Chenjie Zeng
Foundation Models to Unlock Real-World Evidence from Nationwide Medical Claims
May 05, 2026Abstract:Evidence derived from large-scale real-world data (RWD) is increasingly informing regulatory evaluation and healthcare decision-making. Administrative claims provide population-scale, longitudinal records of healthcare utilization, expenditure, and detailed coding of diagnoses, procedures, and medications, yet their potential as a substrate for healthcare foundation models remains largely unexplored. Here we present ReClaim, a generative transformer trained from scratch on 43.8 billion medical events from more than 200 million enrollees in the MarketScan claims data spanning 2008-2022. ReClaim models longitudinal trajectories across diagnoses, procedures, medications, and expenditure, and was scaled to 140 million, 700 million, and 1.7 billion parameters. Across over 1,000 disease-onset prediction tasks, ReClaim achieved a mean AUC of 75.6%, substantially outperforming disease-specific LightGBM (66.3%) and the transformer-based Delphi model (69.4%), with the largest gains for rare diseases. These advantages held across retrospective and prospective evaluations and in external validation on two independent datasets. Performance improved monotonically with scale, and post-training added 13.8 percentage points over pre-training alone. Beyond disease prediction, ReClaim captured financial outcomes and improved real-world evidence (RWE) analyses: for healthcare expenditure forecasting it increased explained variance from 0.28 to 0.37 relative to LightGBM, and in a target trial emulation it reduced systematic bias by 72% on average relative to Delphi. Together, these results establish administrative claims as a scalable substrate for healthcare foundation models and show that learned representations generalize across time periods and data sources, supporting disease surveillance, expenditure forecasting, and RWE generation.
Ontologizing Health Systems Data at Scale: Making Translational Discovery a Reality
Sep 10, 2022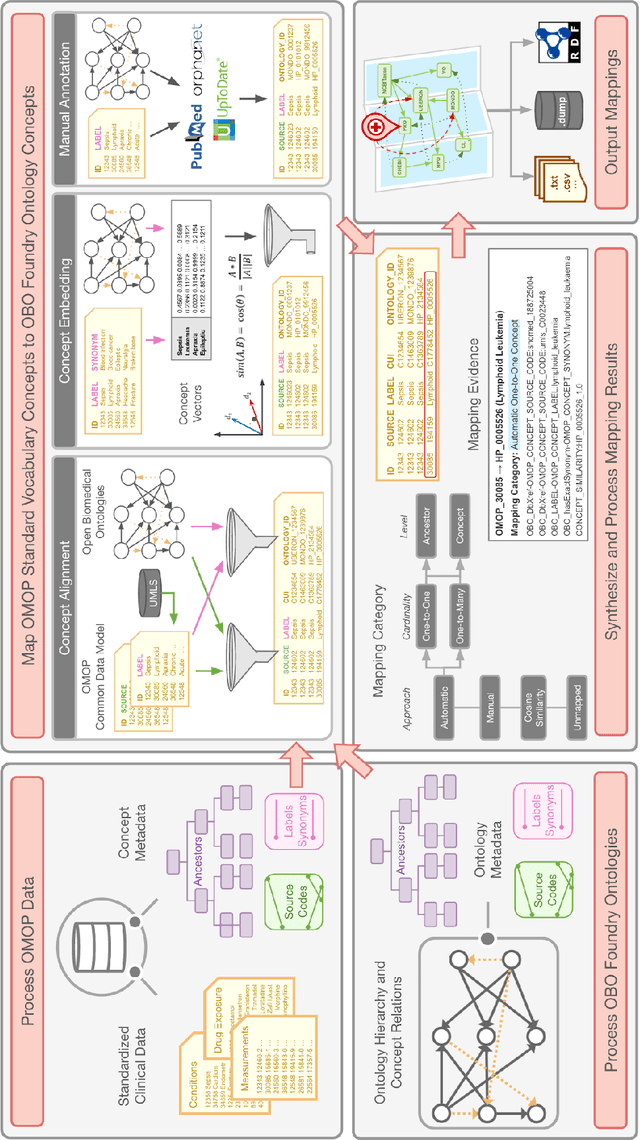
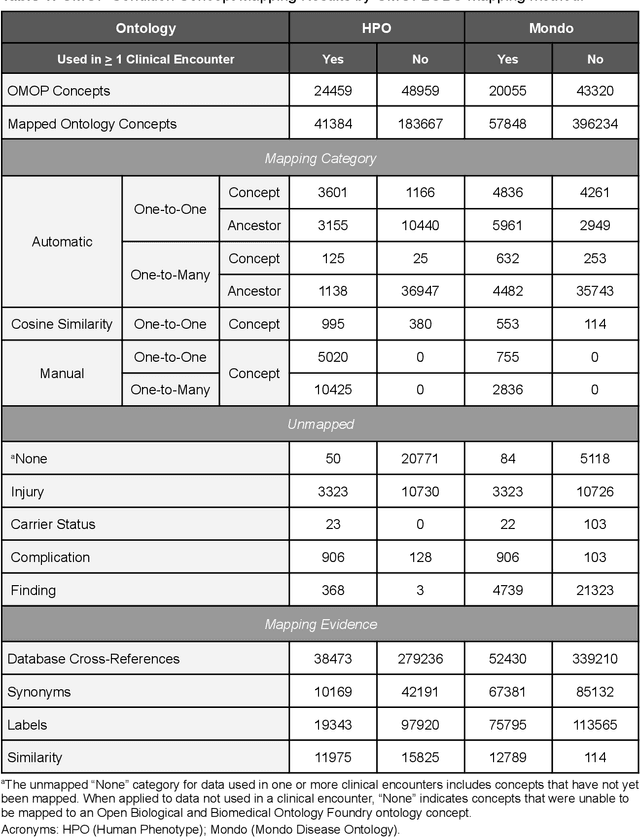
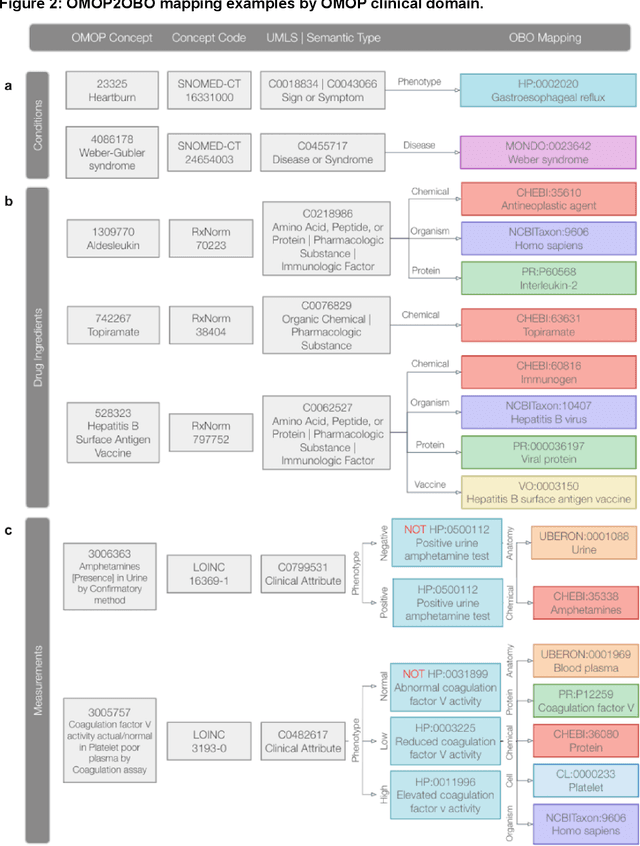
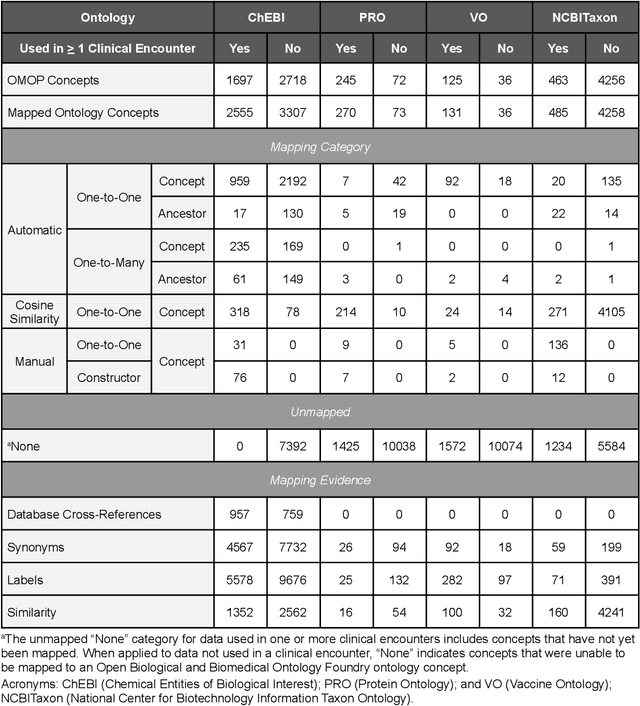
Abstract:Common data models solve many challenges of standardizing electronic health record (EHR) data, but are unable to semantically integrate the resources needed for deep phenotyping. Open Biological and Biomedical Ontology (OBO) Foundry ontologies provide semantically computable representations of biological knowledge and enable the integration of a variety of biomedical data. However, mapping EHR data to OBO Foundry ontologies requires significant manual curation and domain expertise. We introduce a framework for mapping Observational Medical Outcomes Partnership (OMOP) standard vocabularies to OBO Foundry ontologies. Using this framework, we produced mappings for 92,367 conditions, 8,615 drug ingredients, and 10,673 measurement results. Mapping accuracy was verified by domain experts and when examined across 24 hospitals, the mappings covered 99% of conditions and drug ingredients and 68% of measurements. Finally, we demonstrate that OMOP2OBO mappings can aid in the systematic identification of undiagnosed rare disease patients who might benefit from genetic testing.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge