Georgios Exarchakis
Dissecting Self-Supervised Learning Methods for Surgical Computer Vision
Jul 01, 2022
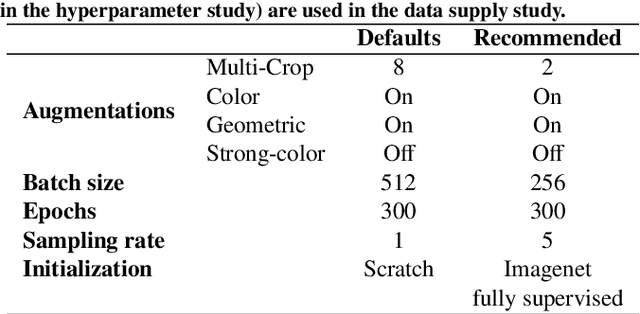
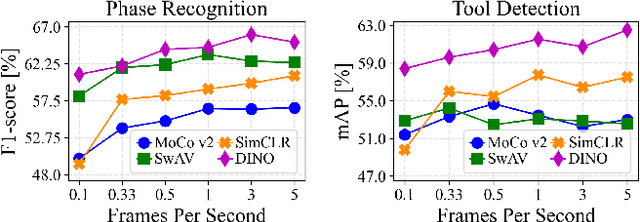
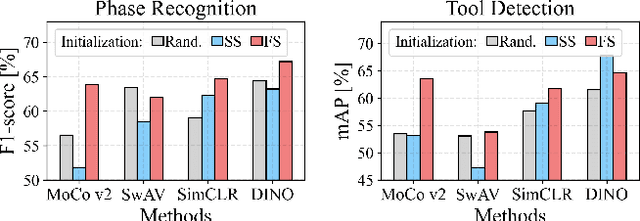
Abstract:The field of surgical computer vision has undergone considerable breakthroughs in recent years with the rising popularity of deep neural network-based methods. However, standard fully-supervised approaches for training such models require vast amounts of annotated data, imposing a prohibitively high cost; especially in the clinical domain. Self-Supervised Learning (SSL) methods, which have begun to gain traction in the general computer vision community, represent a potential solution to these annotation costs, allowing to learn useful representations from only unlabeled data. Still, the effectiveness of SSL methods in more complex and impactful domains, such as medicine and surgery, remains limited and unexplored. In this work, we address this critical need by investigating four state-of-the-art SSL methods (MoCo v2, SimCLR, DINO, SwAV) in the context of surgical computer vision. We present an extensive analysis of the performance of these methods on the Cholec80 dataset for two fundamental and popular tasks in surgical context understanding, phase recognition and tool presence detection. We examine their parameterization, then their behavior with respect to training data quantities in semi-supervised settings. Correct transfer of these methods to surgery, as described and conducted in this work, leads to substantial performance gains over generic uses of SSL - up to 7% on phase recognition and 20% on tool presence detection - as well as state-of-the-art semi-supervised phase recognition approaches by up to 14%. The code will be made available at https://github.com/CAMMA-public/SelfSupSurg.
A sampling-based approach for efficient clustering in large datasets
Dec 29, 2021
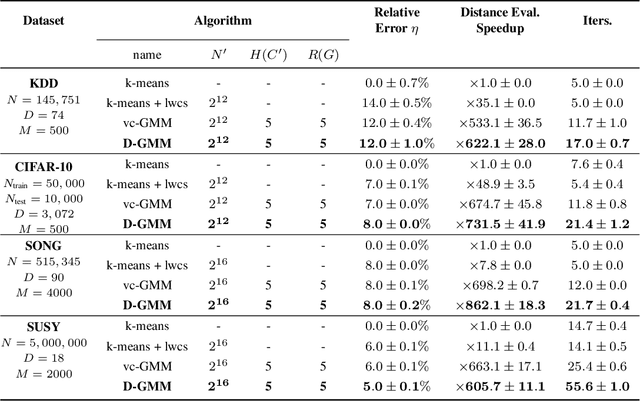
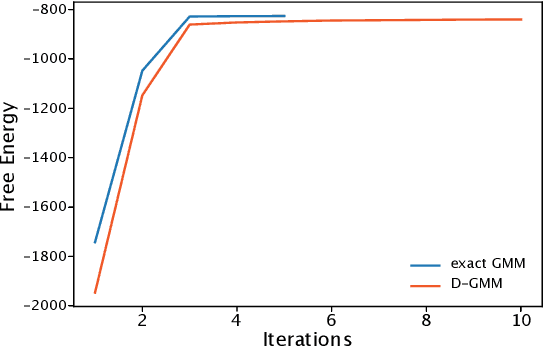
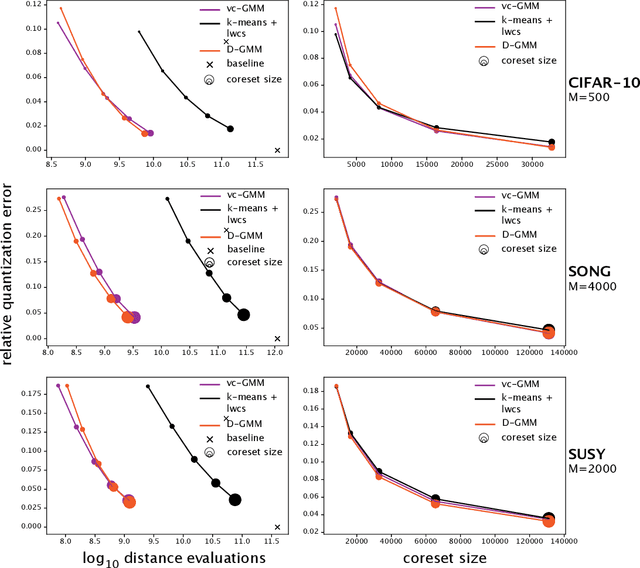
Abstract:We propose a simple and efficient clustering method for high-dimensional data with a large number of clusters. Our algorithm achieves high-performance by evaluating distances of datapoints with a subset of the cluster centres. Our contribution is substantially more efficient than k-means as it does not require an all to all comparison of data points and clusters. We show that the optimal solutions of our approximation are the same as in the exact solution. However, our approach is considerably more efficient at extracting these clusters compared to the state-of-the-art. We compare our approximation with the exact k-means and alternative approximation approaches on a series of standardised clustering tasks. For the evaluation, we consider the algorithmic complexity, including number of operations to convergence, and the stability of the results.
ProSper -- A Python Library for Probabilistic Sparse Coding with Non-Standard Priors and Superpositions
Aug 01, 2019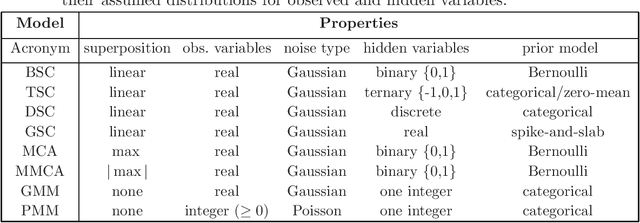
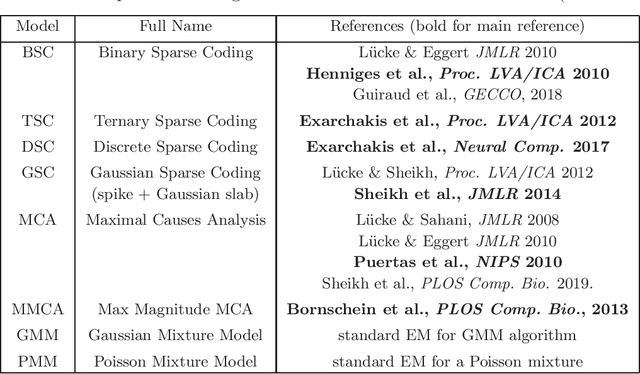
Abstract:ProSper is a python library containing probabilistic algorithms to learn dictionaries. Given a set of data points, the implemented algorithms seek to learn the elementary components that have generated the data. The library widens the scope of dictionary learning approaches beyond implementations of standard approaches such as ICA, NMF or standard L1 sparse coding. The implemented algorithms are especially well-suited in cases when data consist of components that combine non-linearly and/or for data requiring flexible prior distributions. Furthermore, the implemented algorithms go beyond standard approaches by inferring prior and noise parameters of the data, and they provide rich a-posteriori approximations for inference. The library is designed to be extendable and it currently includes: Binary Sparse Coding (BSC), Ternary Sparse Coding (TSC), Discrete Sparse Coding (DSC), Maximal Causes Analysis (MCA), Maximum Magnitude Causes Analysis (MMCA), and Gaussian Sparse Coding (GSC, a recent spike-and-slab sparse coding approach). The algorithms are scalable due to a combination of variational approximations and parallelization. Implementations of all algorithms allow for parallel execution on multiple CPUs and multiple machines for medium to large-scale applications. Typical large-scale runs of the algorithms can use hundreds of CPUs to learn hundreds of dictionary elements from data with tens of millions of floating-point numbers such that models with several hundred thousand parameters can be optimized. The library is designed to have minimal dependencies and to be easy to use. It targets users of dictionary learning algorithms and Machine Learning researchers.
Kymatio: Scattering Transforms in Python
Dec 28, 2018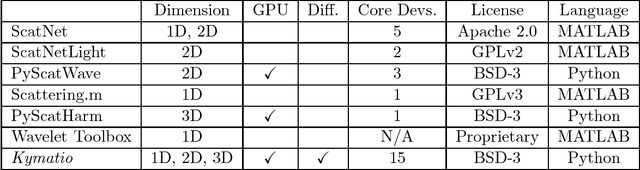
Abstract:The wavelet scattering transform is an invariant signal representation suitable for many signal processing and machine learning applications. We present the Kymatio software package, an easy-to-use, high-performance Python implementation of the scattering transform in 1D, 2D, and 3D that is compatible with modern deep learning frameworks. All transforms may be executed on a GPU (in addition to CPU), offering a considerable speed up over CPU implementations. The package also has a small memory footprint, resulting inefficient memory usage. The source code, documentation, and examples are available undera BSD license at https://www.kymat.io/
Solid Harmonic Wavelet Scattering for Predictions of Molecule Properties
May 01, 2018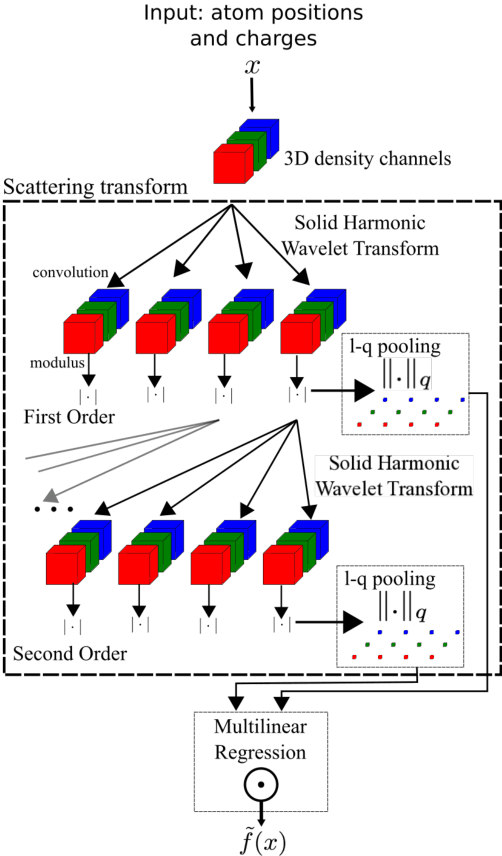
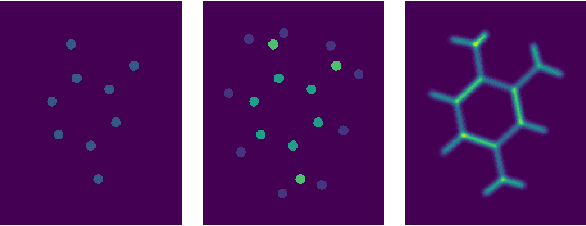

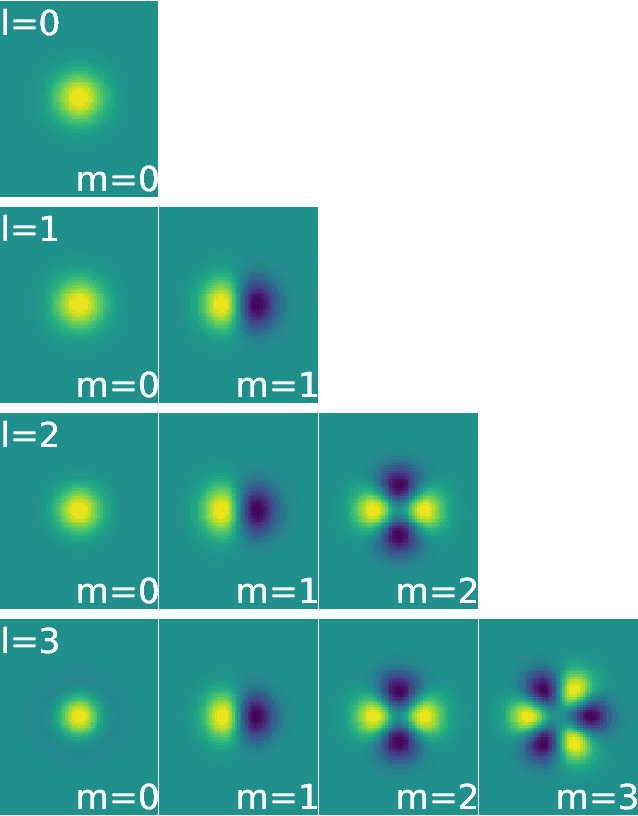
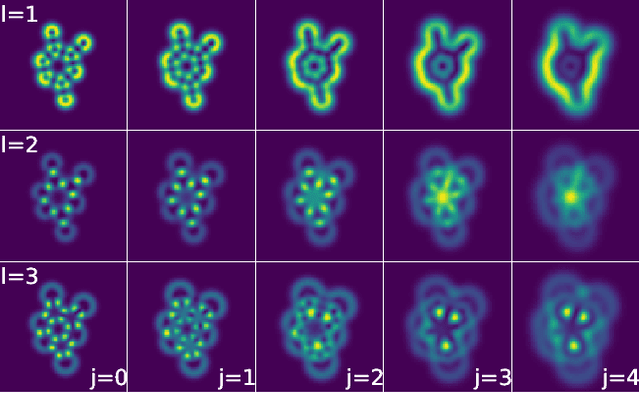
Abstract:We present a machine learning algorithm for the prediction of molecule properties inspired by ideas from density functional theory. Using Gaussian-type orbital functions, we create surrogate electronic densities of the molecule from which we compute invariant "solid harmonic scattering coefficients" that account for different types of interactions at different scales. Multi-linear regressions of various physical properties of molecules are computed from these invariant coefficients. Numerical experiments show that these regressions have near state of the art performance, even with relatively few training examples. Predictions over small sets of scattering coefficients can reach a DFT precision while being interpretable.
Truncated Variational Sampling for "Black Box" Optimization of Generative Models
Feb 22, 2018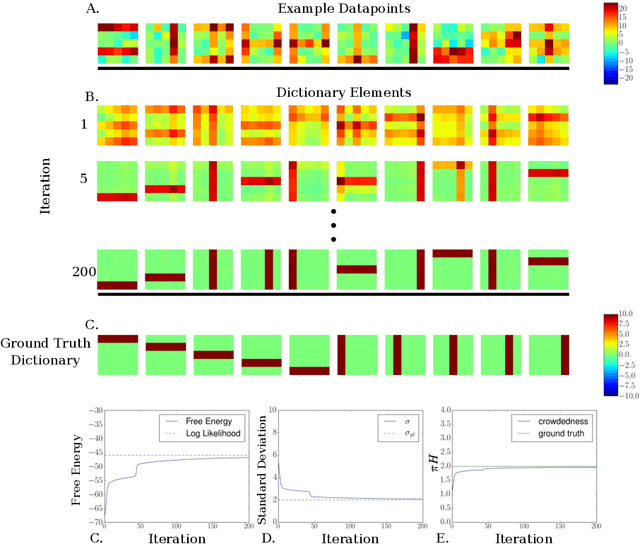
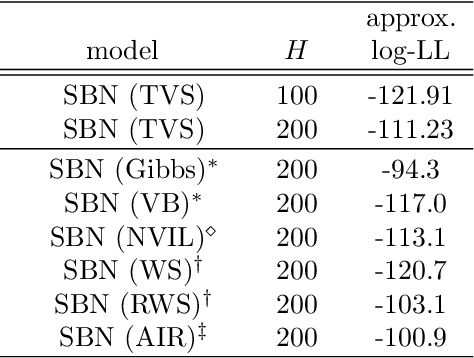
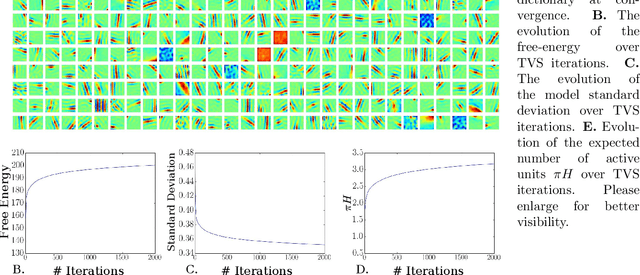
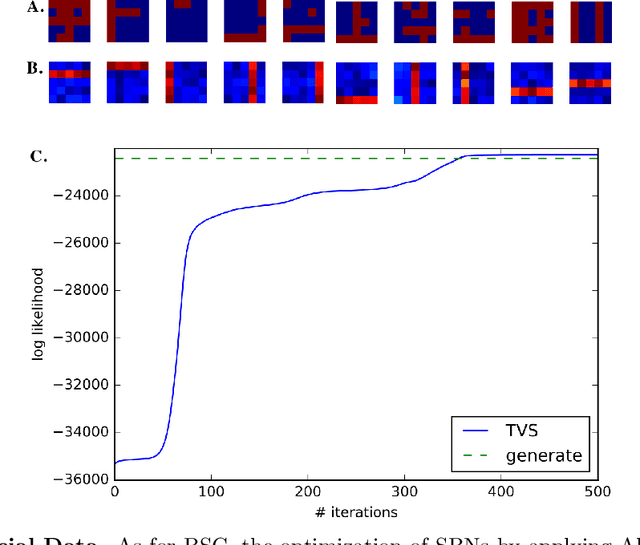
Abstract:We investigate the optimization of two probabilistic generative models with binary latent variables using a novel variational EM approach. The approach distinguishes itself from previous variational approaches by using latent states as variational parameters. Here we use efficient and general purpose sampling procedures to vary the latent states, and investigate the "black box" applicability of the resulting optimization procedure. For general purpose applicability, samples are drawn from approximate marginal distributions of the considered generative model as well as from the model's prior distribution. As such, variational sampling is defined in a generic form, and is directly executable for a given model. As a proof of concept, we then apply the novel procedure (A) to Binary Sparse Coding (a model with continuous observables), and (B) to basic Sigmoid Belief Networks (which are models with binary observables). Numerical experiments verify that the investigated approach efficiently as well as effectively increases a variational free energy objective without requiring any additional analytical steps.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge