Arturo Chiti
Virtual Scanning for NSCLC Histology: Investigating the Discriminatory Power of Synthetic PET
May 04, 2026Abstract:Accurate histological differentiation between adenocarcinoma (ADC) and squamous cell carcinoma (SCC) is critical for personalized treatment in non-small cell lung cancer (NSCLC). While [$^{18}$F]FDG PET/CT is a standard tool for the clinical evaluation of lung cancer, its utility is often limited by high costs and radiation exposure. In this paper, we investigate the feasibility of "virtual scanning" as a feature-enhancement strategy by evaluating whether synthetic PET data can provide complementary feature representations to supplement anatomical CT scans in histological subtype classification. We propose a framework that leverages a 3D Pix2Pix Generative Adversarial Network (GAN), pretrained on the FDG-PET/CT Lesions dataset, to synthesize pseudo-PET volumes from anatomical CT scans. These synthetic volumes are integrated with structural CT data within the MINT framework, a multi-stage intermediate fusion architecture. Our experiments, conducted on a multi-center dataset of 714 subjects, demonstrate that the inclusion of synthetic metabolic features significantly improves classification performance over a CT-only baseline. The multimodal approach achieved a statistically significant increase in the Area Under the Curve (AUC) from 0.489 to 0.591 and improved the Geometric Mean (GMean) from 0.305 to 0.524. These results suggest that synthetic PET scans provide discriminatory metabolic cues that enable deep learning models to exploit complementary cross-modal information, offering a potential feature-enhancement strategy for clinical scenarios where physical PET scans are unavailable.
Context-Gated Cross-Modal Perception with Visual Mamba for PET-CT Lung Tumor Segmentation
Oct 31, 2025Abstract:Accurate lung tumor segmentation is vital for improving diagnosis and treatment planning, and effectively combining anatomical and functional information from PET and CT remains a major challenge. In this study, we propose vMambaX, a lightweight multimodal framework integrating PET and CT scan images through a Context-Gated Cross-Modal Perception Module (CGM). Built on the Visual Mamba architecture, vMambaX adaptively enhances inter-modality feature interaction, emphasizing informative regions while suppressing noise. Evaluated on the PCLT20K dataset, the model outperforms baseline models while maintaining lower computational complexity. These results highlight the effectiveness of adaptive cross-modal gating for multimodal tumor segmentation and demonstrate the potential of vMambaX as an efficient and scalable framework for advanced lung cancer analysis. The code is available at https://github.com/arco-group/vMambaX.
Chest X-Rays Image Classification from beta-Variational Autoencoders Latent Features
Sep 29, 2021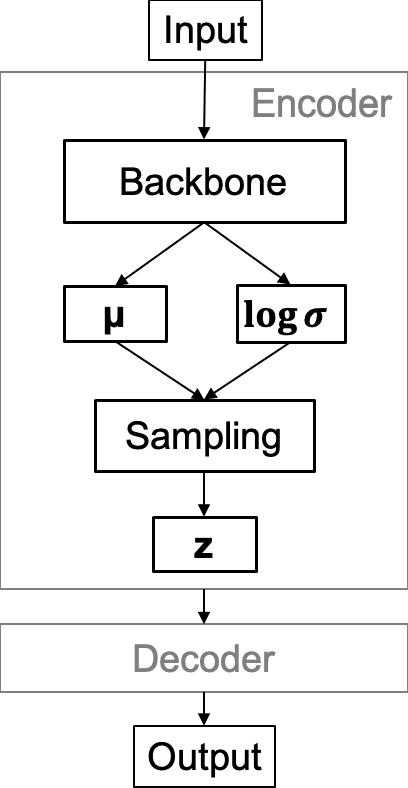
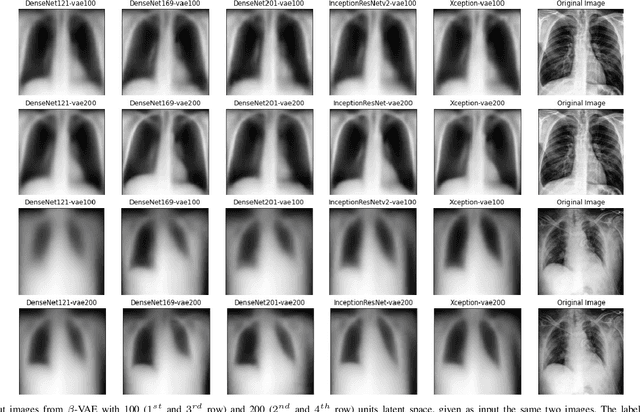
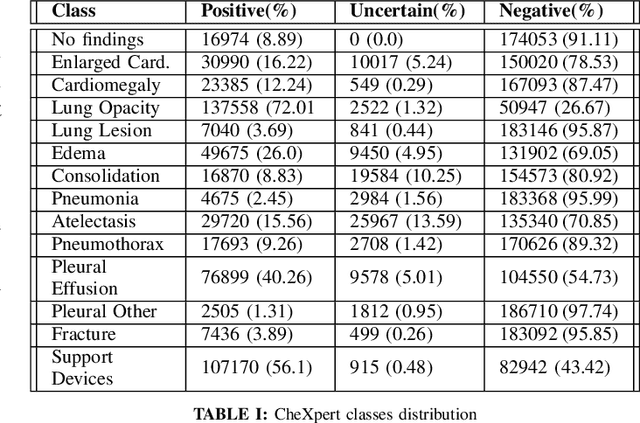
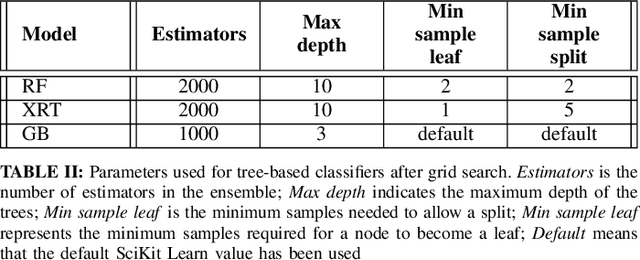
Abstract:Chest X-Ray (CXR) is one of the most common diagnostic techniques used in everyday clinical practice all around the world. We hereby present a work which intends to investigate and analyse the use of Deep Learning (DL) techniques to extract information from such images and allow to classify them, trying to keep our methodology as general as possible and possibly also usable in a real world scenario without much effort, in the future. To move in this direction, we trained several beta-Variational Autoencoder (beta-VAE) models on the CheXpert dataset, one of the largest publicly available collection of labeled CXR images; from these models, latent features have been extracted and used to train other Machine Learning models, able to classify the original images from the features extracted by the beta-VAE. Lastly, tree-based models have been combined together in ensemblings to improve the results without the necessity of further training or models engineering. Expecting some drop in pure performance with the respect to state of the art classification specific models, we obtained encouraging results, which show the viability of our approach and the usability of the high level features extracted by the autoencoders for classification tasks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge