David Sontag
PClean: Bayesian Data Cleaning at Scale with Domain-Specific Probabilistic Programming
Aug 07, 2020
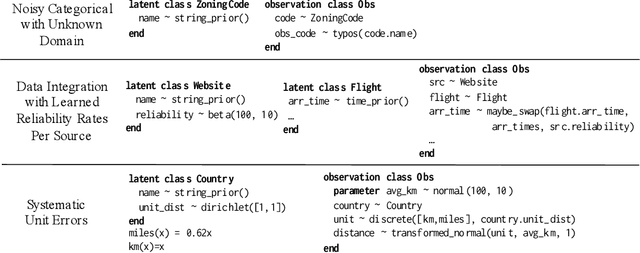
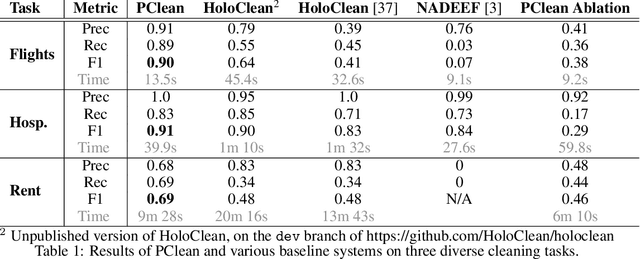

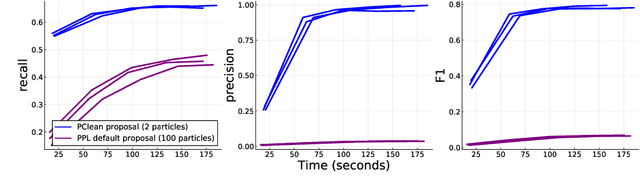
Abstract:Data cleaning is naturally framed as probabilistic inference in a generative model, combining a prior distribution over ground-truth databases with a likelihood that models the noisy channel by which the data are filtered, corrupted, and joined to yield incomplete, dirty, and denormalized datasets. Based on this view, we present PClean, a unified generative modeling architecture for cleaning and normalizing dirty data in diverse domains. Given an unclean dataset and a probabilistic program encoding relevant domain knowledge, PClean learns a structured representation of the data as a relational database of interrelated objects, and uses this latent structure to impute missing values, identify duplicates, detect errors, and propose corrections in the original data table. PClean makes three modeling and inference contributions: (i) a domain-general non-parametric generative model of relational data, for inferring latent objects and their network of latent connections; (ii) a domain-specific probabilistic programming language, for encoding domain knowledge specific to each dataset being cleaned; and (iii) a domain-general inference engine that adapts to each PClean program by constructing data-driven proposals used in sequential Monte Carlo and particle Gibbs. We show empirically that short (< 50-line) PClean programs deliver higher accuracy than state-of-the-art data cleaning systems based on machine learning and weighted logic; that PClean's inference algorithm is faster than generic particle Gibbs inference for probabilistic programs; and that PClean scales to large real-world datasets with millions of rows.
Robust Benchmarking for Machine Learning of Clinical Entity Extraction
Jul 31, 2020
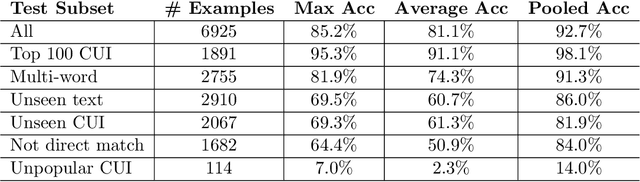
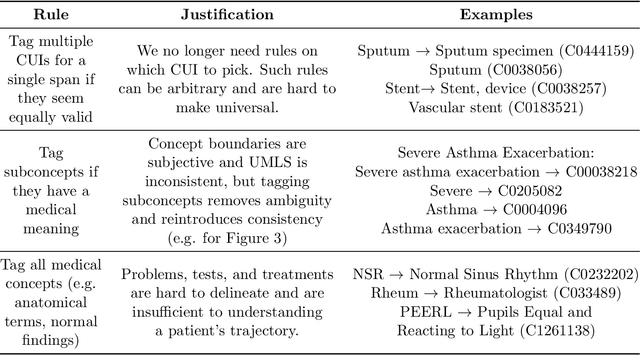
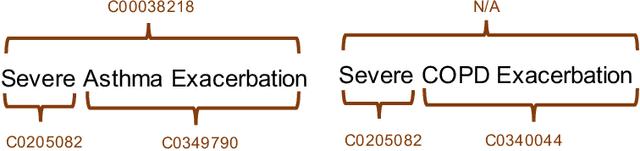
Abstract:Clinical studies often require understanding elements of a patient's narrative that exist only in free text clinical notes. To transform notes into structured data for downstream use, these elements are commonly extracted and normalized to medical vocabularies. In this work, we audit the performance of and indicate areas of improvement for state-of-the-art systems. We find that high task accuracies for clinical entity normalization systems on the 2019 n2c2 Shared Task are misleading, and underlying performance is still brittle. Normalization accuracy is high for common concepts (95.3%), but much lower for concepts unseen in training data (69.3%). We demonstrate that current approaches are hindered in part by inconsistencies in medical vocabularies, limitations of existing labeling schemas, and narrow evaluation techniques. We reformulate the annotation framework for clinical entity extraction to factor in these issues to allow for robust end-to-end system benchmarking. We evaluate concordance of annotations from our new framework between two annotators and achieve a Jaccard similarity of 0.73 for entity recognition and an agreement of 0.83 for entity normalization. We propose a path forward to address the demonstrated need for the creation of a reference standard to spur method development in entity recognition and normalization.
Fast, Structured Clinical Documentation via Contextual Autocomplete
Jul 29, 2020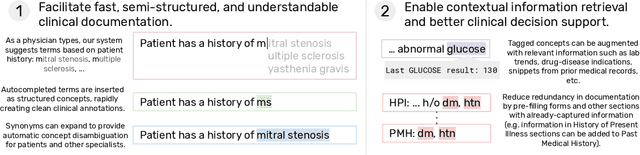

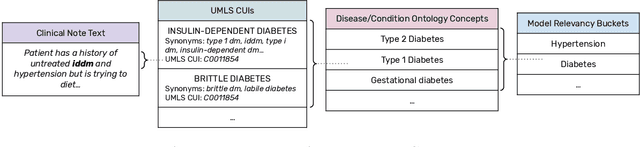

Abstract:We present a system that uses a learned autocompletion mechanism to facilitate rapid creation of semi-structured clinical documentation. We dynamically suggest relevant clinical concepts as a doctor drafts a note by leveraging features from both unstructured and structured medical data. By constraining our architecture to shallow neural networks, we are able to make these suggestions in real time. Furthermore, as our algorithm is used to write a note, we can automatically annotate the documentation with clean labels of clinical concepts drawn from medical vocabularies, making notes more structured and readable for physicians, patients, and future algorithms. To our knowledge, this system is the only machine learning-based documentation utility for clinical notes deployed in a live hospital setting, and it reduces keystroke burden of clinical concepts by 67% in real environments.
Deep Contextual Clinical Prediction with Reverse Distillation
Jul 10, 2020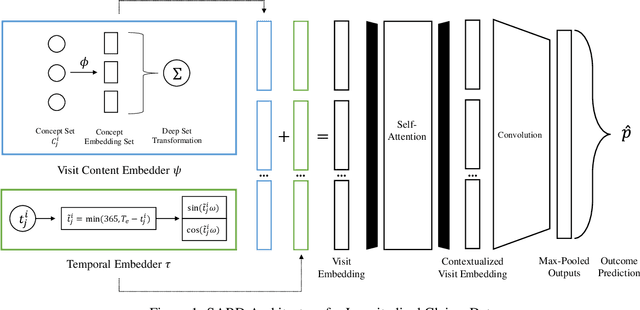
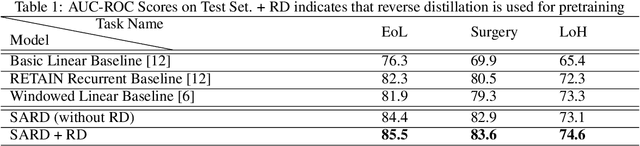

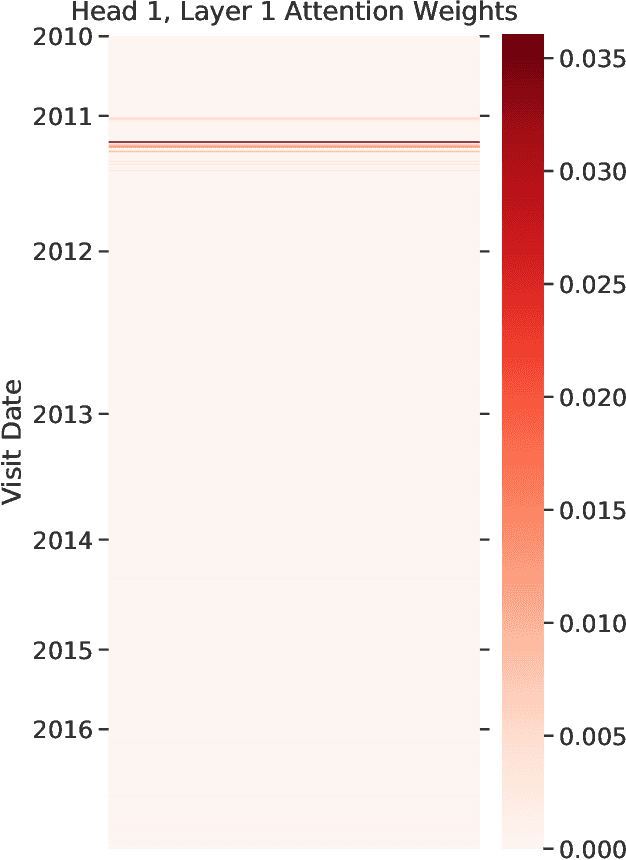
Abstract:Healthcare providers are increasingly using learned methods to predict and understand long-term patient outcomes in order to make meaningful interventions. However, despite innovations in this area, deep learning models often struggle to match performance of shallow linear models in predicting these outcomes, making it difficult to leverage such techniques in practice. In this work, motivated by the task of clinical prediction from insurance claims, we present a new technique called reverse distillation which pretrains deep models by using high-performing linear models for initialization. We make use of the longitudinal structure of insurance claims datasets to develop Self Attention with Reverse Distillation, or SARD, an architecture that utilizes a combination of contextual embedding, temporal embedding and self-attention mechanisms and most critically is trained via reverse distillation. SARD outperforms state-of-the-art methods on multiple clinical prediction outcomes, with ablation studies revealing that reverse distillation is a primary driver of these improvements.
Consistent Estimators for Learning to Defer to an Expert
Jun 02, 2020
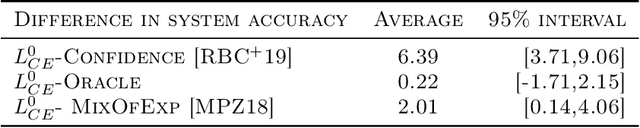
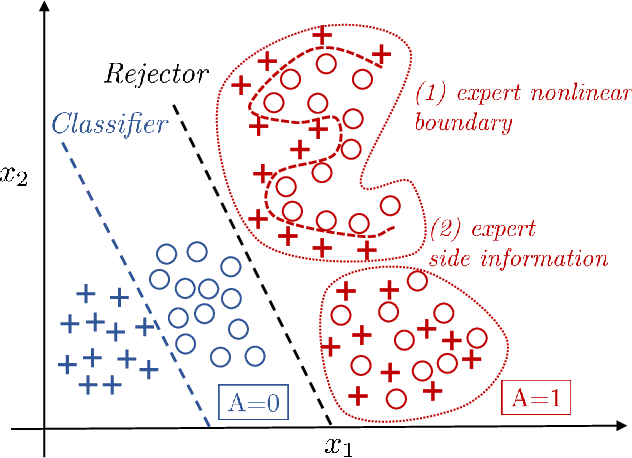

Abstract:Learning algorithms are often used in conjunction with expert decision makers in practical scenarios, however this fact is largely ignored when designing these algorithms. In this paper we explore how to learn predictors that can either predict or choose to defer the decision to a downstream expert. Given only samples of the expert's decisions, we give a procedure based on learning a classifier and a rejector and analyze it theoretically. Our approach is based on a novel reduction to cost sensitive learning where we give a consistent surrogate loss for cost sensitive learning that generalizes the cross entropy loss. We show the effectiveness of our approach on a variety of experimental tasks.
Treatment Policy Learning in Multiobjective Settings with Fully Observed Outcomes
Jun 01, 2020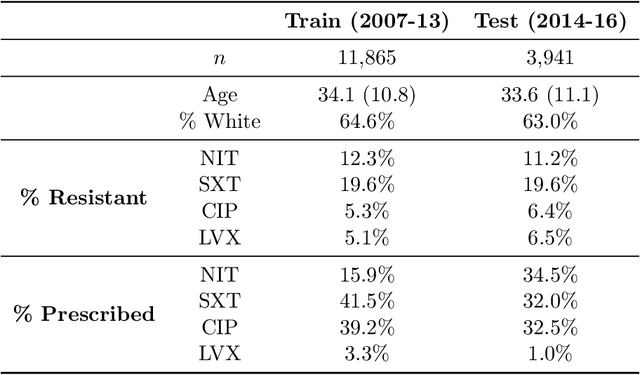
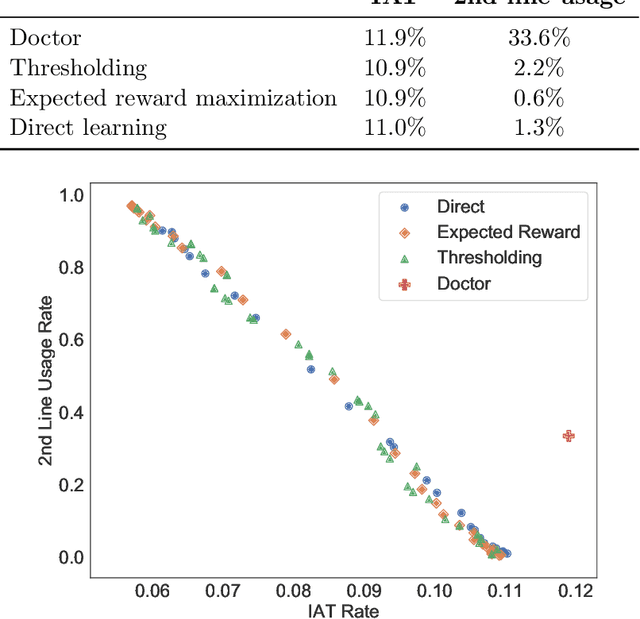
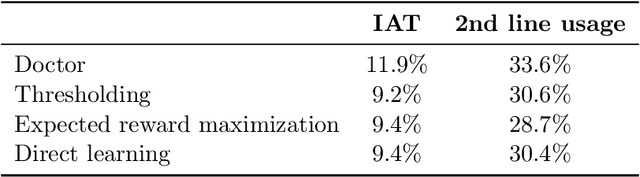
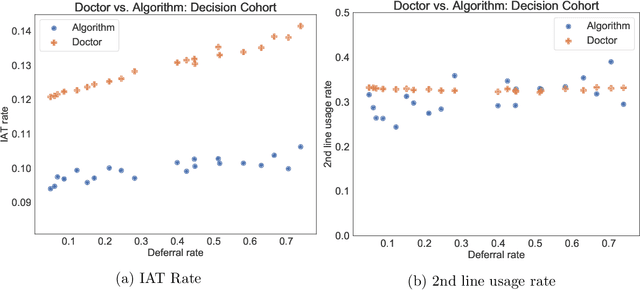
Abstract:In several medical decision-making problems, such as antibiotic prescription, laboratory testing can provide precise indications for how a patient will respond to different treatment options. This enables us to "fully observe" all potential treatment outcomes, but while present in historical data, these results are infeasible to produce in real-time at the point of the initial treatment decision. Moreover, treatment policies in these settings often need to trade off between multiple competing objectives, such as effectiveness of treatment and harmful side effects. We present, compare, and evaluate three approaches for learning individualized treatment policies in this setting: First, we consider two indirect approaches, which use predictive models of treatment response to construct policies optimal for different trade-offs between objectives. Second, we consider a direct approach that constructs such a set of policies without any intermediate models of outcomes. Using a medical dataset of Urinary Tract Infection (UTI) patients, we show that all approaches are able to find policies that achieve strictly better performance on all outcomes than clinicians, while also trading off between different objectives as desired. We demonstrate additional benefits of the direct approach, including flexibly incorporating other goals such as deferral to physicians on simple cases.
Knowledge Base Completion for Constructing Problem-Oriented Medical Records
Apr 27, 2020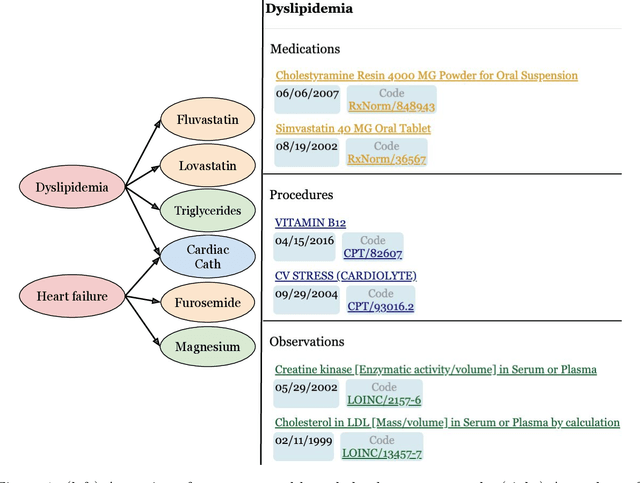
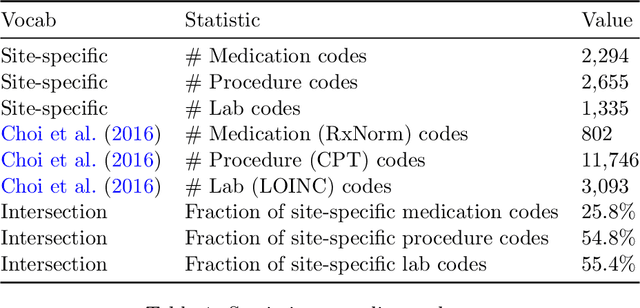

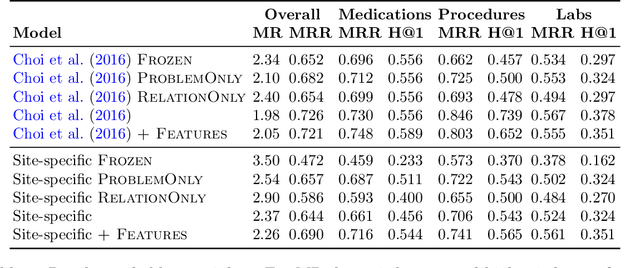
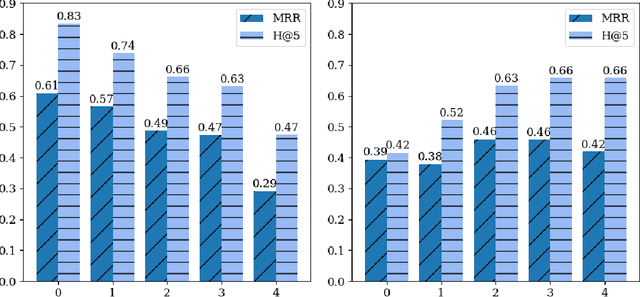
Abstract:Both electronic health records and personal health records are typically organized by data type, with medical problems, medications, procedures, and laboratory results chronologically sorted in separate areas of the chart. As a result, it can be difficult to find all of the relevant information for answering a clinical question about a given medical problem. A promising alternative is to instead organize by problems, with related medications, procedures, and other pertinent information all grouped together. A recent effort by Buchanan (2017) manually defined, through expert consensus, 11 medical problems and the relevant labs and medications for each. We show how to use machine learning on electronic health records to instead automatically construct these problem-based groupings of relevant medications, procedures, and laboratory tests. We formulate the learning task as one of knowledge base completion, and annotate a dataset that expands the set of problems from 11 to 32. We develop a model architecture that exploits both pre-trained concept embeddings and usage data relating the concepts contained in a longitudinal dataset from a large health system. We evaluate our algorithms' ability to suggest relevant medications, procedures, and lab tests, and find that the approach provides feasible suggestions even for problems that are hidden during training.
Generalization Bounds and Representation Learning for Estimation of Potential Outcomes and Causal Effects
Jan 21, 2020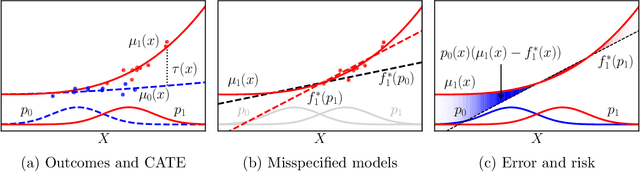
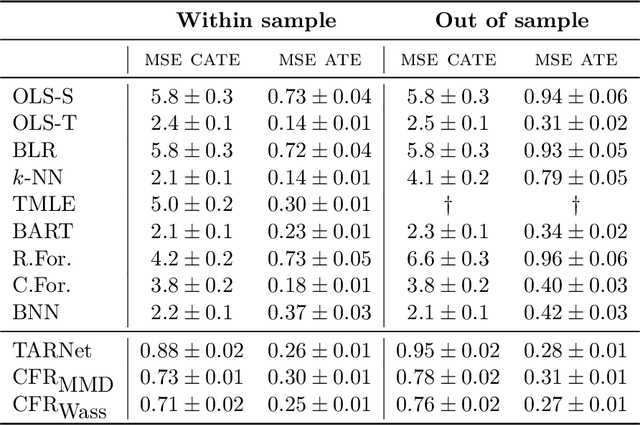
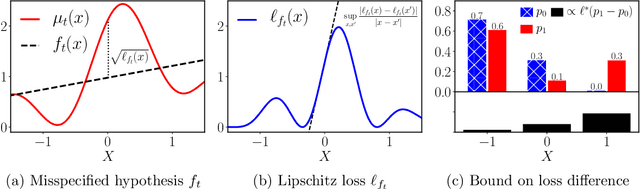
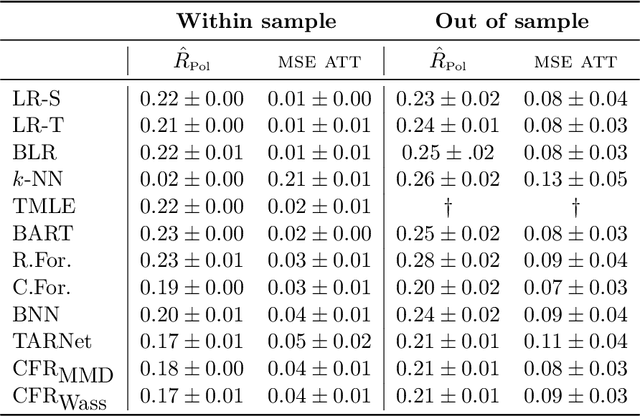
Abstract:Practitioners in diverse fields such as healthcare, economics and education are eager to apply machine learning to improve decision making. The cost and impracticality of performing experiments and a recent monumental increase in electronic record keeping has brought attention to the problem of evaluating decisions based on non-experimental observational data. This is the setting of this work. In particular, we study estimation of individual-level causal effects, such as a single patient's response to alternative medication, from recorded contexts, decisions and outcomes. We give generalization bounds on the error in estimated effects based on distance measures between groups receiving different treatments, allowing for sample re-weighting. We provide conditions under which our bound is tight and show how it relates to results for unsupervised domain adaptation. Led by our theoretical results, we devise representation learning algorithms that minimize our bound, by regularizing the representation's induced treatment group distance, and encourage sharing of information between treatment groups. We extend these algorithms to simultaneously learn a weighted representation to further reduce treatment group distances. Finally, an experimental evaluation on real and synthetic data shows the value of our proposed representation architecture and regularization scheme.
Estimation of Utility-Maximizing Bounds on Potential Outcomes
Oct 10, 2019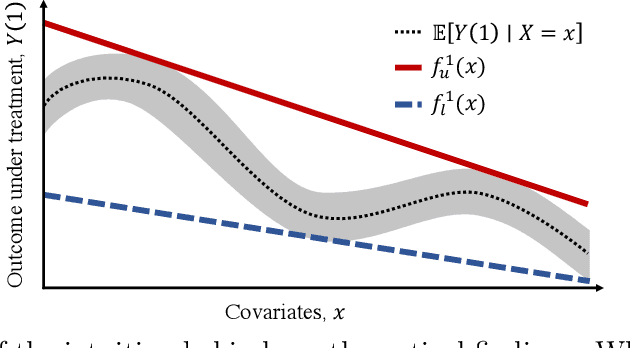
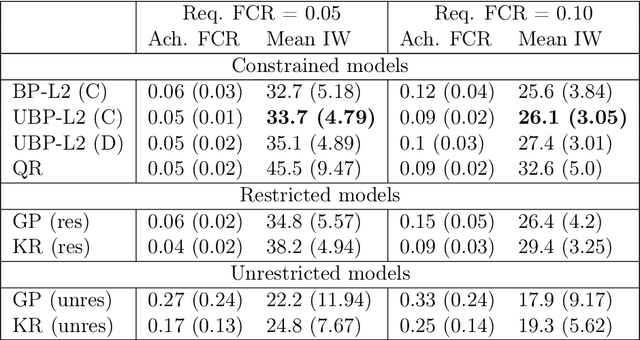
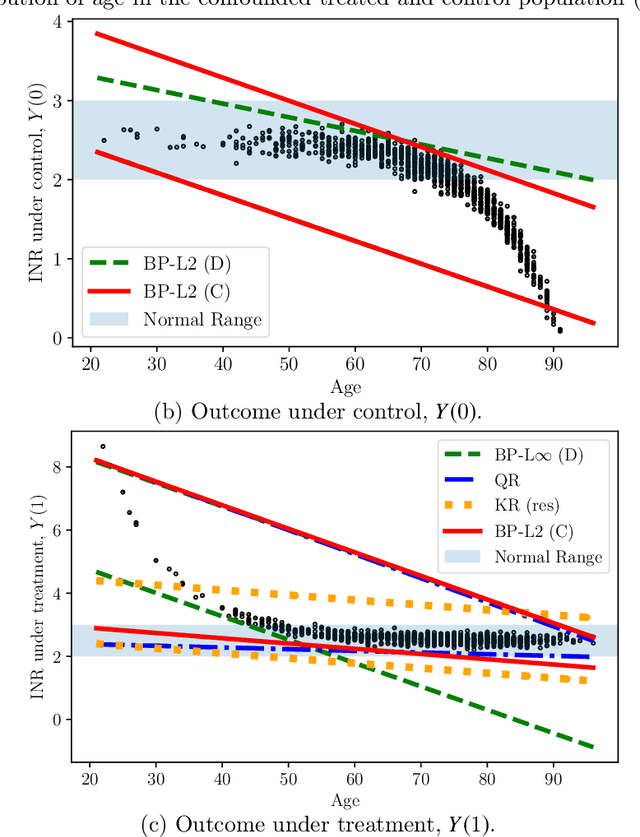
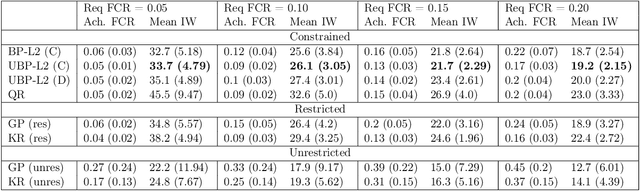
Abstract:Estimation of individual treatment effects is often used as the basis for contextual decision making in fields such as healthcare, education, and economics. However, in many real-world applications it is sufficient for the decision maker to have upper and lower bounds on the potential outcomes of decision alternatives, allowing them to evaluate the trade-off between benefit and risk. With this in mind, we develop an algorithm for directly learning upper and lower bounds on the potential outcomes under treatment and non-treatment. Our theoretical analysis highlights a trade-off between the complexity of the learning task and the confidence with which the resulting bounds cover the true potential outcomes; the more confident we wish to be, the more complex the learning task is. We suggest a novel algorithm that maximizes a utility function while maintaining valid potential outcome bounds. We illustrate different properties of our algorithm, and highlight how it can be used to guide decision making using two semi-simulated datasets.
Open Set Medical Diagnosis
Oct 07, 2019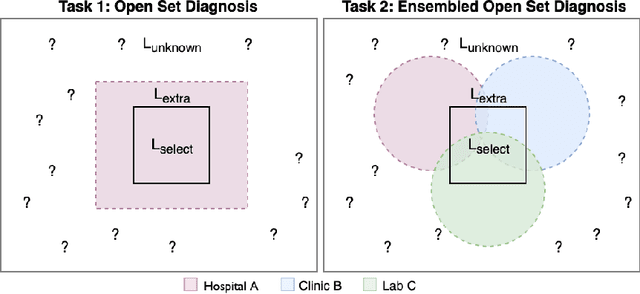
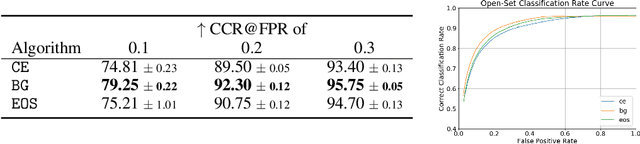
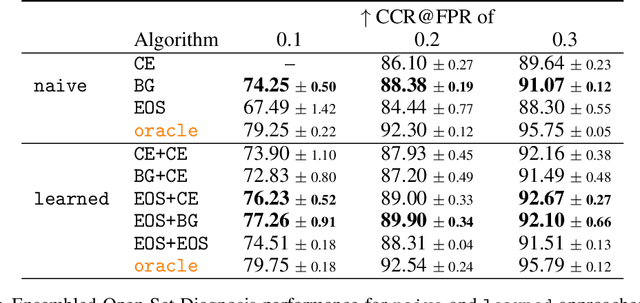
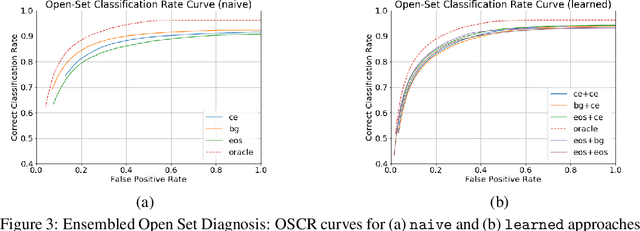
Abstract:Machine-learned diagnosis models have shown promise as medical aides but are trained under a closed-set assumption, i.e. that models will only encounter conditions on which they have been trained. However, it is practically infeasible to obtain sufficient training data for every human condition, and once deployed such models will invariably face previously unseen conditions. We frame machine-learned diagnosis as an open-set learning problem, and study how state-of-the-art approaches compare. Further, we extend our study to a setting where training data is distributed across several healthcare sites that do not allow data pooling, and experiment with different strategies of building open-set diagnostic ensembles. Across both settings, we observe consistent gains from explicitly modeling unseen conditions, but find the optimal training strategy to vary across settings.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge