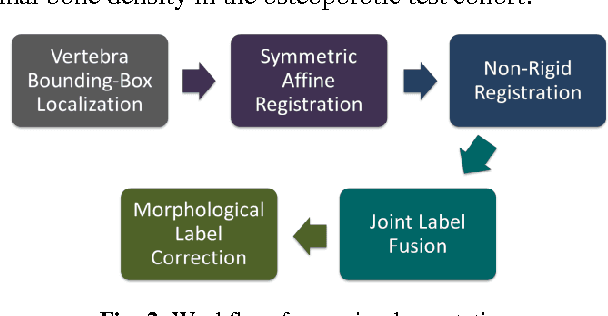
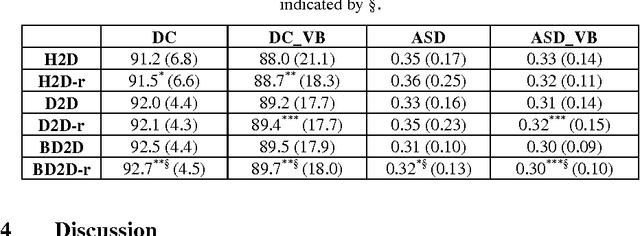
Abstract:The precise and accurate segmentation of the vertebral column is essential in the diagnosis and treatment of various orthopedic, neurological, and oncological traumas and pathologies. Segmentation is especially challenging in the presence of pathology such as vertebral compression fractures. In this paper, we propose a method to produce segmentations for osteoporotic compression fractured vertebrae by applying a multi-atlas joint label fusion technique for clinical CT images. A total of 170 thoracic and lumbar vertebrae were evaluated using atlases from five patients with varying degrees of spinal degeneration. In an osteoporotic cohort of bundled atlases, registration provided an average Dice coefficient and mean absolute surface distance of 2.7$\pm$4.5% and 0.32$\pm$0.13mm for osteoporotic vertebrae, respectively, and 90.9$\pm$3.0% and 0.36$\pm$0.11mm for compression fractured vertebrae.
Improving Computer-aided Detection using Convolutional Neural Networks and Random View Aggregation
Sep 15, 2015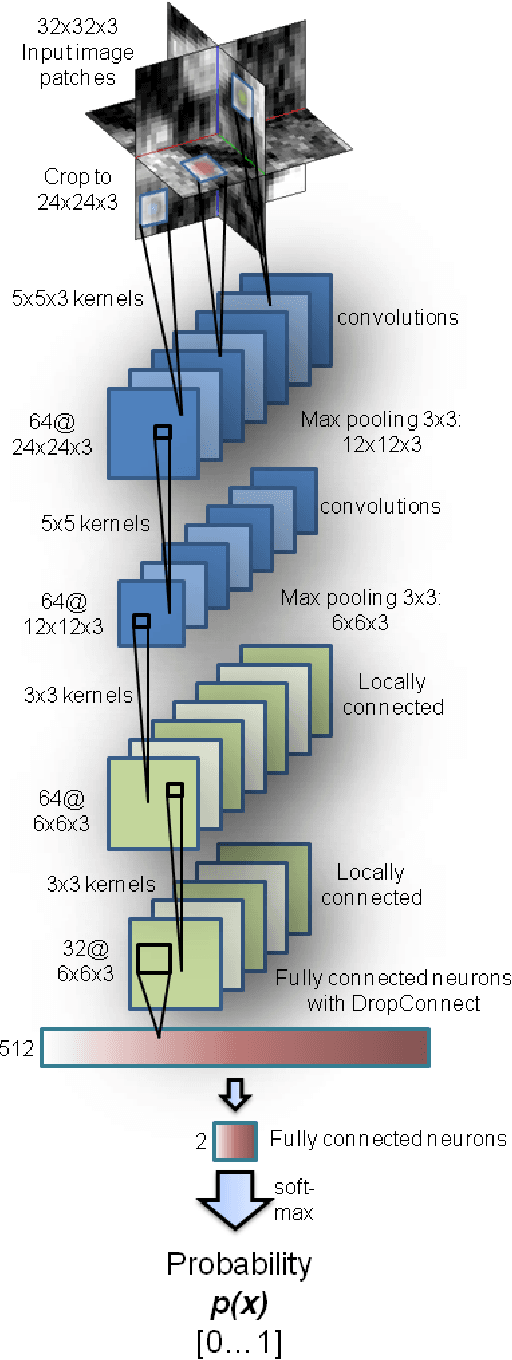
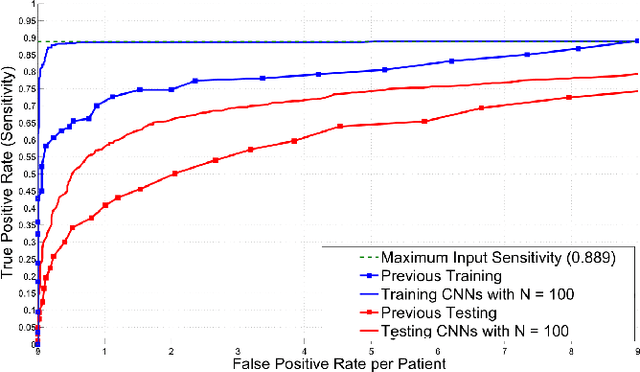
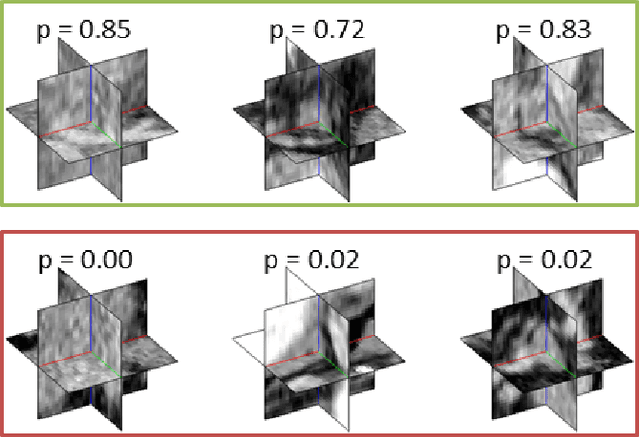
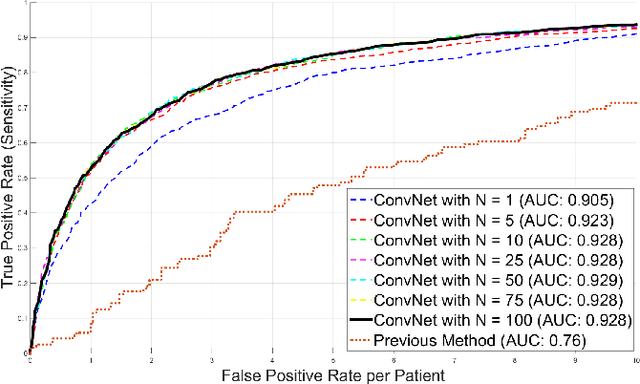
Abstract:Automated computer-aided detection (CADe) in medical imaging has been an important tool in clinical practice and research. State-of-the-art methods often show high sensitivities but at the cost of high false-positives (FP) per patient rates. We design a two-tiered coarse-to-fine cascade framework that first operates a candidate generation system at sensitivities of $\sim$100% but at high FP levels. By leveraging existing CAD systems, coordinates of regions or volumes of interest (ROI or VOI) for lesion candidates are generated in this step and function as input for a second tier, which is our focus in this study. In this second stage, we generate $N$ 2D (two-dimensional) or 2.5D views via sampling through scale transformations, random translations and rotations with respect to each ROI's centroid coordinates. These random views are used to train deep convolutional neural network (ConvNet) classifiers. In testing, the trained ConvNets are employed to assign class (e.g., lesion, pathology) probabilities for a new set of $N$ random views that are then averaged at each ROI to compute a final per-candidate classification probability. This second tier behaves as a highly selective process to reject difficult false positives while preserving high sensitivities. The methods are evaluated on three different data sets with different numbers of patients: 59 patients for sclerotic metastases detection, 176 patients for lymph node detection, and 1,186 patients for colonic polyp detection. Experimental results show the ability of ConvNets to generalize well to different medical imaging CADe applications and scale elegantly to various data sets. Our proposed methods improve CADe performance markedly in all cases. CADe sensitivities improved from 57% to 70%, from 43% to 77% and from 58% to 75% at 3 FPs per patient for sclerotic metastases, lymph nodes and colonic polyps, respectively.
* 2D vs 2.5D vs 3D inputs and comparison to other standard classifiers such as SVM have been addressed by more experimentation and two completely new sections and figures. Results and Discussions have been updated accordingly
DeepOrgan: Multi-level Deep Convolutional Networks for Automated Pancreas Segmentation
Jun 22, 2015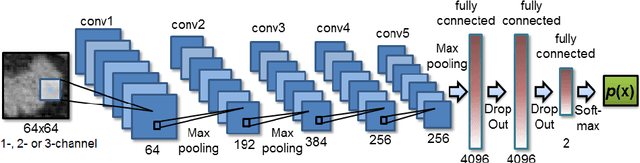
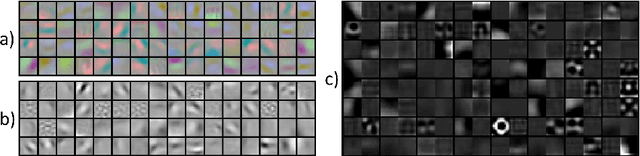
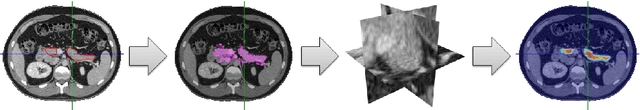
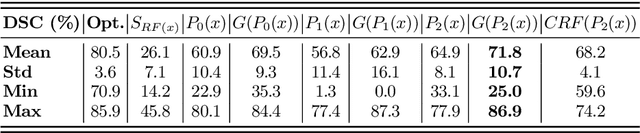


Abstract:Automatic organ segmentation is an important yet challenging problem for medical image analysis. The pancreas is an abdominal organ with very high anatomical variability. This inhibits previous segmentation methods from achieving high accuracies, especially compared to other organs such as the liver, heart or kidneys. In this paper, we present a probabilistic bottom-up approach for pancreas segmentation in abdominal computed tomography (CT) scans, using multi-level deep convolutional networks (ConvNets). We propose and evaluate several variations of deep ConvNets in the context of hierarchical, coarse-to-fine classification on image patches and regions, i.e. superpixels. We first present a dense labeling of local image patches via $P{-}\mathrm{ConvNet}$ and nearest neighbor fusion. Then we describe a regional ConvNet ($R_1{-}\mathrm{ConvNet}$) that samples a set of bounding boxes around each image superpixel at different scales of contexts in a "zoom-out" fashion. Our ConvNets learn to assign class probabilities for each superpixel region of being pancreas. Last, we study a stacked $R_2{-}\mathrm{ConvNet}$ leveraging the joint space of CT intensities and the $P{-}\mathrm{ConvNet}$ dense probability maps. Both 3D Gaussian smoothing and 2D conditional random fields are exploited as structured predictions for post-processing. We evaluate on CT images of 82 patients in 4-fold cross-validation. We achieve a Dice Similarity Coefficient of 83.6$\pm$6.3% in training and 71.8$\pm$10.7% in testing.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge