Daphna Link Sourani
Bootstrapped Physically-Primed Neural Networks for Robust T2 Distribution Estimation in Low-SNR Pancreatic MRI
Mar 14, 2026Abstract:Estimating multi-component T2 relaxation distributions from Multi-Echo Spin Echo (MESE) MRI is a severely ill-posed inverse problem, traditionally solved using regularized non-negative least squares (NNLS). In abdominal imaging, particularly the pancreas, low SNR and residual uncorrelated noise challenge classical solvers and deterministic deep learning models. We introduce a bootstrap-based inference framework for robust distributional T2 estimation that performs stochastic resampling of the echo train and aggregates predictions across multiple subsets. This treats the acquisition as a distribution rather than a fixed input, yielding variance-reduced, physically consistent estimates and converting deterministic relaxometry networks into probabilistic ensemble predictors. Applied to the P2T2 architecture, our method uses inference-time bootstrapping to smooth noise artifacts and enhance fidelity to the underlying relaxation distribution. Noninvasive pancreatic evaluation is limited by location and biopsy risks, highlighting the need for biomarkers capable of capturing early pathophysiological changes. In type 1 diabetes (T1DM), progressive beta-cell destruction begins years before overt hyperglycemia, yet current imaging cannot assess early islet decline. We evaluate clinical utility via a test-retest reproducibility study (N=7) and a T1DM versus healthy differentiation task (N=8). Our approach achieves the lowest Wasserstein distances across repeated scans and superior sensitivity to physiology-driven shifts in the relaxation-time distribution, outperforming NNLS and deterministic deep learning baselines. These results establish inference-time bootstrapping as an effective enhancement for quantitative T2 relaxometry in low-SNR abdominal imaging.
Contour Dice loss for structures with Fuzzy and Complex Boundaries in Fetal MRI
Sep 25, 2022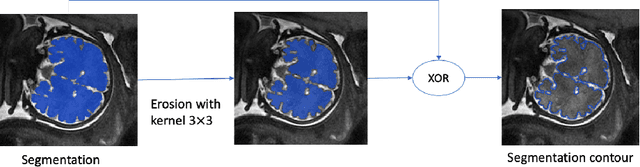
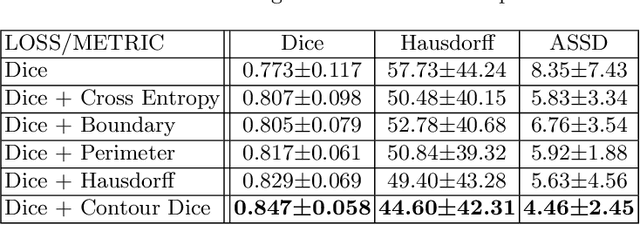
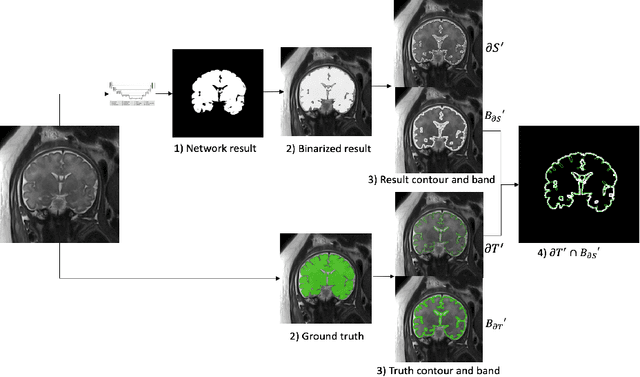
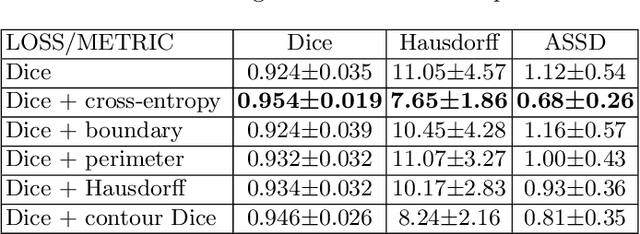
Abstract:Volumetric measurements of fetal structures in MRI are time consuming and error prone and therefore require automatic segmentation. Placenta segmentation and accurate fetal brain segmentation for gyrification assessment are particularly challenging because of the placenta fuzzy boundaries and the fetal brain cortex complex foldings. In this paper, we study the use of the Contour Dice loss for both problems and compare it to other boundary losses and to the combined Dice and Cross-Entropy loss. The loss is computed efficiently for each slice via erosion, dilation and XOR operators. We describe a new formulation of the loss akin to the Contour Dice metric. The combination of the Dice loss and the Contour Dice yielded the best performance for placenta segmentation. For fetal brain segmentation, the best performing loss was the combined Dice with Cross-Entropy loss followed by the Dice with Contour Dice loss, which performed better than other boundary losses.
Partial annotations for the segmentation of large structures with low annotation cost
Sep 25, 2022Abstract:Deep learning methods have been shown to be effective for the automatic segmentation of structures and pathologies in medical imaging. However, they require large annotated datasets, whose manual segmentation is a tedious and time-consuming task, especially for large structures. We present a new method of partial annotations that uses a small set of consecutive annotated slices from each scan with an annotation effort that is equal to that of only few annotated cases. The training with partial annotations is performed by using only annotated blocks, incorporating information about slices outside the structure of interest and modifying a batch loss function to consider only the annotated slices. To facilitate training in a low data regime, we use a two-step optimization process. We tested the method with the popular soft Dice loss for the fetal body segmentation task in two MRI sequences, TRUFI and FIESTA, and compared full annotation regime to partial annotations with a similar annotation effort. For TRUFI data, the use of partial annotations yielded slightly better performance on average compared to full annotations with an increase in Dice score from 0.936 to 0.942, and a substantial decrease in Standard Deviations (STD) of Dice score by 22% and Average Symmetric Surface Distance (ASSD) by 15%. For the FIESTA sequence, partial annotations also yielded a decrease in STD of the Dice score and ASSD metrics by 27.5% and 33% respectively for in-distribution data, and a substantial improvement also in average performance on out-of-distribution data, increasing Dice score from 0.84 to 0.9 and decreasing ASSD from 7.46 to 4.01 mm. The two-step optimization process was helpful for partial annotations for both in-distribution and out-of-distribution data. The partial annotations method with the two-step optimizer is therefore recommended to improve segmentation performance under low data regime.
* 10 pages, 4 figures
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge