Ender Konukoglu
for the Alzheimer's Disease Neuroimaging Initiative
FedFA: Federated Feature Augmentation
Jan 30, 2023Abstract:Federated learning is a distributed paradigm that allows multiple parties to collaboratively train deep models without exchanging the raw data. However, the data distribution among clients is naturally non-i.i.d., which leads to severe degradation of the learnt model. The primary goal of this paper is to develop a robust federated learning algorithm to address feature shift in clients' samples, which can be caused by various factors, e.g., acquisition differences in medical imaging. To reach this goal, we propose FedFA to tackle federated learning from a distinct perspective of federated feature augmentation. FedFA is based on a major insight that each client's data distribution can be characterized by statistics (i.e., mean and standard deviation) of latent features; and it is likely to manipulate these local statistics globally, i.e., based on information in the entire federation, to let clients have a better sense of the underlying distribution and therefore alleviate local data bias. Based on this insight, we propose to augment each local feature statistic probabilistically based on a normal distribution, whose mean is the original statistic and variance quantifies the augmentation scope. Key to our approach is the determination of a meaningful Gaussian variance, which is accomplished by taking into account not only biased data of each individual client, but also underlying feature statistics characterized by all participating clients. We offer both theoretical and empirical justifications to verify the effectiveness of FedFA. Our code is available at https://github.com/tfzhou/FedFA.
Unsupervised Superpixel Generation using Edge-Sparse Embedding
Nov 29, 2022



Abstract:Partitioning an image into superpixels based on the similarity of pixels with respect to features such as colour or spatial location can significantly reduce data complexity and improve subsequent image processing tasks. Initial algorithms for unsupervised superpixel generation solely relied on local cues without prioritizing significant edges over arbitrary ones. On the other hand, more recent methods based on unsupervised deep learning either fail to properly address the trade-off between superpixel edge adherence and compactness or lack control over the generated number of superpixels. By using random images with strong spatial correlation as input, \ie, blurred noise images, in a non-convolutional image decoder we can reduce the expected number of contrasts and enforce smooth, connected edges in the reconstructed image. We generate edge-sparse pixel embeddings by encoding additional spatial information into the piece-wise smooth activation maps from the decoder's last hidden layer and use a standard clustering algorithm to extract high quality superpixels. Our proposed method reaches state-of-the-art performance on the BSDS500, PASCAL-Context and a microscopy dataset.
Neural Vector Fields for Surface Representation and Inference
Apr 13, 2022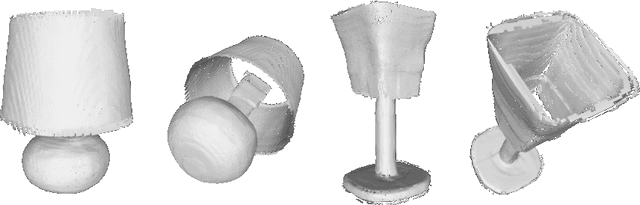
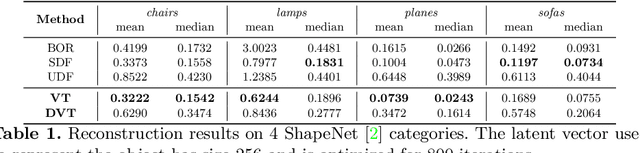
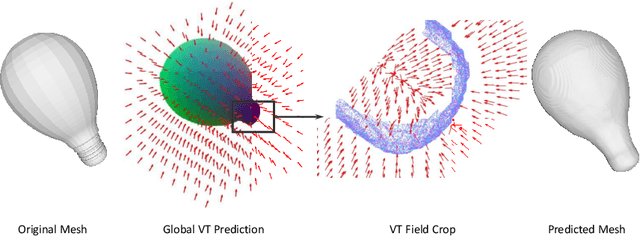
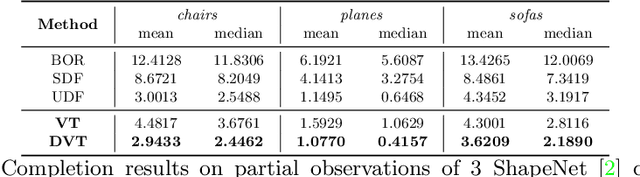
Abstract:Neural implicit fields have recently been shown to represent 3D shapes accurately, opening up various applications in 3D shape analysis. Up to now, such implicit fields for 3D representation are scalar, encoding the signed distance or binary volume occupancy and more recently the unsigned distance. However, the first two can only represent closed shapes, while the unsigned distance has difficulties in accurate and fast shape inference. In this paper, we propose a Neural Vector Field for shape representation in order to overcome the two aforementioned problems. Mapping each point in space to the direction towards the closest surface, we can represent any type of shape. Similarly the shape mesh can be reconstructed by applying the marching cubes algorithm, with proposed small changes, on top of the inferred vector field. We compare the method on ShapeNet where the proposed new neural implicit field shows superior accuracy in representing both closed and open shapes outperforming previous methods.
Rethinking Semantic Segmentation: A Prototype View
Apr 04, 2022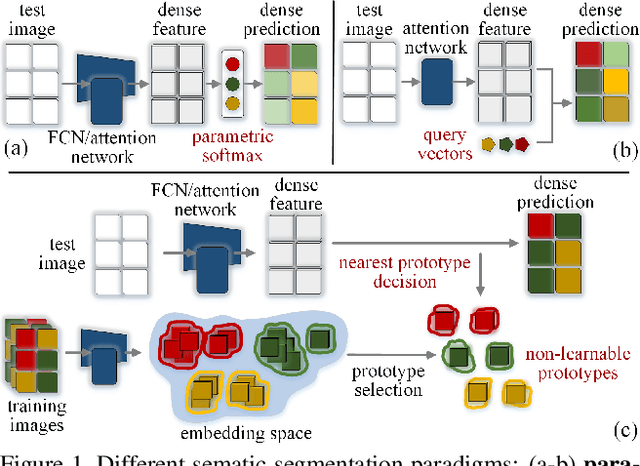
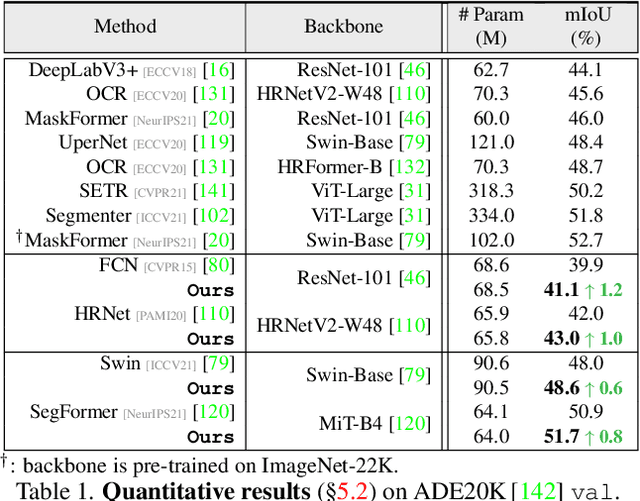
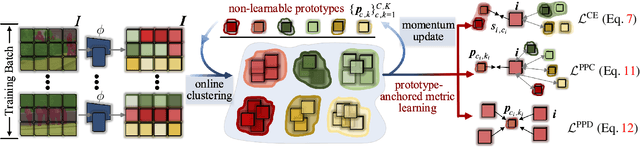
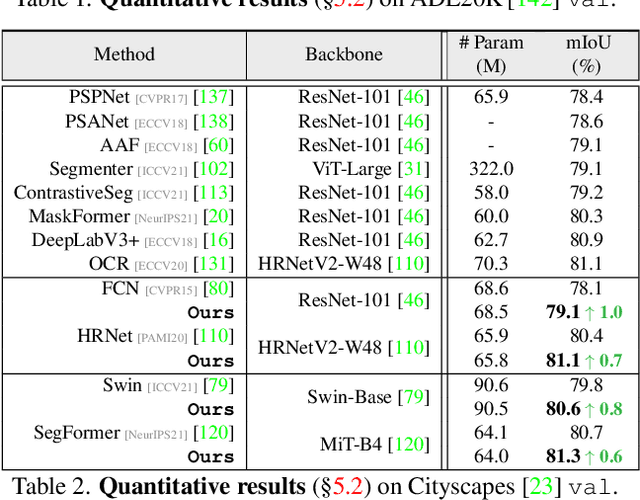
Abstract:Prevalent semantic segmentation solutions, despite their different network designs (FCN based or attention based) and mask decoding strategies (parametric softmax based or pixel-query based), can be placed in one category, by considering the softmax weights or query vectors as learnable class prototypes. In light of this prototype view, this study uncovers several limitations of such parametric segmentation regime, and proposes a nonparametric alternative based on non-learnable prototypes. Instead of prior methods learning a single weight/query vector for each class in a fully parametric manner, our model represents each class as a set of non-learnable prototypes, relying solely on the mean features of several training pixels within that class. The dense prediction is thus achieved by nonparametric nearest prototype retrieving. This allows our model to directly shape the pixel embedding space, by optimizing the arrangement between embedded pixels and anchored prototypes. It is able to handle arbitrary number of classes with a constant amount of learnable parameters. We empirically show that, with FCN based and attention based segmentation models (i.e., HRNet, Swin, SegFormer) and backbones (i.e., ResNet, HRNet, Swin, MiT), our nonparametric framework yields compelling results over several datasets (i.e., ADE20K, Cityscapes, COCO-Stuff), and performs well in the large-vocabulary situation. We expect this work will provoke a rethink of the current de facto semantic segmentation model design.
Zero Pixel Directional Boundary by Vector Transform
Mar 16, 2022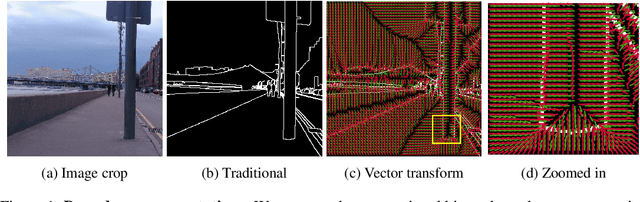
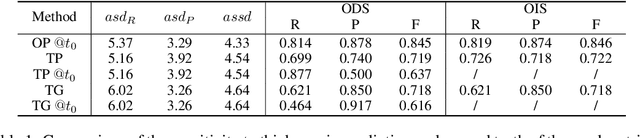
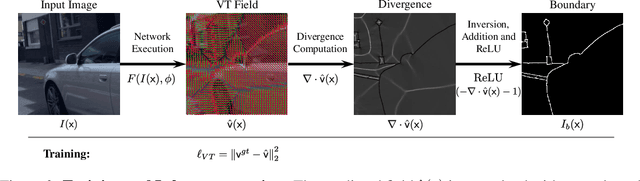
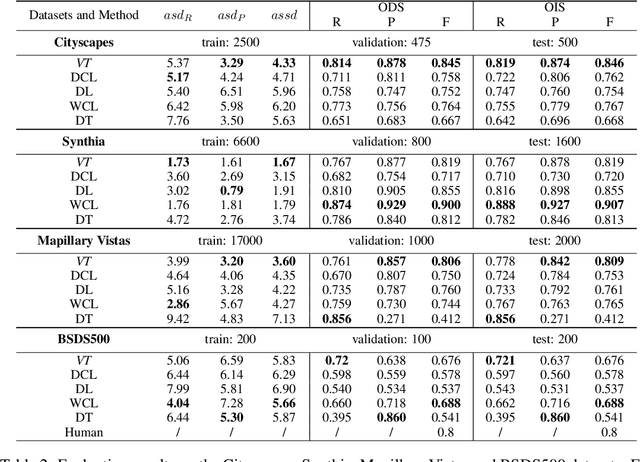
Abstract:Boundaries are among the primary visual cues used by human and computer vision systems. One of the key problems in boundary detection is the label representation, which typically leads to class imbalance and, as a consequence, to thick boundaries that require non-differential post-processing steps to be thinned. In this paper, we re-interpret boundaries as 1-D surfaces and formulate a one-to-one vector transform function that allows for training of boundary prediction completely avoiding the class imbalance issue. Specifically, we define the boundary representation at any point as the unit vector pointing to the closest boundary surface. Our problem formulation leads to the estimation of direction as well as richer contextual information of the boundary, and, if desired, the availability of zero-pixel thin boundaries also at training time. Our method uses no hyper-parameter in the training loss and a fixed stable hyper-parameter at inference. We provide theoretical justification/discussions of the vector transform representation. We evaluate the proposed loss method using a standard architecture and show the excellent performance over other losses and representations on several datasets.
A Field of Experts Prior for Adapting Neural Networks at Test Time
Feb 10, 2022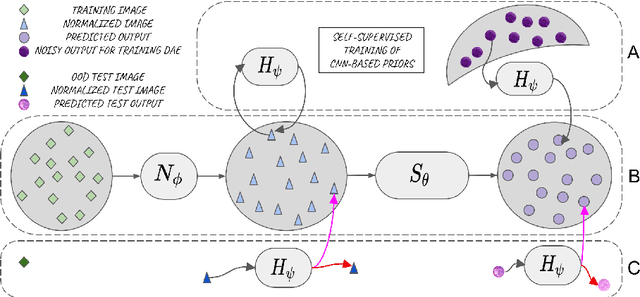
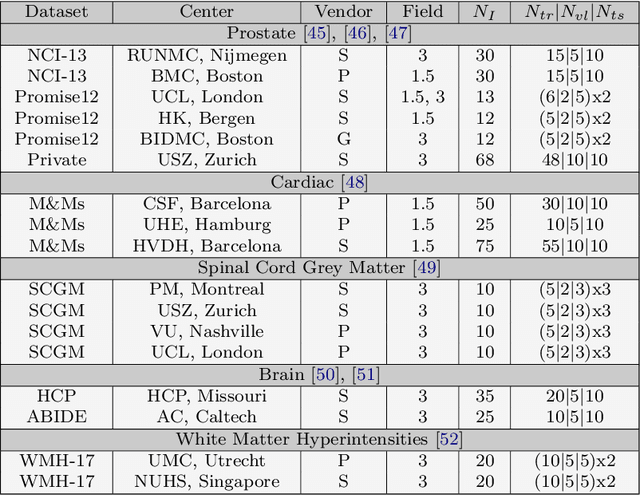
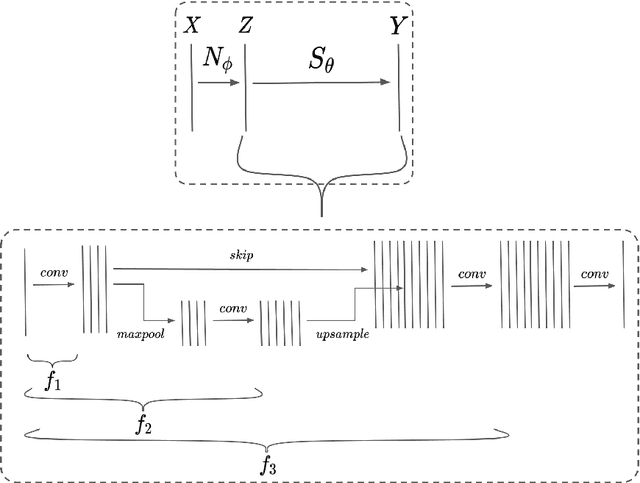
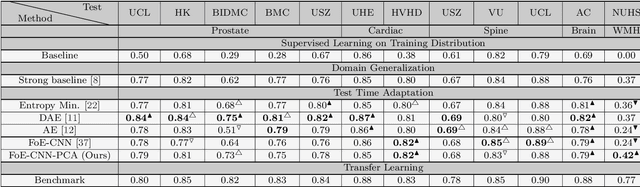
Abstract:Performance of convolutional neural networks (CNNs) in image analysis tasks is often marred in the presence of acquisition-related distribution shifts between training and test images. Recently, it has been proposed to tackle this problem by fine-tuning trained CNNs for each test image. Such test-time-adaptation (TTA) is a promising and practical strategy for improving robustness to distribution shifts as it requires neither data sharing between institutions nor annotating additional data. Previous TTA methods use a helper model to increase similarity between outputs and/or features extracted from a test image with those of the training images. Such helpers, which are typically modeled using CNNs, can be task-specific and themselves vulnerable to distribution shifts in their inputs. To overcome these problems, we propose to carry out TTA by matching the feature distributions of test and training images, as modelled by a field-of-experts (FoE) prior. FoEs model complicated probability distributions as products of many simpler expert distributions. We use 1D marginal distributions of a trained task CNN's features as experts in the FoE model. Further, we compute principal components of patches of the task CNN's features, and consider the distributions of PCA loadings as additional experts. We validate the method on 5 MRI segmentation tasks (healthy tissues in 4 anatomical regions and lesions in 1 one anatomy), using data from 17 clinics, and on a MRI registration task, using data from 3 clinics. We find that the proposed FoE-based TTA is generically applicable in multiple tasks, and outperforms all previous TTA methods for lesion segmentation. For healthy tissue segmentation, the proposed method outperforms other task-agnostic methods, but a previous TTA method which is specifically designed for segmentation performs the best for most of the tested datasets. Our code is publicly available.
Wiener Guided DIP for Unsupervised Blind Image Deconvolution
Dec 19, 2021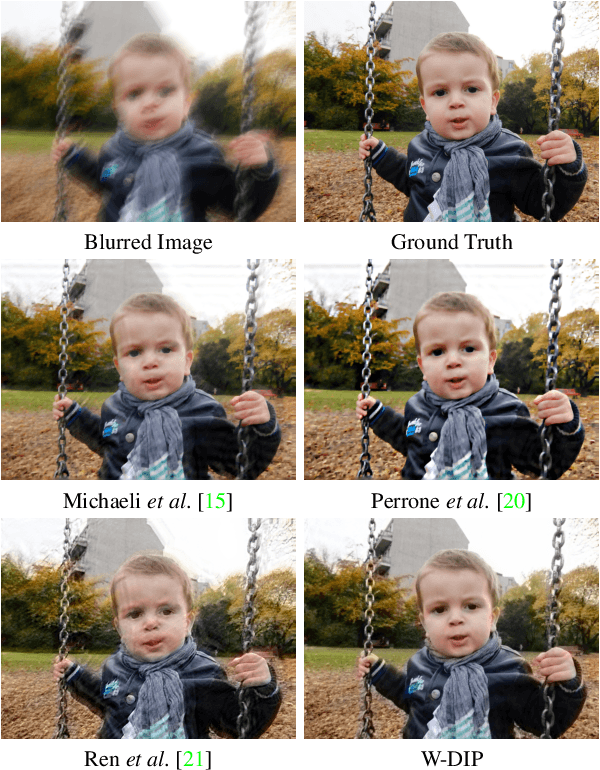
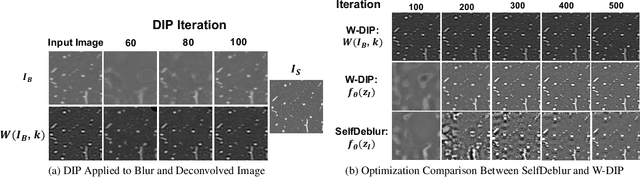
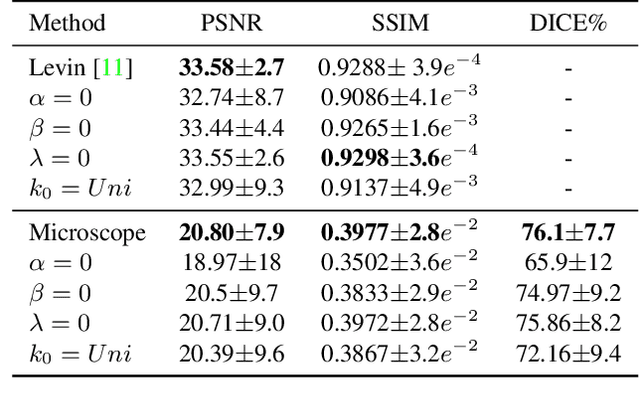

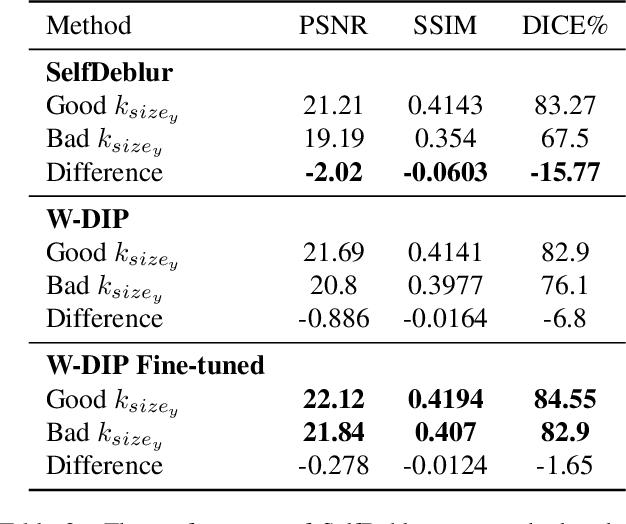
Abstract:Blind deconvolution is an ill-posed problem arising in various fields ranging from microscopy to astronomy. The ill-posed nature of the problem requires adequate priors to arrive to a desirable solution. Recently, it has been shown that deep learning architectures can serve as an image generation prior during unsupervised blind deconvolution optimization, however often exhibiting a performance fluctuation even on a single image. We propose to use Wiener-deconvolution to guide the image generator during optimization by providing it a sharpened version of the blurry image using an auxiliary kernel estimate starting from a Gaussian. We observe that the high-frequency artifacts of deconvolution are reproduced with a delay compared to low-frequency features. In addition, the image generator reproduces low-frequency features of the deconvolved image faster than that of a blurry image. We embed the computational process in a constrained optimization framework and show that the proposed method yields higher stability and performance across multiple datasets. In addition, we provide the code.
Local contrastive loss with pseudo-label based self-training for semi-supervised medical image segmentation
Dec 17, 2021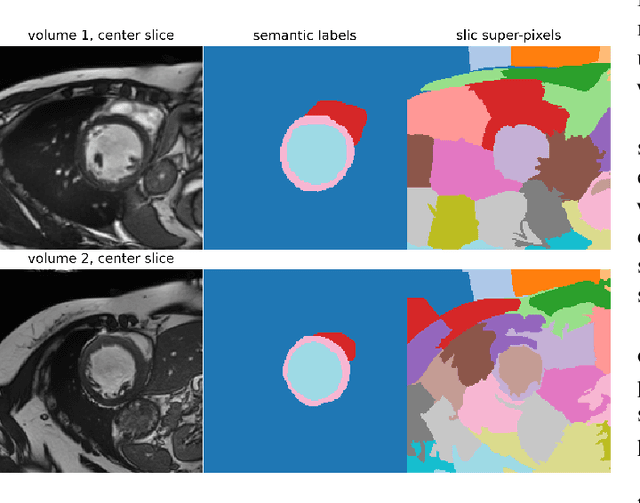
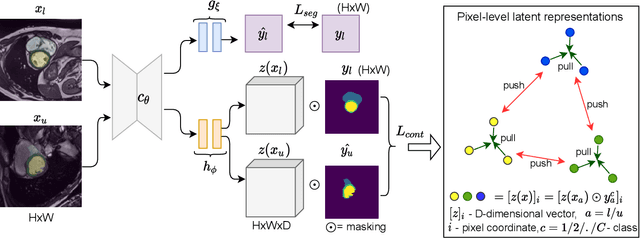
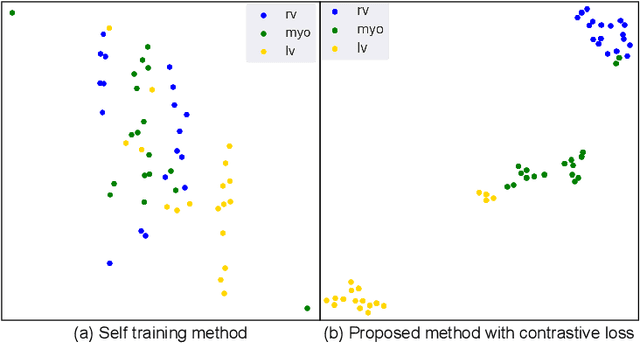
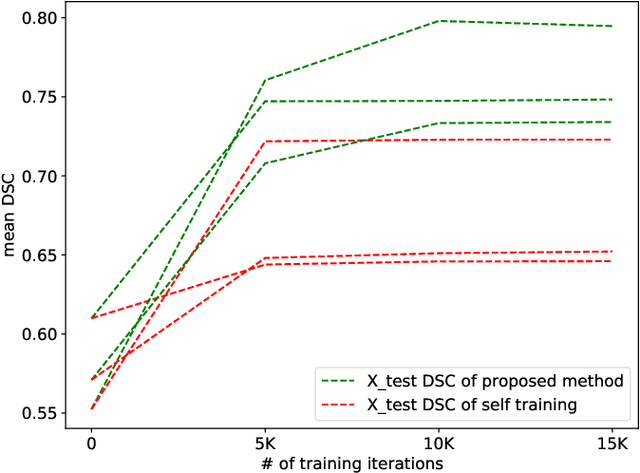
Abstract:Supervised deep learning-based methods yield accurate results for medical image segmentation. However, they require large labeled datasets for this, and obtaining them is a laborious task that requires clinical expertise. Semi/self-supervised learning-based approaches address this limitation by exploiting unlabeled data along with limited annotated data. Recent self-supervised learning methods use contrastive loss to learn good global level representations from unlabeled images and achieve high performance in classification tasks on popular natural image datasets like ImageNet. In pixel-level prediction tasks such as segmentation, it is crucial to also learn good local level representations along with global representations to achieve better accuracy. However, the impact of the existing local contrastive loss-based methods remains limited for learning good local representations because similar and dissimilar local regions are defined based on random augmentations and spatial proximity; not based on the semantic label of local regions due to lack of large-scale expert annotations in the semi/self-supervised setting. In this paper, we propose a local contrastive loss to learn good pixel level features useful for segmentation by exploiting semantic label information obtained from pseudo-labels of unlabeled images alongside limited annotated images. In particular, we define the proposed loss to encourage similar representations for the pixels that have the same pseudo-label/ label while being dissimilar to the representation of pixels with different pseudo-label/label in the dataset. We perform pseudo-label based self-training and train the network by jointly optimizing the proposed contrastive loss on both labeled and unlabeled sets and segmentation loss on only the limited labeled set. We evaluated on three public cardiac and prostate datasets, and obtain high segmentation performance.
ISNAS-DIP: Image-Specific Neural Architecture Search for Deep Image Prior
Nov 27, 2021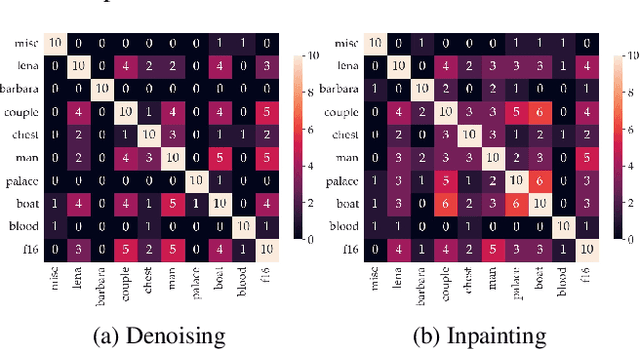
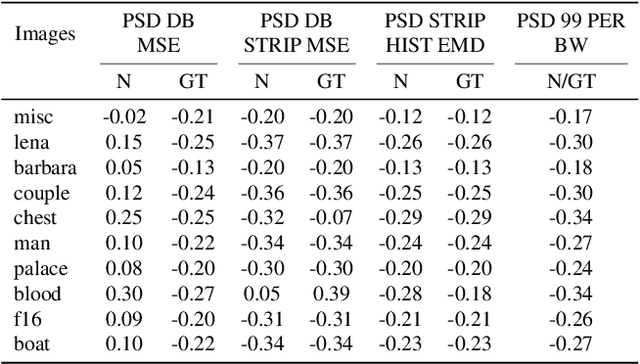
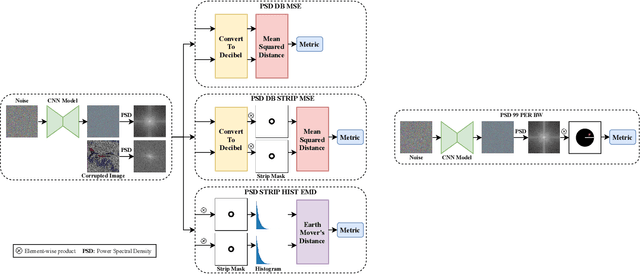
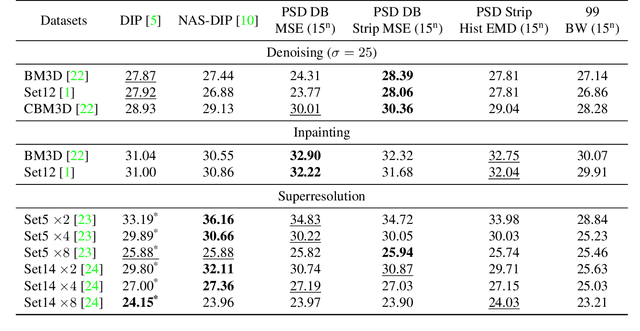
Abstract:Recent works show that convolutional neural network (CNN) architectures have a spectral bias towards lower frequencies, which has been leveraged for various image restoration tasks in the Deep Image Prior (DIP) framework. The benefit of the inductive bias the network imposes in the DIP framework depends on the architecture. Therefore, researchers have studied how to automate the search to determine the best-performing model. However, common neural architecture search (NAS) techniques are resource and time-intensive. Moreover, best-performing models are determined for a whole dataset of images instead of for each image independently, which would be prohibitively expensive. In this work, we first show that optimal neural architectures in the DIP framework are image-dependent. Leveraging this insight, we then propose an image-specific NAS strategy for the DIP framework that requires substantially less training than typical NAS approaches, effectively enabling image-specific NAS. For a given image, noise is fed to a large set of untrained CNNs, and their outputs' power spectral densities (PSD) are compared to that of the corrupted image using various metrics. Based on this, a small cohort of image-specific architectures is chosen and trained to reconstruct the corrupted image. Among this cohort, the model whose reconstruction is closest to the average of the reconstructed images is chosen as the final model. We justify the proposed strategy's effectiveness by (1) demonstrating its performance on a NAS Dataset for DIP that includes 500+ models from a particular search space (2) conducting extensive experiments on image denoising, inpainting, and super-resolution tasks. Our experiments show that image-specific metrics can reduce the search space to a small cohort of models, of which the best model outperforms current NAS approaches for image restoration.
Quality-Aware Memory Network for Interactive Volumetric Image Segmentation
Jul 05, 2021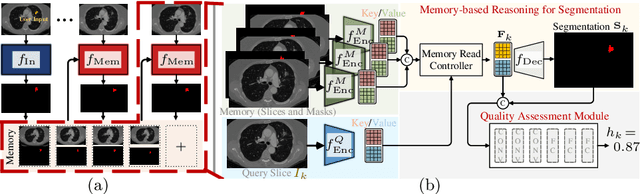
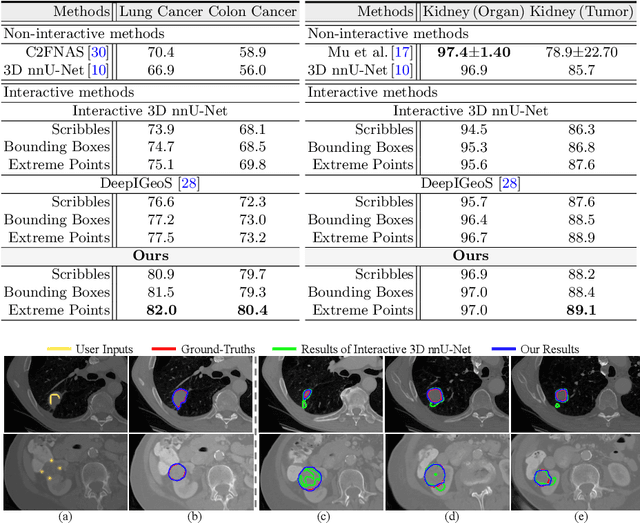
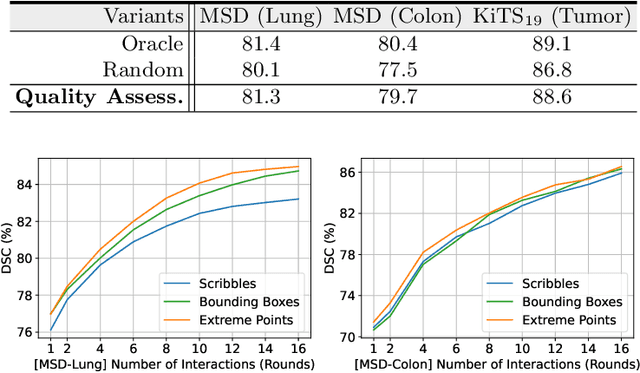
Abstract:Despite recent progress of automatic medical image segmentation techniques, fully automatic results usually fail to meet the clinical use and typically require further refinement. In this work, we propose a quality-aware memory network for interactive segmentation of 3D medical images. Provided by user guidance on an arbitrary slice, an interaction network is firstly employed to obtain an initial 2D segmentation. The quality-aware memory network subsequently propagates the initial segmentation estimation bidirectionally over the entire volume. Subsequent refinement based on additional user guidance on other slices can be incorporated in the same manner. To further facilitate interactive segmentation, a quality assessment module is introduced to suggest the next slice to segment based on the current segmentation quality of each slice. The proposed network has two appealing characteristics: 1) The memory-augmented network offers the ability to quickly encode past segmentation information, which will be retrieved for the segmentation of other slices; 2) The quality assessment module enables the model to directly estimate the qualities of segmentation predictions, which allows an active learning paradigm where users preferentially label the lowest-quality slice for multi-round refinement. The proposed network leads to a robust interactive segmentation engine, which can generalize well to various types of user annotations (e.g., scribbles, boxes). Experimental results on various medical datasets demonstrate the superiority of our approach in comparison with existing techniques.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge