Christopher Nguan
Open-H-Embodiment: A Large-Scale Dataset for Enabling Foundation Models in Medical Robotics
Apr 22, 2026Abstract:Autonomous medical robots hold promise to improve patient outcomes, reduce provider workload, democratize access to care, and enable superhuman precision. However, autonomous medical robotics has been limited by a fundamental data problem: existing medical robotic datasets are small, single-embodiment, and rarely shared openly, restricting the development of foundation models that the field needs to advance. We introduce Open-H-Embodiment, the largest open dataset of medical robotic video with synchronized kinematics to date, spanning more than 49 institutions and multiple robotic platforms including the CMR Versius, Intuitive Surgical's da Vinci, da Vinci Research Kit (dVRK), Rob Surgical BiTrack, Virtual Incision's MIRA, Moon Surgical Maestro, and a variety of custom systems, spanning surgical manipulation, robotic ultrasound, and endoscopy procedures. We demonstrate the research enabled by this dataset through two foundation models. GR00T-H is the first open foundation vision-language-action model for medical robotics, which is the only evaluated model to achieve full end-to-end task completion on a structured suturing benchmark (25% of trials vs. 0% for all others) and achieves 64% average success across a 29-step ex vivo suturing sequence. We also train Cosmos-H-Surgical-Simulator, the first action-conditioned world model to enable multi-embodiment surgical simulation from a single checkpoint, spanning nine robotic platforms and supporting in silico policy evaluation and synthetic data generation for the medical domain. These results suggest that open, large-scale medical robot data collection can serve as critical infrastructure for the research community, enabling advances in robot learning, world modeling, and beyond.
The Open Kidney Ultrasound Data Set
Jun 14, 2022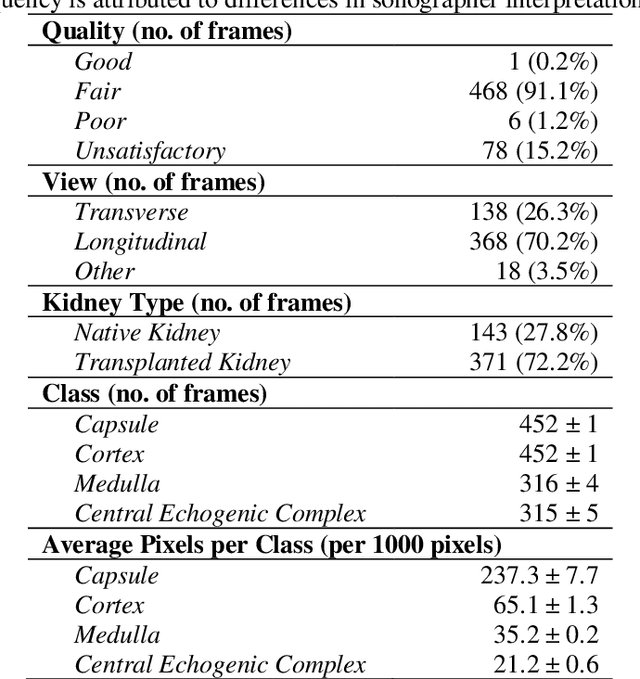
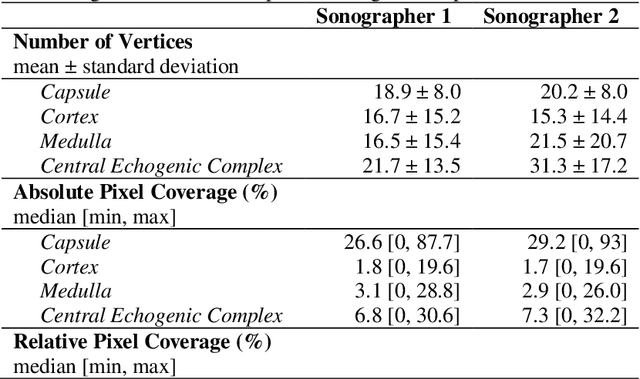

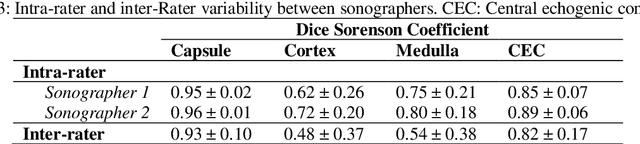
Abstract:Ultrasound use is because of its low cost, non-ionizing, and non-invasive characteristics, and has established itself as a cornerstone radiological examination. Research on ultrasound applications has also expanded, especially with image analysis with machine learning. However, ultrasound data are frequently restricted to closed data sets, with only a few openly available. Despite being a frequently examined organ, the kidney lacks a publicly available ultrasonography data set. The proposed Open Kidney Ultrasound Data Set is the first publicly available set of kidney B-mode ultrasound data that includes annotations for multi-class semantic segmentation. It is based on data retrospectively collected in a 5-year period from over 500 patients with a mean age of 53.2 +/- 14.7 years, body mass index of 27.0 +/- 5.4 kg/m2, and most common primary diseases being diabetes mellitus, IgA nephropathy, and hypertension. There are labels for the view and fine-grained manual annotations from two expert sonographers. Notably, this data includes native and transplanted kidneys. Initial benchmarking measurements are performed, demonstrating a state-of-the-art algorithm achieving a Dice Sorenson Coefficient of 0.74 for the kidney capsule. This data set is a high-quality data set, including two sets of expert annotations, with a larger breadth of images than previously available. In increasing access to kidney ultrasound data, future researchers may be able to create novel image analysis techniques for tissue characterization, disease detection, and prognostication.
The Kidneys Are Not All Normal: Investigating the Speckle Distributions of Transplanted Kidneys
Jun 14, 2022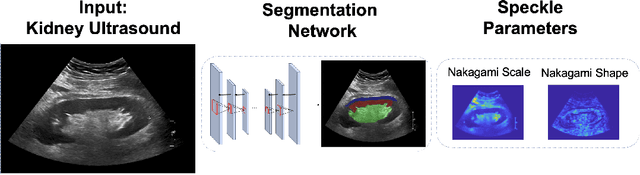
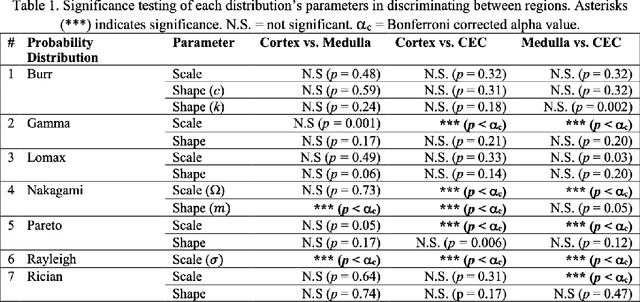
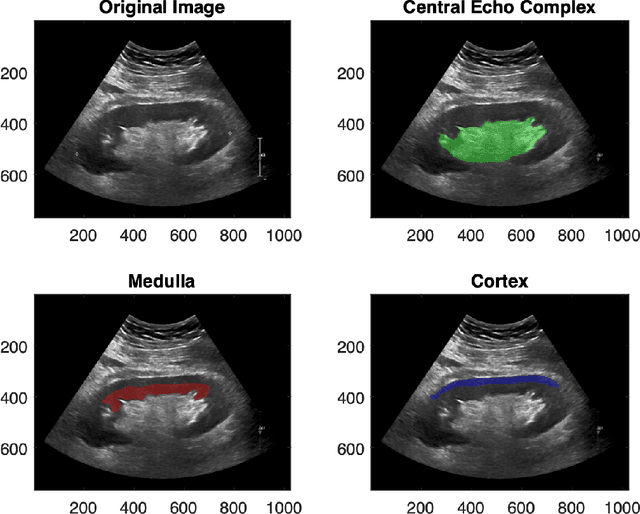
Abstract:Modelling ultrasound speckle has generated considerable interest for its ability to characterize tissue properties. As speckle is dependent on the underlying tissue architecture, modelling it may aid in tasks like segmentation or disease detection. However, for the transplanted kidney where ultrasound is commonly used to investigate dysfunction, it is currently unknown which statistical distribution best characterises such speckle. This is especially true for the regions of the transplanted kidney: the cortex, the medulla and the central echogenic complex. Furthermore, it is unclear how these distributions vary by patient variables such as age, sex, body mass index, primary disease, or donor type. These traits may influence speckle modelling given their influence on kidney anatomy. We are the first to investigate these two aims. N=821 kidney transplant recipient B-mode images were automatically segmented into the cortex, medulla, and central echogenic complex using a neural network. Seven distinct probability distributions were fitted to each region. The Rayleigh and Nakagami distributions had model parameters that differed significantly between the three regions (p <= 0.05). While both had excellent goodness of fit, the Nakagami had higher Kullbeck-Leibler divergence. Recipient age correlated weakly with scale in the cortex (Omega: rho = 0.11, p = 0.004), while body mass index correlated weakly with shape in the medulla (m: rho = 0.08, p = 0.04). Neither sex, primary disease, nor donor type demonstrated any correlation. We propose the Nakagami distribution be used to characterize transplanted kidneys regionally independent of disease etiology and most patient characteristics based on our findings.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge