Evangelos E. Papalexakis
Ensemble Node Embeddings using Tensor Decomposition: A Case-Study on DeepWalk
Aug 17, 2020
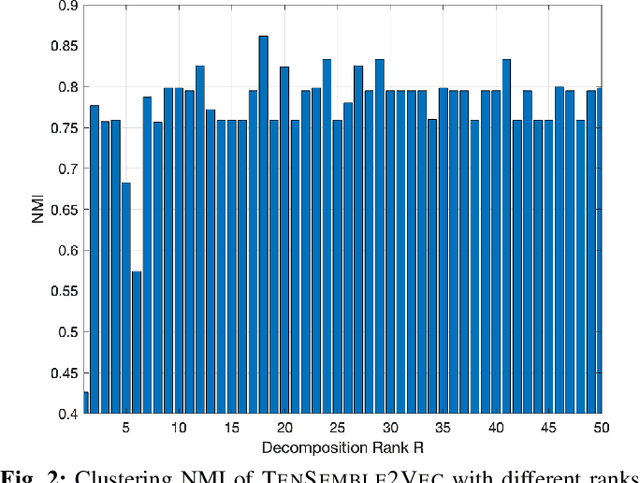
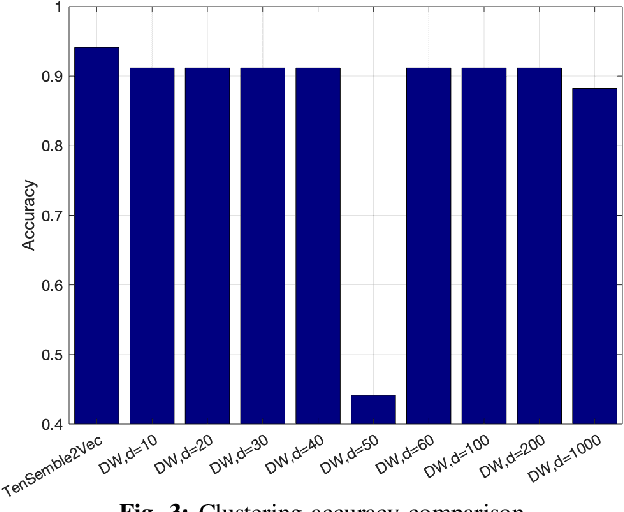
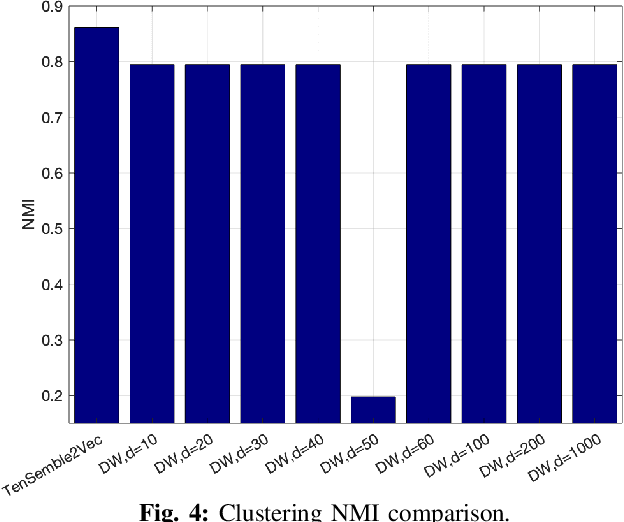
Abstract:Node embeddings have been attracting increasing attention during the past years. In this context, we propose a new ensemble node embedding approach, called TenSemble2Vec, by first generating multiple embeddings using the existing techniques and taking them as multiview data input of the state-of-art tensor decomposition model namely PARAFAC2 to learn the shared lower-dimensional representations of the nodes. Contrary to other embedding methods, our TenSemble2Vec takes advantage of the complementary information from different methods or the same method with different hyper-parameters, which bypasses the challenge of choosing models. Extensive tests using real-world data validates the efficiency of the proposed method.
HiJoD: Semi-Supervised Multi-aspect Detection of Misinformation using Hierarchical Joint Decomposition
May 08, 2020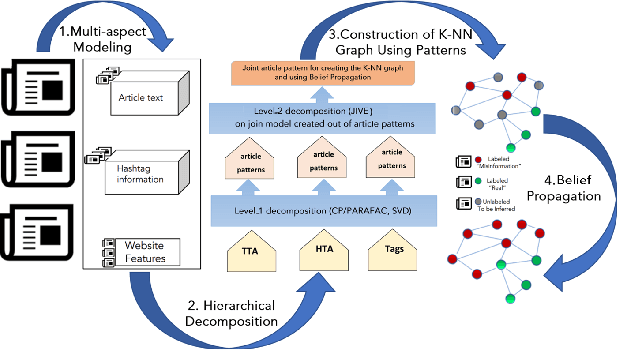
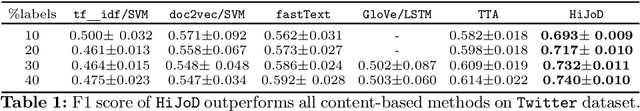
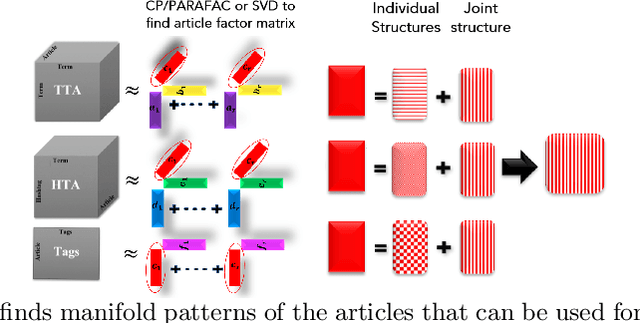

Abstract:Distinguishing between misinformation and real information is one of the most challenging problems in today's interconnected world. The vast majority of the state-of-the-art in detecting misinformation is fully supervised, requiring a large number of high-quality human annotations. However, the availability of such annotations cannot be taken for granted, since it is very costly, time-consuming, and challenging to do so in a way that keeps up with the proliferation of misinformation. In this work, we are interested in exploring scenarios where the number of annotations is limited. In such scenarios, we investigate how tapping on a diverse number of resources that characterize a news article, henceforth referred to as "aspects" can compensate for the lack of labels. In particular, our contributions in this paper are twofold: 1) We propose the use of three different aspects: article content, context of social sharing behaviors, and host website/domain features, and 2) We introduce a principled tensor based embedding framework that combines all those aspects effectively. We propose HiJoD a 2-level decomposition pipeline which not only outperforms state-of-the-art methods with F1-scores of 74% and 81% on Twitter and Politifact datasets respectively but also is an order of magnitude faster than similar ensemble approaches.
TensorShield: Tensor-based Defense Against Adversarial Attacks on Images
Feb 18, 2020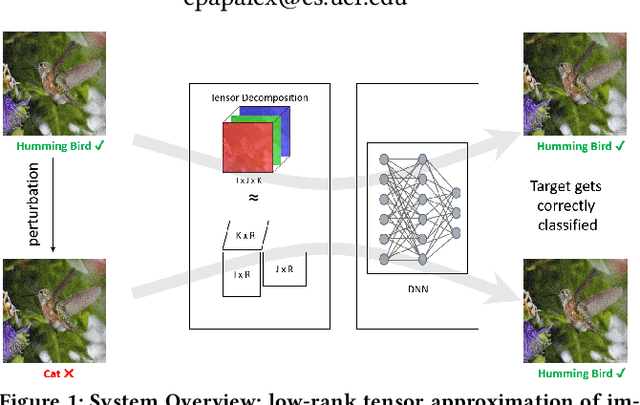
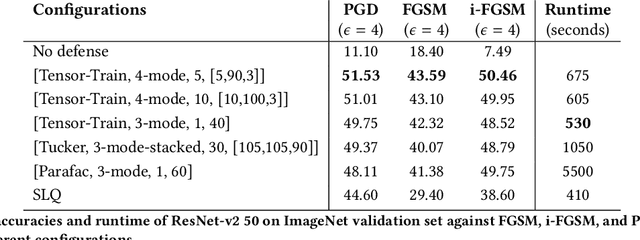
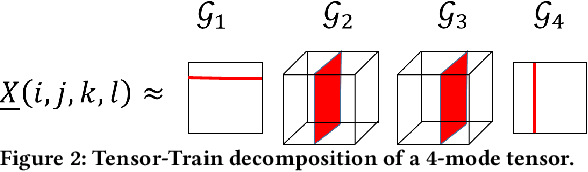
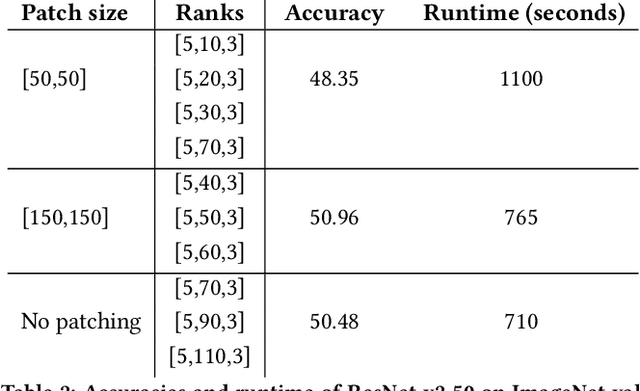
Abstract:Recent studies have demonstrated that machine learning approaches like deep neural networks (DNNs) are easily fooled by adversarial attacks. Subtle and imperceptible perturbations of the data are able to change the result of deep neural networks. Leveraging vulnerable machine learning methods raises many concerns especially in domains where security is an important factor. Therefore, it is crucial to design defense mechanisms against adversarial attacks. For the task of image classification, unnoticeable perturbations mostly occur in the high-frequency spectrum of the image. In this paper, we utilize tensor decomposition techniques as a preprocessing step to find a low-rank approximation of images which can significantly discard high-frequency perturbations. Recently a defense framework called Shield could "vaccinate" Convolutional Neural Networks (CNN) against adversarial examples by performing random-quality JPEG compressions on local patches of images on the ImageNet dataset. Our tensor-based defense mechanism outperforms the SLQ method from Shield by 14% against FastGradient Descent (FGSM) adversarial attacks, while maintaining comparable speed.
REST: A thread embedding approach for identifying and classifying user-specified information in security forums
Jan 08, 2020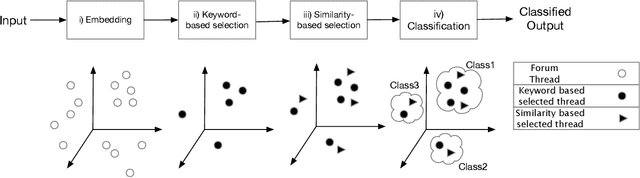
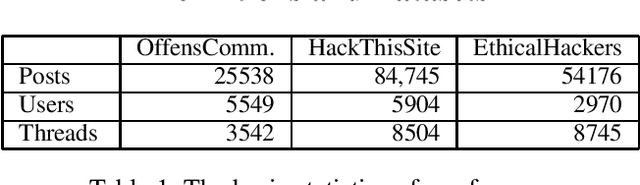
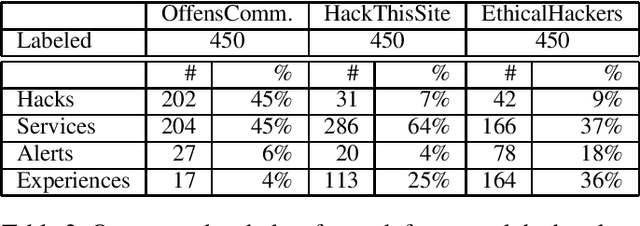
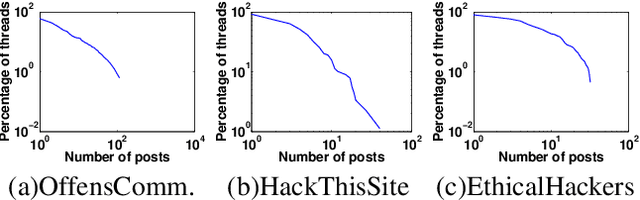
Abstract:How can we extract useful information from a security forum? We focus on identifying threads of interest to a security professional: (a) alerts of worrisome events, such as attacks, (b) offering of malicious services and products, (c) hacking information to perform malicious acts, and (d) useful security-related experiences. The analysis of security forums is in its infancy despite several promising recent works. Novel approaches are needed to address the challenges in this domain: (a) the difficulty in specifying the "topics" of interest efficiently, and (b) the unstructured and informal nature of the text. We propose, REST, a systematic methodology to: (a) identify threads of interest based on a, possibly incomplete, bag of words, and (b) classify them into one of the four classes above. The key novelty of the work is a multi-step weighted embedding approach: we project words, threads and classes in appropriate embedding spaces and establish relevance and similarity there. We evaluate our method with real data from three security forums with a total of 164k posts and 21K threads. First, REST robustness to initial keyword selection can extend the user-provided keyword set and thus, it can recover from missing keywords. Second, REST categorizes the threads into the classes of interest with superior accuracy compared to five other methods: REST exhibits an accuracy between 63.3-76.9%. We see our approach as a first step for harnessing the wealth of information of online forums in a user-friendly way, since the user can loosely specify her keywords of interest.
Adaptive Granularity in Tensors: A Quest for Interpretable Structure
Dec 19, 2019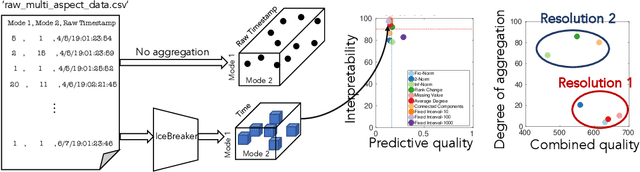
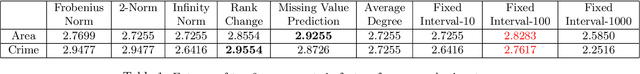
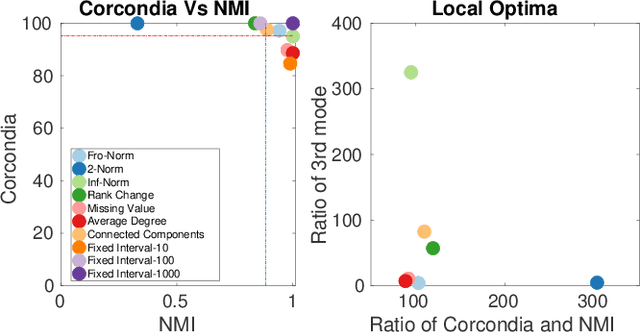
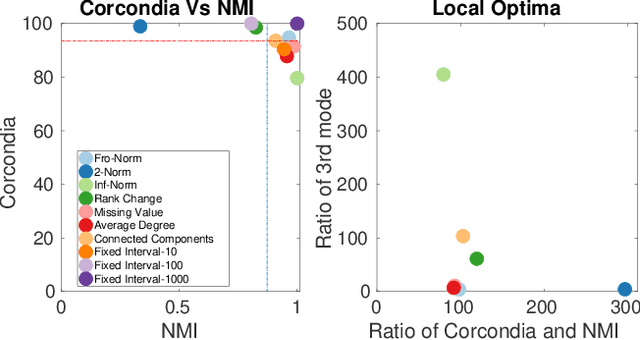
Abstract:Data collected at very frequent intervals is usually extremely sparse and has no structure that is exploitable by modern tensor decomposition algorithms. Thus the utility of such tensors is low, in terms of the amount of interpretable and exploitable structure that one can extract from them. In this paper, we introduce the problem of finding a tensor of adaptive aggregated granularity that can be decomposed to reveal meaningful latent concepts (structures) from datasets that, in their original form, are not amenable to tensor analysis. Such datasets fall under the broad category of sparse point processes that evolve over space and/or time. To the best of our knowledge, this is the first work that explores adaptive granularity aggregation in tensors. Furthermore, we formally define the problem and discuss what different definitions of "good structure" can be in practice, and show that optimal solution is of prohibitive combinatorial complexity. Subsequently, we propose an efficient and effective greedy algorithm which follows a number of intuitive decision criteria that locally maximize the "goodness of structure", resulting in high-quality tensors. We evaluate our method on both semi-synthetic data where ground truth is known and real datasets for which we do not have any ground truth. In both cases, our proposed method constructs tensors that have very high structure quality. Finally, our proposed method is able to discover different natural resolutions of a multi-aspect dataset, which can lead to multi-resolution analysis.
The core consistency of a compressed tensor
Nov 18, 2018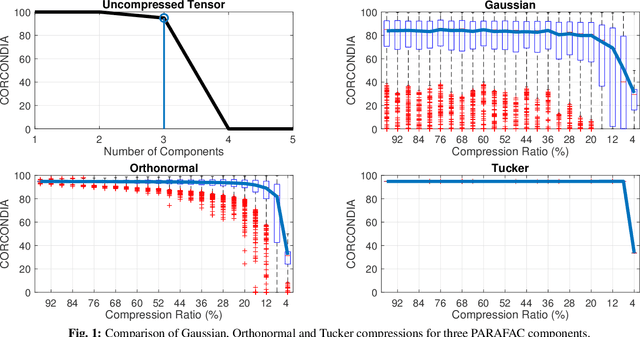
Abstract:Tensor decomposition on big data has attracted significant attention recently. Among the most popular methods is a class of algorithms that leverages compression in order to reduce the size of the tensor and potentially parallelize computations. A fundamental requirement for such methods to work properly is that the low-rank tensor structure is retained upon compression. In lieu of efficient and realistic means of computing and studying the effects of compression on the low rank of a tensor, we study the effects of compression on the core consistency; a widely used heuristic that has been used as a proxy for estimating that low rank. We provide theoretical analysis, where we identify sufficient conditions for the compression such that the core consistency is preserved, and we conduct extensive experiments that validate our analysis. Further, we explore popular compression schemes and how they affect the core consistency.
Representation Learning by Reconstructing Neighborhoods
Nov 06, 2018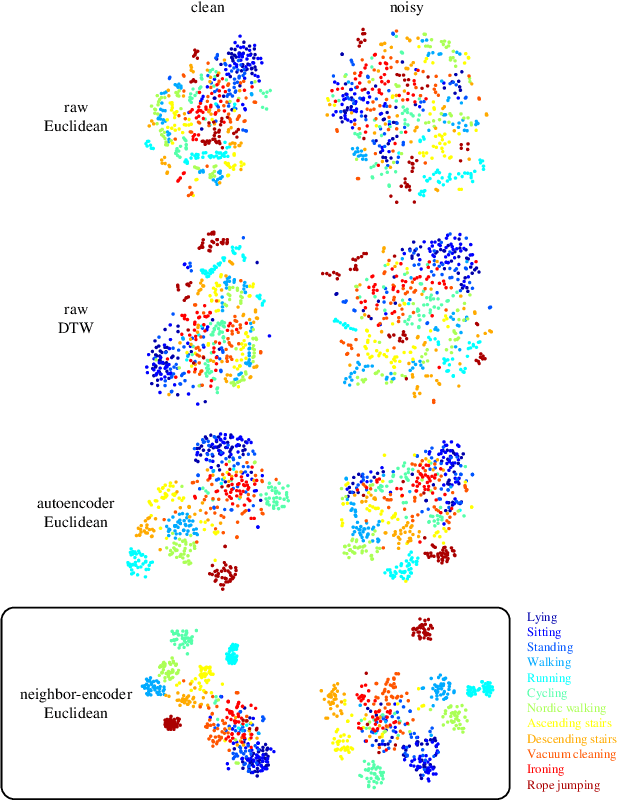
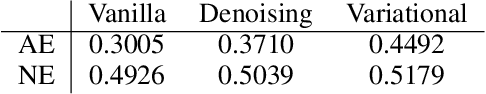
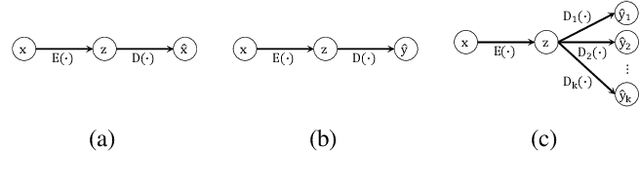
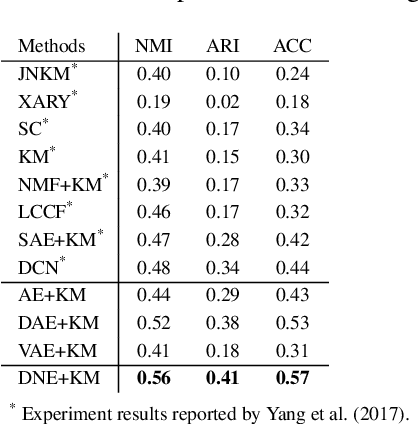
Abstract:Since its introduction, unsupervised representation learning has attracted a lot of attention from the research community, as it is demonstrated to be highly effective and easy-to-apply in tasks such as dimension reduction, clustering, visualization, information retrieval, and semi-supervised learning. In this work, we propose a novel unsupervised representation learning framework called neighbor-encoder, in which domain knowledge can be easily incorporated into the learning process without modifying the general encoder-decoder architecture of the classic autoencoder.In contrast to autoencoder, which reconstructs the input data itself, neighbor-encoder reconstructs the input data's neighbors. As the proposed representation learning problem is essentially a neighbor reconstruction problem, domain knowledge can be easily incorporated in the form of an appropriate definition of similarity between objects. Based on that observation, our framework can leverage any off-the-shelf similarity search algorithms or side information to find the neighbor of an input object. Applications of other algorithms (e.g., association rule mining) in our framework are also possible, given that the appropriate definition of neighbor can vary in different contexts. We have demonstrated the effectiveness of our framework in many diverse domains, including images, text, and time series, and for various data mining tasks including classification, clustering, and visualization. Experimental results show that neighbor-encoder not only outperforms autoencoder in most of the scenarios we consider, but also achieves the state-of-the-art performance on text document clustering.
RECS: Robust Graph Embedding Using Connection Subgraphs
Sep 05, 2018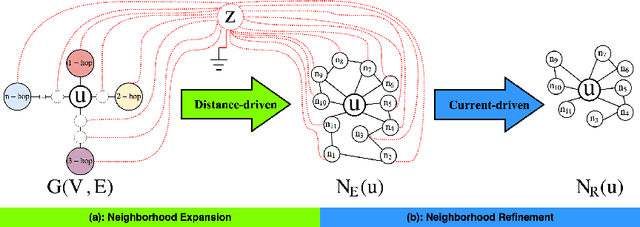
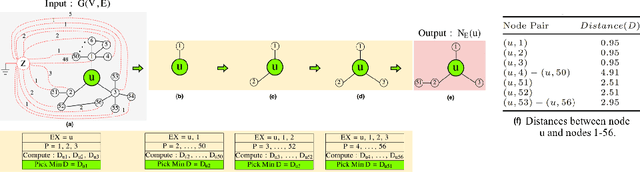
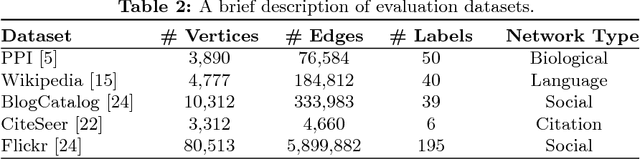
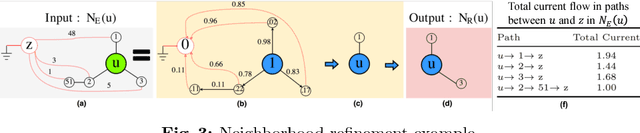
Abstract:The success of graph embeddings or node representation learning in a variety of downstream tasks, such as node classification, link prediction, and recommendation systems, has led to their popularity in recent years. Representation learning algorithms aim to preserve local and global network structure by identifying node neighborhood notions. However, many existing algorithms generate embeddings that fail to properly preserve the network structure, or lead to unstable representations due to random processes (e.g., random walks to generate context) and, thus, cannot generate to multi-graph problems. In this paper, we propose RECS, a novel, stable graph embedding algorithmic framework. RECS learns graph representations using connection subgraphs by employing the analogy of graphs with electrical circuits. It preserves both local and global connectivity patterns, and addresses the issue of high-degree nodes. Further, it exploits the strength of weak ties and meta-data that have been neglected by baselines. The experiments show that RECS outperforms state-of-the-art algorithms by up to 36.85% on multi-label classification problem. Further, in contrast to baselines, RECS, being deterministic, is completely stable.
COPA: Constrained PARAFAC2 for Sparse & Large Datasets
Aug 27, 2018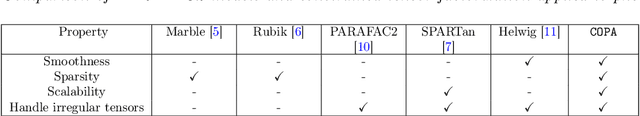
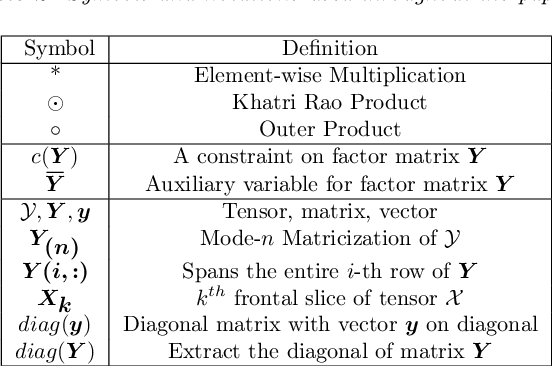
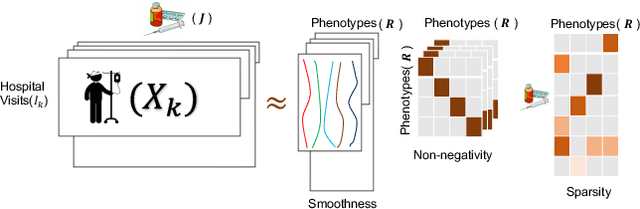

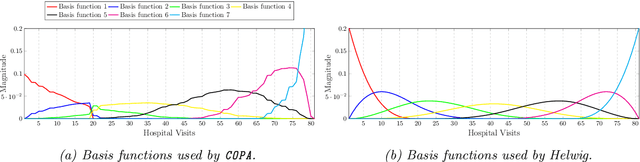
Abstract:PARAFAC2 has demonstrated success in modeling irregular tensors, where the tensor dimensions vary across one of the modes. An example scenario is modeling treatments across a set of patients with the varying number of medical encounters over time. Despite recent improvements on unconstrained PARAFAC2, its model factors are usually dense and sensitive to noise which limits their interpretability. As a result, the following open challenges remain: a) various modeling constraints, such as temporal smoothness, sparsity and non-negativity, are needed to be imposed for interpretable temporal modeling and b) a scalable approach is required to support those constraints efficiently for large datasets. To tackle these challenges, we propose a {\it CO}nstrained {\it PA}RAFAC2 (COPA) method, which carefully incorporates optimization constraints such as temporal smoothness, sparsity, and non-negativity in the resulting factors. To efficiently support all those constraints, COPA adopts a hybrid optimization framework using alternating optimization and alternating direction method of multiplier (AO-ADMM). As evaluated on large electronic health record (EHR) datasets with hundreds of thousands of patients, COPA achieves significant speedups (up to 36 times faster) over prior PARAFAC2 approaches that only attempt to handle a subset of the constraints that COPA enables. Overall, our method outperforms all the baselines attempting to handle a subset of the constraints in terms of speed, while achieving the same level of accuracy. Through a case study on temporal phenotyping of medically complex children, we demonstrate how the constraints imposed by COPA reveal concise phenotypes and meaningful temporal profiles of patients. The clinical interpretation of both the phenotypes and the temporal profiles was confirmed by a medical expert.
Webly Supervised Joint Embedding for Cross-Modal Image-Text Retrieval
Aug 23, 2018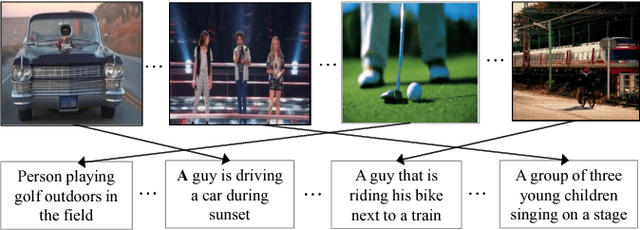
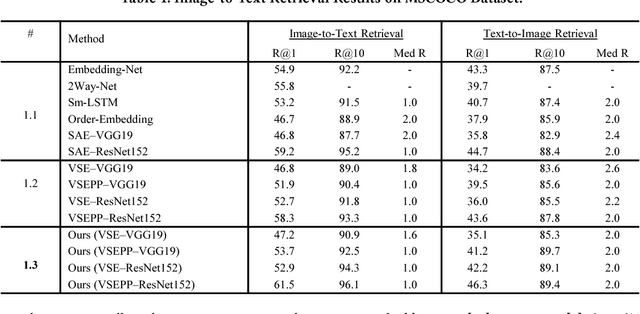
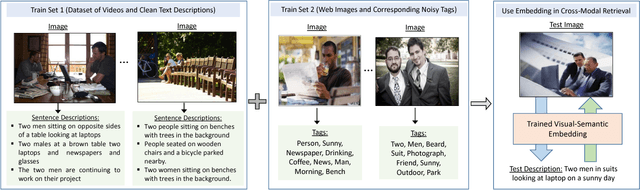
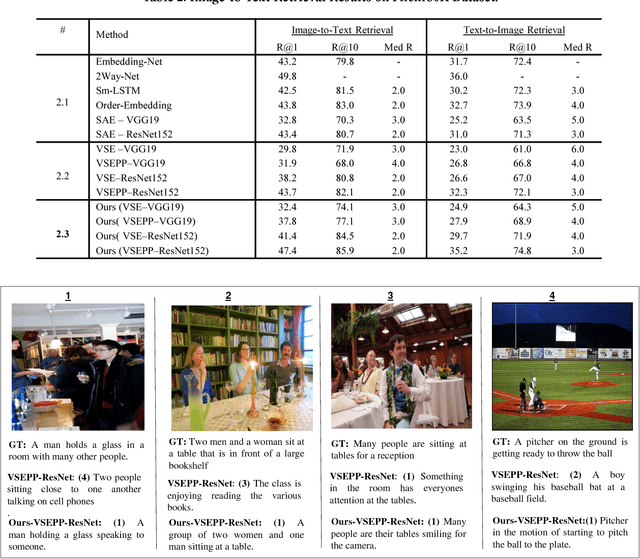
Abstract:Cross-modal retrieval between visual data and natural language description remains a long-standing challenge in multimedia. While recent image-text retrieval methods offer great promise by learning deep representations aligned across modalities, most of these methods are plagued by the issue of training with small-scale datasets covering a limited number of images with ground-truth sentences. Moreover, it is extremely expensive to create a larger dataset by annotating millions of images with sentences and may lead to a biased model. Inspired by the recent success of webly supervised learning in deep neural networks, we capitalize on readily-available web images with noisy annotations to learn robust image-text joint representation. Specifically, our main idea is to leverage web images and corresponding tags, along with fully annotated datasets, in training for learning the visual-semantic joint embedding. We propose a two-stage approach for the task that can augment a typical supervised pair-wise ranking loss based formulation with weakly-annotated web images to learn a more robust visual-semantic embedding. Experiments on two standard benchmark datasets demonstrate that our method achieves a significant performance gain in image-text retrieval compared to state-of-the-art approaches.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge