Donghyeon Kim
Adverse Weather Image Translation with Asymmetric and Uncertainty-aware GAN
Dec 08, 2021
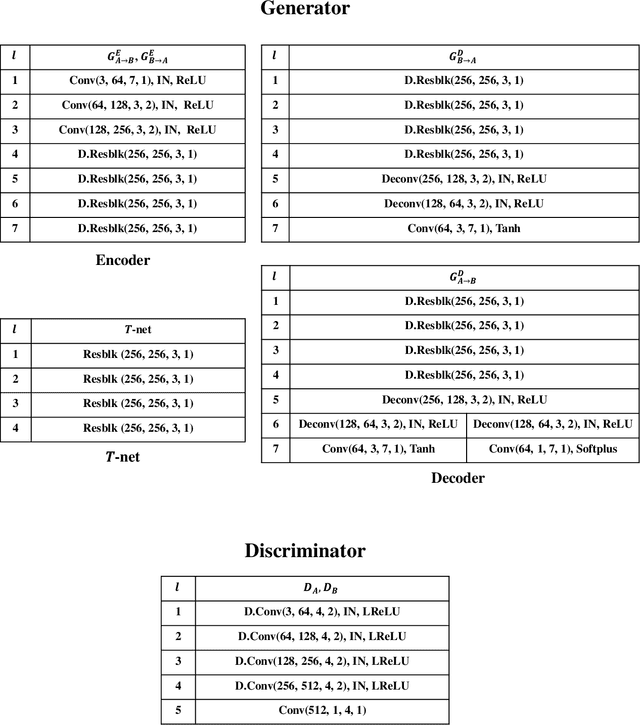


Abstract:Adverse weather image translation belongs to the unsupervised image-to-image (I2I) translation task which aims to transfer adverse condition domain (eg, rainy night) to standard domain (eg, day). It is a challenging task because images from adverse domains have some artifacts and insufficient information. Recently, many studies employing Generative Adversarial Networks (GANs) have achieved notable success in I2I translation but there are still limitations in applying them to adverse weather enhancement. Symmetric architecture based on bidirectional cycle-consistency loss is adopted as a standard framework for unsupervised domain transfer methods. However, it can lead to inferior translation result if the two domains have imbalanced information. To address this issue, we propose a novel GAN model, i.e., AU-GAN, which has an asymmetric architecture for adverse domain translation. We insert a proposed feature transfer network (${T}$-net) in only a normal domain generator (i.e., rainy night-> day) to enhance encoded features of the adverse domain image. In addition, we introduce asymmetric feature matching for disentanglement of encoded features. Finally, we propose uncertainty-aware cycle-consistency loss to address the regional uncertainty of a cyclic reconstructed image. We demonstrate the effectiveness of our method by qualitative and quantitative comparisons with state-of-the-art models. Codes are available at https://github.com/jgkwak95/AU-GAN.
Lightweight dynamic filter for keyword spotting
Sep 27, 2021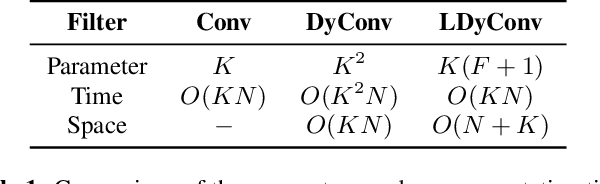
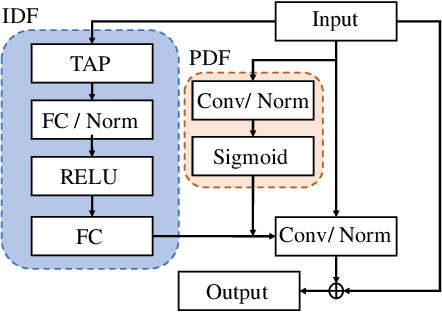
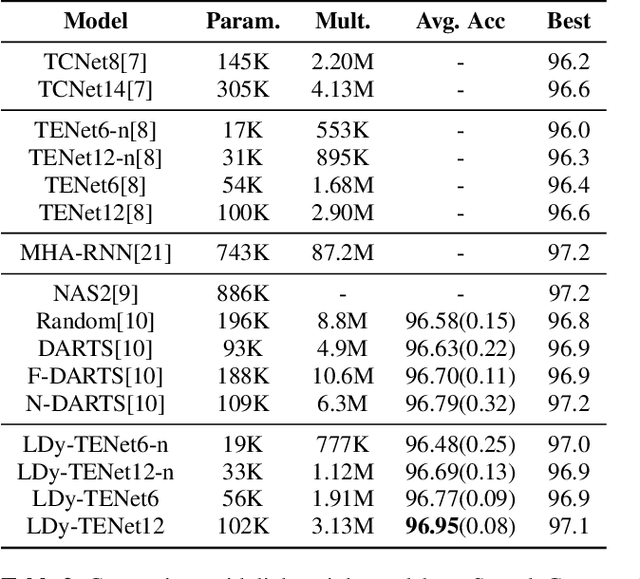
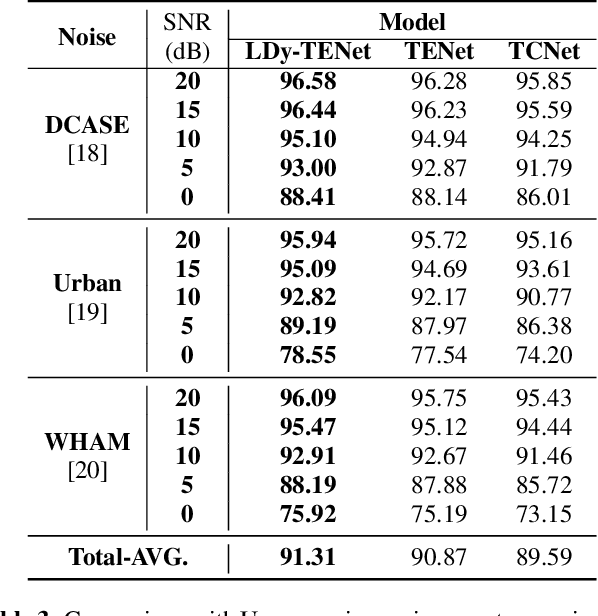
Abstract:Keyword Spotting (KWS) from speech signal is widely applied for being fully hands free speech recognition. The KWS network is designed as a small footprint model to be constantly monitored. Recently, dynamic filter based models are applied in deep learning applications to enhance a system's robustness or accuracy. However, as a dynamic filter framework requires high computational cost, the usage is limited to the condition of the device. In this paper, we proposed a lightweight dynamic filter to improve the performance of KWS. Our proposed model divides dynamic filter as two branches to reduce the computational complexity. This lightweight dynamic filter is applied to the front-end of KWS to enhance the separability of the input data. The experiments show that our model is robustly working on unseen noise and small training data environment by using small computational resource.
Transferability of Natural Language Inference to Biomedical Question Answering
Jul 01, 2020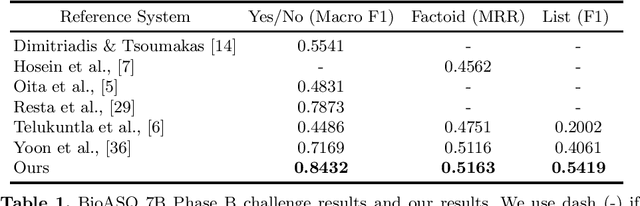
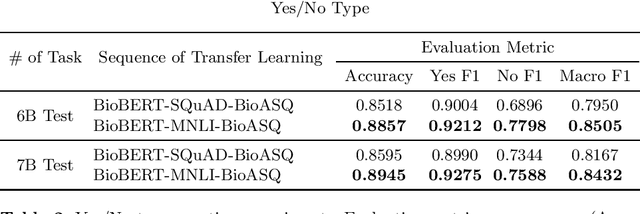
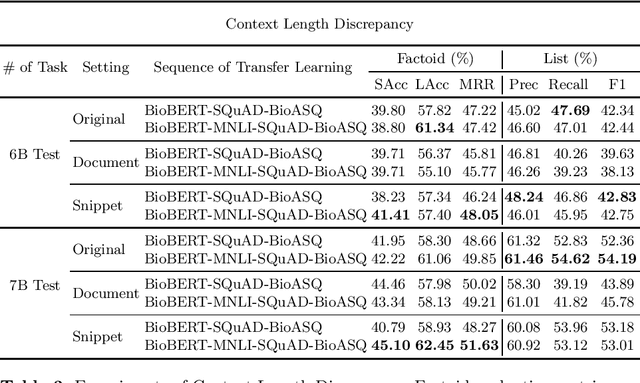
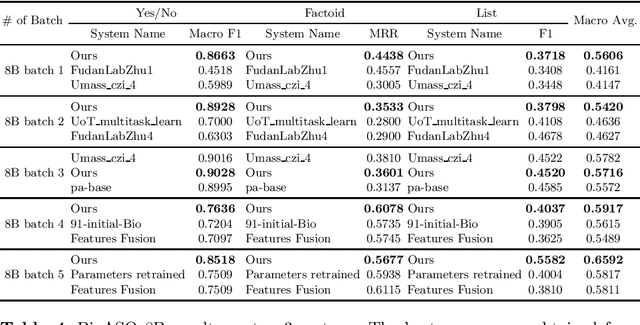
Abstract:Biomedical question answering (QA) is a challenging problem due to the scarcity of data and the requirement of domain expertise. Growing interests of using pre-trained language models with transfer learning address the issue to some extent. Recently, learning linguistic knowledge of entailment in sentence pairs enhances the performance in general domain QA by leveraging such transferability between the two tasks. In this paper, we focus on facilitating the transferability by unifying the experimental setup from natural language inference (NLI) to biomedical QA. We observe that transferring from entailment data shows effective performance on Yes/No (+5.59%), Factoid (+0.53%), List (+13.58%) type questions compared to previous challenge reports (BioASQ 7B Phase B). We also observe that our method generally performs well in the 8th BioASQ Challenge (Phase B). For sequential transfer learning, the order of how tasks are fine-tuned is important. In factoid- and list-type questions, we thoroughly analyze an intrinsic limitation of the extractive QA setting when these questions are converted to the same format of the Stanford Question Answering Dataset (SQuAD).
Pre-trained Language Model for Biomedical Question Answering
Sep 18, 2019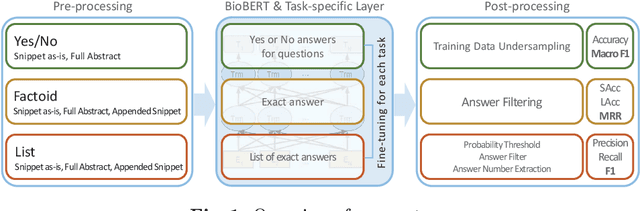
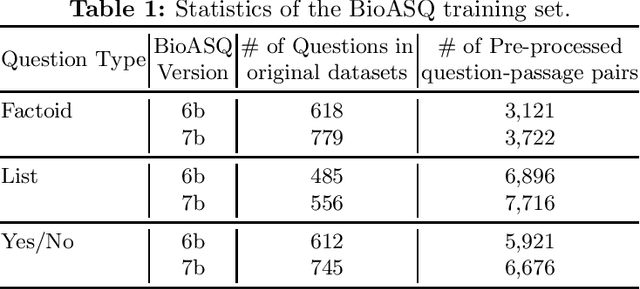
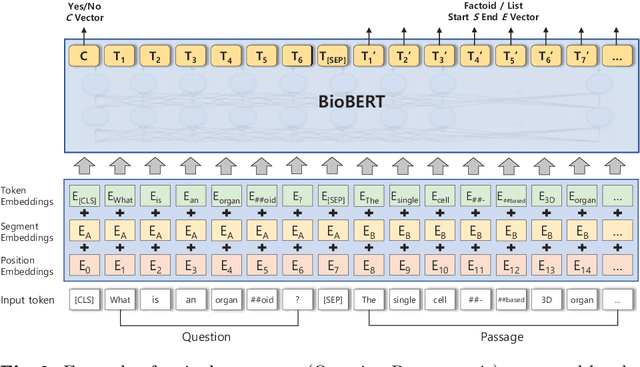
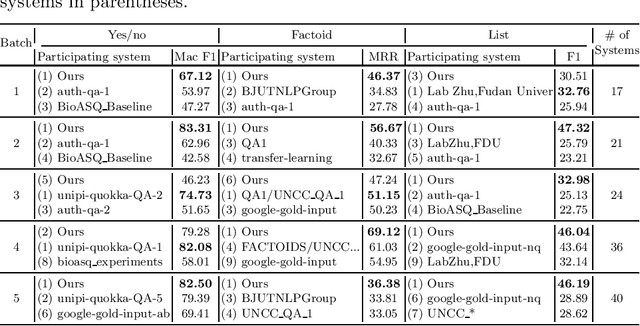
Abstract:The recent success of question answering systems is largely attributed to pre-trained language models. However, as language models are mostly pre-trained on general domain corpora such as Wikipedia, they often have difficulty in understanding biomedical questions. In this paper, we investigate the performance of BioBERT, a pre-trained biomedical language model, in answering biomedical questions including factoid, list, and yes/no type questions. BioBERT uses almost the same structure across various question types and achieved the best performance in the 7th BioASQ Challenge (Task 7b, Phase B). BioBERT pre-trained on SQuAD or SQuAD 2.0 easily outperformed previous state-of-the-art models. BioBERT obtains the best performance when it uses the appropriate pre-/post-processing strategies for questions, passages, and answers.
BioBERT: a pre-trained biomedical language representation model for biomedical text mining
Feb 03, 2019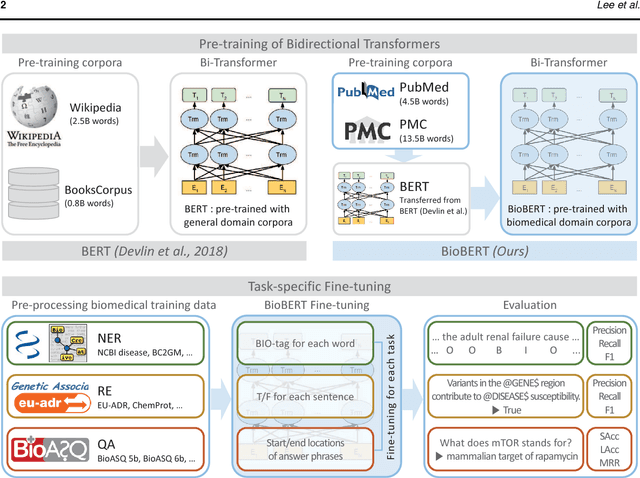
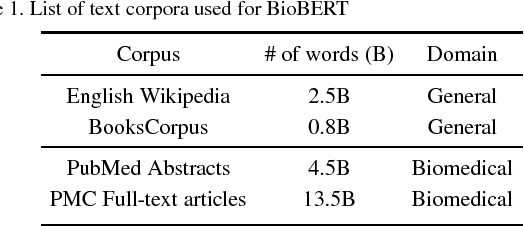

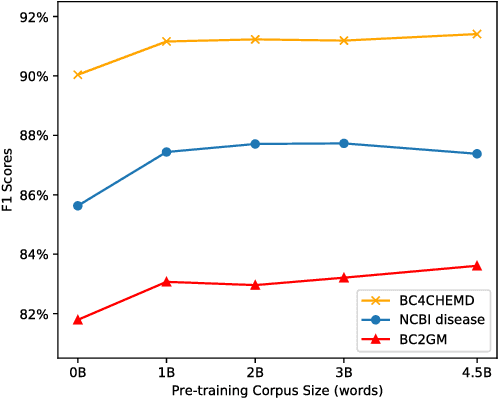
Abstract:Biomedical text mining is becoming increasingly important as the number of biomedical documents rapidly grows. With the progress in machine learning, extracting valuable information from biomedical literature has gained popularity among researchers, and deep learning has boosted the development of effective biomedical text mining models. However, as deep learning models require a large amount of training data, applying deep learning to biomedical text mining is often unsuccessful due to the lack of training data in biomedical fields. Recent researches on training contextualized language representation models on text corpora shed light on the possibility of leveraging a large number of unannotated biomedical text corpora. We introduce BioBERT (Bidirectional Encoder Representations from Transformers for Biomedical Text Mining), which is a domain specific language representation model pre-trained on large-scale biomedical corpora. Based on the BERT architecture, BioBERT effectively transfers the knowledge from a large amount of biomedical texts to biomedical text mining models with minimal task-specific architecture modifications. While BERT shows competitive performances with previous state-of-the-art models, BioBERT significantly outperforms them on the following three representative biomedical text mining tasks: biomedical named entity recognition (0.51% absolute improvement), biomedical relation extraction (3.49% absolute improvement), and biomedical question answering (9.61% absolute improvement). We make the pre-trained weights of BioBERT freely available at https://github.com/naver/biobert-pretrained, and the source code for fine-tuning BioBERT available at https://github.com/dmis-lab/biobert.
Learning User Preferences and Understanding Calendar Contexts for Event Scheduling
Oct 17, 2018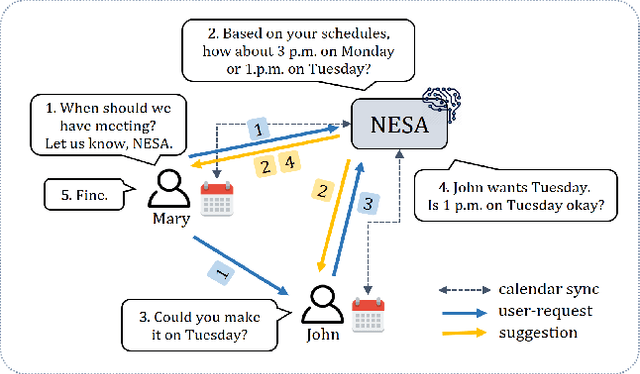
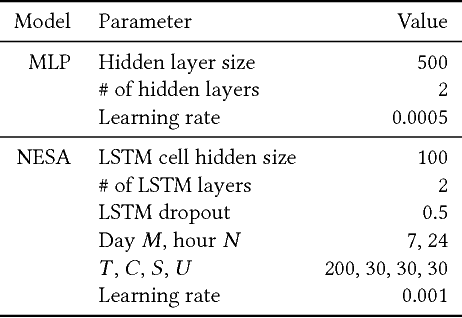
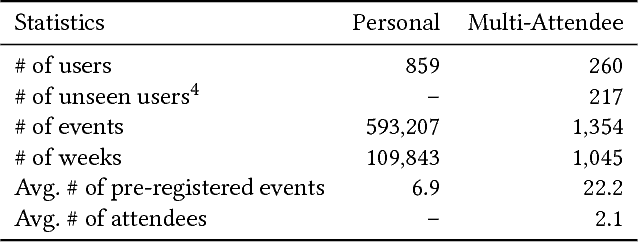
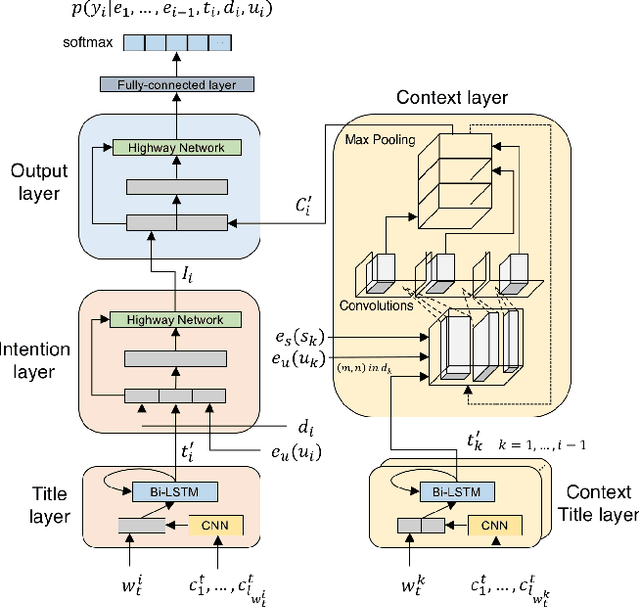

Abstract:With online calendar services gaining popularity worldwide, calendar data has become one of the richest context sources for understanding human behavior. However, event scheduling is still time-consuming even with the development of online calendars. Although machine learning based event scheduling models have automated scheduling processes to some extent, they often fail to understand subtle user preferences and complex calendar contexts with event titles written in natural language. In this paper, we propose Neural Event Scheduling Assistant (NESA) which learns user preferences and understands calendar contexts, directly from raw online calendars for fully automated and highly effective event scheduling. We leverage over 593K calendar events for NESA to learn scheduling personal events, and we further utilize NESA for multi-attendee event scheduling. NESA successfully incorporates deep neural networks such as Bidirectional Long Short-Term Memory, Convolutional Neural Network, and Highway Network for learning the preferences of each user and understanding calendar context based on natural languages. The experimental results show that NESA significantly outperforms previous baseline models in terms of various evaluation metrics on both personal and multi-attendee event scheduling tasks. Our qualitative analysis demonstrates the effectiveness of each layer in NESA and learned user preferences.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge