Fatih Aksu
Virtual Scanning for NSCLC Histology: Investigating the Discriminatory Power of Synthetic PET
May 04, 2026Abstract:Accurate histological differentiation between adenocarcinoma (ADC) and squamous cell carcinoma (SCC) is critical for personalized treatment in non-small cell lung cancer (NSCLC). While [$^{18}$F]FDG PET/CT is a standard tool for the clinical evaluation of lung cancer, its utility is often limited by high costs and radiation exposure. In this paper, we investigate the feasibility of "virtual scanning" as a feature-enhancement strategy by evaluating whether synthetic PET data can provide complementary feature representations to supplement anatomical CT scans in histological subtype classification. We propose a framework that leverages a 3D Pix2Pix Generative Adversarial Network (GAN), pretrained on the FDG-PET/CT Lesions dataset, to synthesize pseudo-PET volumes from anatomical CT scans. These synthetic volumes are integrated with structural CT data within the MINT framework, a multi-stage intermediate fusion architecture. Our experiments, conducted on a multi-center dataset of 714 subjects, demonstrate that the inclusion of synthetic metabolic features significantly improves classification performance over a CT-only baseline. The multimodal approach achieved a statistically significant increase in the Area Under the Curve (AUC) from 0.489 to 0.591 and improved the Geometric Mean (GMean) from 0.305 to 0.524. These results suggest that synthetic PET scans provide discriminatory metabolic cues that enable deep learning models to exploit complementary cross-modal information, offering a potential feature-enhancement strategy for clinical scenarios where physical PET scans are unavailable.
Learning from Limited and Incomplete Data: A Multimodal Framework for Predicting Pathological Response in NSCLC
Mar 16, 2026Abstract:Major pathological response (pR) following neoadjuvant therapy is a clinically meaningful endpoint in non-small cell lung cancer, strongly associated with improved survival. However, accurate preoperative prediction of pR remains challenging, particularly in real-world clinical settings characterized by limited data availability and incomplete clinical profiles. In this study, we propose a multimodal deep learning framework designed to address these constraints by integrating foundation model-based CT feature extraction with a missing-aware architecture for clinical variables. This approach enables robust learning from small cohorts while explicitly modeling missing clinical information, without relying on conventional imputation strategies. A weighted fusion mechanism is employed to leverage the complementary contributions of imaging and clinical modalities, yielding a multimodal model that consistently outperforms both unimodal imaging and clinical baselines. These findings underscore the added value of integrating heterogeneous data sources and highlight the potential of multimodal, missing-aware systems to support pR prediction under realistic clinical conditions.
A Systematic Review of Intermediate Fusion in Multimodal Deep Learning for Biomedical Applications
Aug 02, 2024


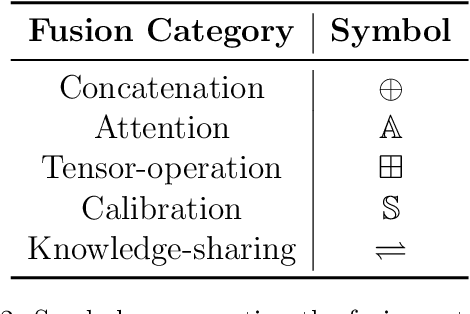
Abstract:Deep learning has revolutionized biomedical research by providing sophisticated methods to handle complex, high-dimensional data. Multimodal deep learning (MDL) further enhances this capability by integrating diverse data types such as imaging, textual data, and genetic information, leading to more robust and accurate predictive models. In MDL, differently from early and late fusion methods, intermediate fusion stands out for its ability to effectively combine modality-specific features during the learning process. This systematic review aims to comprehensively analyze and formalize current intermediate fusion methods in biomedical applications. We investigate the techniques employed, the challenges faced, and potential future directions for advancing intermediate fusion methods. Additionally, we introduce a structured notation to enhance the understanding and application of these methods beyond the biomedical domain. Our findings are intended to support researchers, healthcare professionals, and the broader deep learning community in developing more sophisticated and insightful multimodal models. Through this review, we aim to provide a foundational framework for future research and practical applications in the dynamic field of MDL.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge